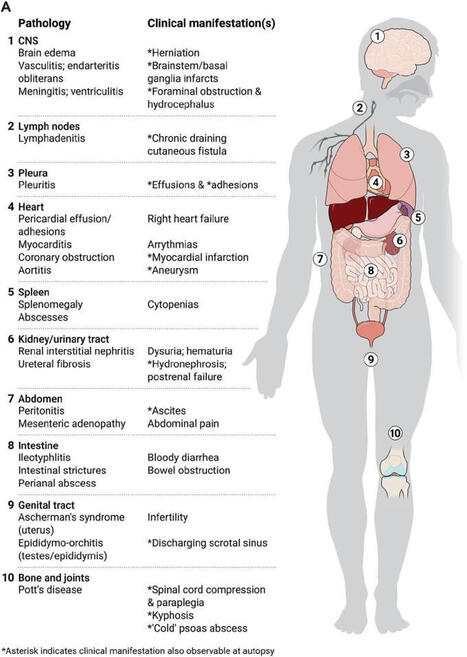
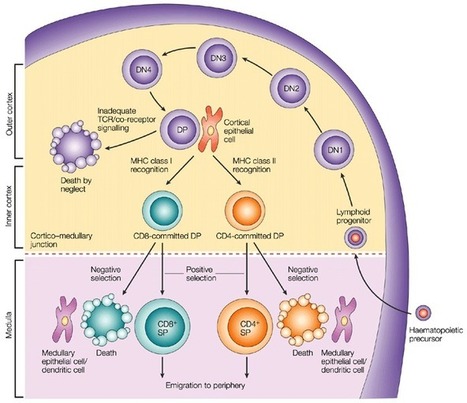
T Lymphocytes Development and markers for major T cell subtypes It is well established that CD4-CD8- T cell precursors migrate to the thymus where they undergo the following phenotypical stages: CD44+CD25- (DN1), CD44+CD25+ (DN2), CD44-CD25+ (DN3), and CD44-CD25- (DN4), followed by the progression of DN4 cells into the double-positive CD4+CD8+ T cells [5] (Figure 1). Infection by various pathogens causes activation and proliferation of naïve T cells, which differentiate into lineages with effector and memory fates. Naive CD4+ T cells recognize antigens presented by major histocompatibility complex (MHC) class II on antigen-presenting cells. Depending on the specific stimuli, the CD4+ T cells can differentiate into various subtypes, including the helper TH1, TH2 and TH17 cells and regulatory T cells (Tregs). A subset of TH2 cells differentiate into allergic disease-related H2A cells, with a CD45RBlow CD27− phenotype and coexpression of the chemoattractant receptor CRTH2, the natural killer cell marker CD161, and the homing receptor CD49d [6]. Memory T cells vary in their surface receptor expression, effector and trafficking abilities. There are four major subsets of memory T cells: central memory, effector memory, tissue-resident memory and stem memory T cells. Multiple signals regulate the differentiation of CD4+ T cells into central and peripheral memory cells. CD4+ T central memory cells express CD62L and CCR7, which are important for their migration [7]. The peripheral T stem cell memory cells express CXCR3 and CD95 molecules. In addition, both naive and memory T-cell subsets express a variety of functional molecules (Table 1). Process Antigen Function Costimulation/Survival CD27 Costimulation CD28 Costimulation CD127 IL-7 signaling PD-1 Inhibition of effector function CD122 IL-2/IL-15 signaling CD132 γc cytokine signaling KLRG-1 Inhibition of effector function Activation HLA DR Peptide presentation CD38 Calcium flux/signal transduction CD69 Proliferation Adhesion CD11a Adhesion to APC/endothelium CD58 Adhesion to APC CD99 Transendothelial migration Migration CD62L Secondary lymphoid tissues homing CD103 Gut homing CCR4 Chemokine response/TH2 associated CCR5 Homing to inflamed tissues CCR6 Chemokine response/TH17 associated CCR9 Gut homing CCR10 Skin homing CXCR3 Homing to inflamed tissues CXCR4 Homing to bone marrow CLA Skin homing Cytolytic molecules Granzyme A Cleavage of cellular proteins Granzyme B Cleavage of cellular proteins Perforin Pore-forming Miscellaneous CD161 Regulation of proliferation/cytotoxicity IL-18Ra 18Ra Response to IL-18 c-Kit Response to SCF CD130 Response to IL-6 The second major group of T cells, CD8+ T cells, mediates direct killing of antigen-presenting target cells. Naive CD8+ T cells are activated upon recognition of antigens presented by MHC class I on dendritic cells in the spleen or lymph nodes. Activated CD8+ T cells expand and become effector CD8+ T cells. The majority of the T cells bear α and β chains in their T cell receptor (TCR). However, there is a population of T cells, which have TCR formed by γ and δ chains. These cells, gamma delta T cells, are significantly enriched in epithelia [9, 10]. Gamma delta T cells regulate immune responses by various mechanisms, including suppression of effector T cell and TH1 cell functions, blockage of neutrophil influx and regulation of antigen-presenting cell activity. Markers for maturation processes Naive T cells are considered as precursors of the majority of antigen-activated T cell subpopulations. Human naïve CD4+ T cells express CD45RA, CCR7, CD62L and CD27. Upon recognition of antigens presented by major histocompatibility complex (MHC) class II on antigen-presenting cells, naïve CD4+ T cells undergo proliferation and differentiation into functionally different T cell subsets including IFN-γ producing helper T cell-1 cells (TH1), IL-4-producing TH2, IL-17-producing TH17 cells, and inducible regulatory T cells (iTregs) (Figure 2). Each T cell subset expresses specific transcription factors, such as T-bet (TH1), GATA3 (TH2), RORγt (TH17), and Foxp3 (CD25+ Tregs). With regard to TH17 cells, their differentiation is under control TGF-β and IL-6-induced differentiation, IL-21-induced activation, and IL-23-regulated stabilization [11, 12]. As to iTregs, FOXP3 was found to an important marker of natural CD4+CD25+ regulatory T cells. Moreover, transfection of CD4+CD25- T cells with Foxp3 stimulates their regulatory activity [13]. In addition, TGF-β was found to be crucial for the differentiation of naive CD4+ T cells into Foxp3+ Tregs [14]. Also, IL-2 is commonly required for TGF-β-regulated iTreg differentiation [15]. With regard to cytotoxic T cells, there are several peripheral subsets of different subsets of CD8+ T cells based on the expression of CD45RA and CCR7: a CD45RA+CCR7+ subset of naive cells, a CD45RA-CCR7+ subset of antigen-experienced memory T cells, a CD45RA-CCR7- effector memory cell subset, and a CD45RA+CCR7− subset of differentiated, antigen-experienced effector cells. Also, there are effector memory CD8+ T cells expressing CD69 and CD103 and residing in non-lymphoid tissues [16, 17]. A subpopulation of CD8+ T cells shows a memory cell phenotype: CD62L-/+CCR7+CD27-/+. Activated cytotoxic CD8+ T cells downregulate expression of L-selectin and CCR7 and upregulate surface expression of CD44, LFA-1 and/or α4β1 integrin. Additional phenotypical markers of Tregs There are several additional phenotypical markers expressed in both human and mouse Tregs. They include CTLA-4, CD103, GITR and OX40. In particular, CTLA-4 is important for both inhibitory functions and homeostasis of Tregs. Intracellular expression of CTLA-4 was observed in CD4+CD25+ human Tregs [18]. Another marker, integrin α (CD103) is expressed by Tregs and CD4+CD25+CD103+ Tregs were demonstrated to produce IL-10 actively [19]. In addition, GITR (CD357) is expressed in CD4+CD25+ human Tregs in peripheral blood [20]. Also, OX40 (CD134) was shown to stimulate the proliferation of CD4+FoxP3+ Tregs [21]. Moreover, OX40 stimulates migration of Tregs into the peripheral lymphoid and other tissues during inflammation [22]. T cell subsets that regulate B cell functions in the germinal centers Several specific T cell subsets, including follicular B helper T cells (TFH), follicular regulatory T cells (TFR) and cytotoxic CD8+ T cells, reside in the germinal centers and regulate the B cell proliferation [23]. Among these T cell subpopulations, TFH cells belong to CD4+ T cells and assist follicular B cells located in secondary lymphoid tissues, such as lymph nodes, spleen, and tonsils. Concerning the specific markers, high expression of CXC-chemokine receptor 5 (CXCR5) characterizes TFH cells [24]. Its interaction with CXC-chemokine ligand 13 (CXCL13) produced by follicular stromal cells mediates the homing of TFH cells into lymphoid follicles [24]. The development of TFH cells is strongly dependent on IL-2 production, as naїve IL2-secreting CD4+ T cells are destined to differentiate into TFH cells, while other CD4+ T cells, which do not produce IL-2, develop into non-TFH cells [25]. In addition to a universal T cell marker Thy1 (CD90) and CXCR5, TFH cells express ICOS and PD-1 molecules. Upregulation of CXCR5 expression stimulates TFH cells to migrate into the germinal centers, where these cells stabilize their phenotype by contacts with local B cells via ICOS-ICOSL binding [26]. Concerning the regulation of TFH functions, γδ T cells (TCRγδ+CXCR5+ T cells), which also reside in the lymph nodes, have recently been shown to present antigens to TFH cells and induce their activation [27]. Cellular interactions with TFH cells regulate the proliferation and maturation of B cells in the germinal centers [28]. Besides, TFH cells secrete Il-4 and IL-21 cytokines, which are crucial for the functioning of the germinal centers [28]. Moreover, in the germinal centers, TFH cells are represented by two distinct subpopulations: IL-21+ T cells regulating the selection of high-affinity B cells and IL-4+ T cells promoting differentiation of plasmocytes [29]. Several studies have shown that chronic viral infection strongly induces differentiation of TFH, which leads to non-specific B cell activation [30]. In addition to TFH cells, researchers have identified TFR cells in the germinal centers. This subset of T cells expresses Foxp3 and also regulates the activity of germinal centers [31]. TFR cells suppress the proliferation of B cells and the production of IgM and IgG antibodies [31, 32] and diminish the secretion of IL-4 and IL-21 by TFH cells in the germinal centers [33]. Measurement of T cell immune responses The standard methods for measurement of T cell immune responses include Enzyme-Linked Immuno Spot assay (ELISpot), Intracellular Cytokine Staining assay (ICS), Tetramer assay and Flow Cytometry. The ELISpot and ICS assays apply in vitro stimulation to analyze the cytokine expression profiles of responding cells. The ELISpot method detects spots of cytokines secreted by individual cells, and ICS examines surface markers and produced cytokines. Multiple approaches can measure the proliferation of T cells in response to specific antigens, including thymidine incorporation assay, flow cytometric analysis of CD38 expression or ELISA detection of BrdU incorporation into DNA of proliferating T cells. T cell immunotherapy T cell immunotherapy has yielded promising results for cancer treatment. Generally speaking, there are two main methods of T cell immunotherapy: 1) application of genetically modified T cell receptors (TCRs) recognizing tumor antigens in relation to HLA and 2) application of chimeric antigen receptors (CARs), which allow binding antigens without HLA recognition [34]. In contrast to CARs, TCRs may recognize both membrane and intracellular antigens. The α and β chains of TCRs recognize T cell targets, and genetic modification of TCR chains modifies antigen specificity. In particular, TCR for MART-1, gp100, and NY ESO-1 have shown anti-cancer activity in patients with melanoma [35]. Other tumors sensitive to TCR modification therapy include lymphoma [36], neuroblastoma [37] and sarcoma [38]. CARs also modify antigen-specific T cell functions and were effective for the treatment of B cell malignancies [39]. The insertion of co-stimulatory signaling regions into the cytoplasmic domain of CARs significantly upregulated the activity of CAR-modified T cells [40, 41]. These co-stimulatory regions include different domains, such as CD28 [42] and OX40 [43], and can modify the T cell cytotoxic activity, proliferation, and survival. The therapeutic method of applying cells expanded ex vivo is named adoptive cell transfer. This treatment uses specific T cells isolated from fragmented tumor tissues. Isolated T cells can be expanded with the help of IL-2, selected and adoptively transferred into patients. Before the adoptive cell transfer, the patients undergo lymphodepletion by either chemotherapy or irradiation [44]. B Lymphocytes Markers for major B cell subtypes There are three main subsets of naïve B lymphocytes: follicular B cells, marginal zone B cells and B1 B cells. Mature follicular B cells migrate through blood and lymph, reside in specific B cell areas of lymph nodes, Peyer’s patches, and the spleen and may present T-dependent antigens to T cells. Marginal zone CD19+CD21+CD23-CD24+IgM+ B cells reside in the marginal sinus of the spleen and mediate the transport of antigen in immune complexes. B1 cells are involved in the development of IgM responses to bacterial T cell-independent antigens. These cells can migrate from the peritoneum and reside in mesenteric lymph nodes. Memory B-cells are represented by three subsets: pre-switch IgD+IgM+CD27+ B cells, IgD-IgM+CD27+ B cells, post-switch IgA+CD27+ and IgG+CD27+ B cells and IgA+CD27- and IgG+CD27- memory B cells [45]. Circulating plasmablasts can be identified by the expression of CD38 and CD138 [46]. Expression of BCR expression is highly important for maintaining B cells in the peripheral immune system. However, only 30% of B cells in spleen develop into mature B cells. Moreover, mice, which have mutations in genes encoding BCR-related proteins, including BLNK, Btk, and Vav, show disruption of the maturation process [47, 48]. B cell maturation markers Lymphoid progenitors Lin-KITlowCSA1lowIL-7R+ are considered to be a lymphoid progenitor group and can differentiate into both B and T cells. Also, in vitro studies have demonstrated that B220-CD19+ cells can differentiate into myeloid or B cells [49] and Lin-KITlowSCA1lowIL-7R+FLT3+CD34- cells or B220-KITlowSCA1+CD24+CD43+ cells contain increased numbers of B cell precursors [50, 51]. Early B220+ precursors of B cells do not express cell surface immunoglobulin (Ig), reside in the bone marrow and include pre-pro-B cells, pro-B cells, and pre-B cells. Immature pre-B cells migrate to the spleen, where they differentiate into mature B cells and plasmocytes (Figure 3). Peripheral B cell subsets, including transitional, mature, memory and antibody-secreting cells, express different surface markers (Table 2). Name Type Phenotype Markers to sub-fractionate Functions Transitional T1 IgD+CD27neg CD10+CD24highCD38highMTG+ Precursor to T2; IL10 production (?) T2 IgD+CD27negCD10+CD24high/+CD38high/+MTG+ Precursor to T3; IL10 production (?) T3 IgD+CD27negCD10negCD24+/lowCD38+/lowMTG+ Precursor to mature-naive; IL10 production (?) Mature-naive IgD+CD27negCD10negCD24+/lowCD38+/lowMTGneg CD23, CD69, CD80, CD86 Precursor to GC, memory, and antibody-secreting cells Memory Double-negative IgDnegCD27neg CD21, CD24, CD95, CXCR3 Recall responses Non-switched IgD+CD27+ CD1c, CD21, CD24 Immunoprotective self antibody, regulatory IgM-only IgM+IgDnegCD27+ CD1c, CD21, CD24 Immunoprotective self antibody, regulatory Switched IgMneg IgDnegCD27+ CD21, CD24, CD95, CXCR3 Pathogen protection; autoimmune pathology Antibody-secreting cell Plasmablast IgDnegCD27highCD38highCD138neg CD20, HLA-DR Antibody secretion Plasma cell IgDnegCD27highCD38highCD138+ CD20, HLA-DR Antibody secretion In addition to IgG production, a subpopulation of splenic B cells can possess regulatory functions. Regulatory B cells (Bregs) affect various parts of the immune system with IL-10 playing a key role in these processes. The B10 subgroup of B cells was shown to act as regulatory cells in experimental models of lupus and autoimmune encephalomyelitis [53]. Moreover, IL-10 producing Bregs with the surface phenotype CD19+CD24hiCD38hi were found in the peripheral blood in SLE patients [54]. In addition, regulatory phenotypes CD19+, CD24+CD27+ and CD19+IgD+CD24hiCD38hiCD5hi were shown to have suppressive functions in humans [55, 56]. Regulatory B cells Several subsets of Bregs were characterized in human peripheral blood. These subsets include B cells with different levels of maturity: transitional CD19+CD24hiCD38hi Bregs [57, 58], CD19+CD27intCD38+, plasmablasts [59] and CD19+CD25+CD71+ B regulatory 1 cells [60]. Recent studies suggest that differentiation and stimulation of Bregs are likely to be induced by inflammation associated with either infection or autoimmune reactions. In particular, toll-like receptor agonists of bacterial origin were shown to activate Bregs in vitro [61, 62]. In addition, the proliferation of Bregs was reported in a murine model of autoimmune arthritis [63]. In addition to the subsets of Bregs mentioned above, Tim-1+ B cells were also shown to regulate immune reactions, since Tim-1 mucin domain-mutated mice develop autoimmune disorders [64]. Tim-1+ Bregs were identified within different B cell subpopulations, including CD19+CD1dhiCD5+, MZ and B1 cells [65]. Also, human CD73−CD25+CD71+ BR1 cells were demonstrated to be involved in the development of allergen tolerance [60]. Membrane regulatory molecules expressed by Bregs include CD25, CD71 and CD274 [54, 66, 67]. Measurement of antibody production One of the most important functions of B cells is antibody production. Enzyme-linked immunosorbent assay (ELISA) can analyze secreted antibodies, plaque-forming cell (PFC) assays can detect antibody-secreting B cells, and ELISPOT can indicate the number of antibody-producing B cells. Antibodies against T and B Cell Markers in the Literature Labome surveys formal publications to develop Validated Antibody Database (VAD). Table 3 lists the most cited antibodies against T cell markers and B cell markers among the 60,000 articles Labome has surveyed as of Jan 2019. Protein Gene ID Num Top three suppliers B220 5788 4117 Invitrogen 14-0452-86 (116), BioLegend 103202 (103), BD Biosciences 560777 (56) c-kit 3815 338 BioLegend 313201 (16), Cell Signaling Technology 3074 (15), Invitrogen MA5-12944 (9) CD1C 911 81 BioLegend 331501 (15), Miltenyi Biotec 130-090-508 (12), Invitrogen AHS0198 (5) CD1D 912 745 BD Biosciences 339186 (79), BioLegend 350302 (3), Santa Cruz Biotechnology sc-19632 (2) CD4 920 2492 Invitrogen MHCD0400 (118), BD Biosciences 555344 (86), BioLegend 317404 (37) CD5 921 147 Invitrogen MA5-13308 (21), Beckman Coulter IM2637U (6), BD Biosciences 644487 (6) CD8 925 2364 Invitrogen MHCD0800 (153), BD Biosciences 339188 (71), Dako M7103 (62) CD10 4311 225 Invitrogen MA5-14050 (56), BD Biosciences 555373 (15), BioLegend 312202 (10) CD11a 3683 74 Invitrogen MA1-19003 (7), BD Biosciences 555381 (4), Abcam ab52895 (3) CD19 930 1362 BioLegend 302202 (53), BD Biosciences 564457 (52), Invitrogen MHCD1921 (39) CD21 1380 89 Invitrogen MA5-11417 (11), BD Biosciences 555421 (9), Dako M0784 (5) CD23 2208 64 Invitrogen MA5-14572 (11), BD Biosciences 550386 (3), Santa Cruz Biotechnology sc-18910 (1) CD24 100133941 311 Invitrogen MA5-11833 (72), BD Biosciences 555428 (26), BioLegend 311102 (5) CD25 3559 778 BD Biosciences 560356 (57), BioLegend 302602 (27), Invitrogen MHCD2506 (17) CD27 939 641 BD Biosciences 561408 (33), Invitrogen 14-0271-82 (26), BioLegend 302839 (25) CD28 940 392 BioLegend 302902 (18), BD Biosciences 556620 (16), Invitrogen 16-0289-85 (13) CD38 952 542 BD Biosciences 646852 (27), Invitrogen MA1-19316 (24), BioLegend 303502 (24) CD44 960 2374 BioLegend 103002 (106), Invitrogen 14-0441-81 (89), BD Biosciences 550392 (31) CD45RB 5788 4117 Invitrogen 14-0452-86 (116), BioLegend 103202 (103), BD Biosciences 560777 (56) CD49d 3676 102 BD Biosciences 555501 (7), BioLegend 304302 (6), R&D Systems BBA37 (6) CD58 965 11 BD Biosciences 555921 (3), Beckman Coulter IM3702 (2), BioLegend 330902 (1) CD62L 6402 306 Invitrogen MA1-10259 (27), BD Biosciences 555542 (22), BioLegend 304802 (15) CD69 969 360 BioLegend 310902 (27), BD Biosciences 560740 (24), Invitrogen MA1-207 (14) CD71 7037 870 Invitrogen 13-6800 (435), BD Biosciences 555534 (13), BioLegend 334102 (4) CD73 4907 173 BD Biosciences 550257 (45), Invitrogen 41-0200 (4), Santa Cruz Biotechnology sc-32299 (3) CD80 941 300 BD Biosciences 557223 (22), BioLegend 305201 (19), Invitrogen MA1-19215 (15) CD86 942 489 Invitrogen MA1-10293 (33), BioLegend 305402 (29), BD Biosciences 555656 (21) CD95 355 334 BD Biosciences 555670 (24), EMD Millipore 05-201 (14), BioLegend 305614 (11) CD99 4267 32 Dako M3601 (16), Invitrogen MA5-12287 (5), BioLegend 915603 (1) CD103 3682 85 Invitrogen 14-1038-82 (7), BioLegend 350202 (7), Beckman Coulter IM1856U (4) CD130 3572 9 BioLegend 362003 (1), BD Biosciences 555757 (1) CD134 7293 27 BD Biosciences 555838 (4), BioLegend 350002 (3), Invitrogen 14-1347-82 (1) CD138 6382 133 Dako M7228 (15), Abcam ab34164 (11), BD Biosciences 650660 (11) CD161 3820 127 BioLegend 339902 (14), BD Biosciences 556079 (9), Miltenyi Biotec 130-092-676 (4) CD127 3575 270 Invitrogen 14-1278-82 (26), BioLegend 351302 (20), BD Biosciences 552853 (16) CD274 29126 338 BioLegend 329701 (16), Invitrogen 14-5983-80 (14), Abcam ab205921 (5) CD357 8784 12 Invitrogen 12-5875-42 (3), BioLegend 311610 (1) CCR4 1233 49 BD Biosciences 551121 (11), BioLegend 359402 (5), R&D Systems MAB1567-100 (4) CCR5 1234 102 BD Biosciences 555991 (12), BioLegend 313712 (4), Invitrogen 12-1957-42 (1) CCR6 1235 154 BD Biosciences 559560 (16), BioLegend 353402 (15), Invitrogen 14-1969-82 (4) CCR7 1236 314 BD Biosciences 552174 (31), BioLegend 353202 (26), Invitrogen 14-1979-82 (13) CCR9 1238 8 R&D Systems MAB1364 (3), BD Biosciences 561607 (2) CCR10 1238 8 R&D Systems MAB1364 (3), BD Biosciences 561607 (2) CLA 6404 25 BD Biosciences 550407 (4), BioLegend 328805 (2), Abcam ab68143 (1) CXCR3 2833 156 BioLegend 353702 (16), BD Biosciences 557183 (8), R&D Systems MAB160-100 (2) CXCR4 7852 275 BioLegend 306502 (16), Invitrogen 35-8800 (15), BD Biosciences 555971 (12) CRTH2 11251 23 BioLegend 350102 (4), BD Biosciences 558412 (4), Beckman Coulter A07413 (1) FoxP3 50943 506 Invitrogen 14-4776-82 (56), Abcam ab20034 (34), BioLegend 320102 (12) Granzyme A 3001 21 BioLegend 507202 (9), BD Biosciences 557449 (1) Granzyme B 3002 320 Invitrogen MA1-80734 (53), BD Biosciences 561151 (28), BioLegend 515406 (21) IL-18Ra 8809 26 BioLegend 313802 (4), R&D Systems MAB840-100 (4), Invitrogen MA1-20257 (1) KLRG-1 10219 27 BioLegend 138429 (6), Santa Cruz Biotechnology sc-32755 L (1), Miltenyi Biotec 130-103-638 (1) MTG 5646 PD-1 5133 509 BioLegend 329902 (42), Invitrogen 14-2799-80 (11), BD Biosciences 562138 (11) Perforin 5551 89 BioLegend 308102 (12), Invitrogen 14-9994-82 (9), BD Biosciences 556434 (8) SCA1 836 1941 Cell Signaling Technology 9664 (362), Novus Biologicals NB100-56708 (41), BD Biosciences 559565 (34) Of Note CD45, also called leukocyte common antigen(LCA), regarded as a pan-immune marker, has also been found in rare epithelial cells in mouse intestine [68], more specifically in tuft-2 cells [69].

|
Suggested by
Société Francaise d'Immunologie
onto Immunology June 11, 2019 1:03 PM
|
No comment yet.
Sign up to comment



 Your new post is loading...
Your new post is loading...