 Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 6:11 PM
|
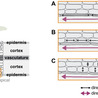
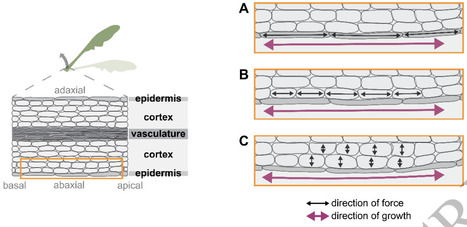
Plants use light both as a resource for photosynthesis and as a signal about their environment. In response to light cues, plants can move their organs via directional growth driven by cell expansion. In dense vegetation where light is available in spatially heterogeneous patterns, plants need to navigate this space to improve the position of their photosynthetic tissues. In canopies blue light irradiance and red to far-red light ratio decrease due to absorption by chloroplasts, and these changes regulate distinct processes within the plant. Changes in light environment are detected by cryptochrome and phytochrome photoreceptors, both regulating Phytochrome Interacting Factors (PIFs) and thereby enhancing elongation in hypocotyls, stems and leaves, and inducing upward leaf movement (hyponasty). An additional class of photoreceptors, phototropins, decode horizontal light gradients to produce directional growth towards the light source (phototropism). Here we review the current state of knowledge on these differential growth responses to light cues, with specific emphasis on the regulatory pathways that translate light signaling in differential cell expansion. Downstream of the photoreceptors, the phytohormone auxin induces cell growth in shoot tissues, but also other phytohormones contribute to balancing light responses. Cell expansion is regulated primarily at the level of cell walls and a comparison of different transcriptome datasets reveals that only a small group of cell wall modifying genes are tightly regulated by shade cues. It remains poorly understood which cell layers are causal to the initiation of cellular expansion. Here we combine insights from different differential growth behaviors in different species and organs to generate different hypotheses for the cellular underpinnings of light-driven leaf movements.

|
Scooped by
mhryu@live.com
Today, 5:04 PM
|
Recent studies offer insights into how the ancient technique works, while newer approaches are bolstering acupuncture with insights from Western medicine.

|
Scooped by
mhryu@live.com
Today, 4:59 PM
|
Spatially regulated membrane constriction is an important milestone in reconstituting minimal cell division. In giant lipid vesicles, bottom-up approaches have reproduced the assembly, mid-cell positioning, and the initial constriction of an FtsZ-based minimal divisome. However, progressive deformation towards giant vesicle scission by near-equatorial Z rings could so far never be observed. One obvious major limitation has been the scale mismatch, as pure reconstituted FtsZ rings typically exhibit bacterial diameters, too small to constrict typical cell-sized vesicles. Therefore, we explore the potential of other key divisome factors to scale up FtsZ-ring functionality in vitro to match the dimensions required for synthetic cell division. We here focus on cytoFtsN, the cytosolic domain of FtsN, and its effect on FtsZ self-organization. Remarkably, a molar excess of cytoFtsN promotes the formation of large, closed equatorial FtsZ rings on giant vesicle membranes, which are able to constrict to almost full closure. By fluorescence imaging and biochemical analysis, we show that cytoFtsN regulates the spatial organization of the FtsZ network primarily by aligning FtsZ filaments while reducing filament depolymerization. Our findings help to define key requirements in a minimal filament-based system for progressive membrane constriction and thus represent a major step forward towards constructing synthetic cells capable of self-division. Spatially regulated membrane constriction is an important milestone in reconstituting minimal cell division. Here the authors engineer a truncated system where the cytosolic domain of FtsN forms large, closed equatorial FtsZ rings on giant vesicle membranes and regulate the FtsZ network formation.

|
Scooped by
mhryu@live.com
Today, 1:30 PM
|
Bio-pesticides constitute a category of natural products and organisms effective against pests. Given their advantages, such as easier clearance and degradation compared to chemical pesticides, along with their lack of detrimental effects on ecosystem health, they represent a promising alternative for sustainable pest management and crop protection. The development of natural biopesticides to substitute synthetic pesticides is therefore of paramount importance for securing food safety and advancing agricultural production. Monoterpenes, primarily present in plant volatile oils, are characterized by a strong aroma and a range of biological activities-including insecticidal, antimicrobial, antiviral, anti-inflammatory, antioxidant, antitumor, and anticancer effects. This profile renders them a promising platform for the research and development of new, more effective pesticides. This article reviews the biosynthetic pathways of monoterpenes and contemporary engineering strategies for their microbial synthesis. Additionally, an evolutionary analysis database for monoterpene synthases (MTPS) has been established. Finally, the biological activities of monoterpenes are summarized and analyzed, with particular emphasis on their potential and advantages as biopesticides, thereby providing direction for the future development of monoterpene-based biopesticides.

|
Scooped by
mhryu@live.com
Today, 1:14 PM
|
The increasing prevalence of antibiotic-resistant bacteria has intensified the demand for novel antimicrobial agents. Antimicrobial peptides (AMP) have emerged as promising alternatives, yet their identification or classification remains challenging due to the lack of multi-perspective information, insufficient feature representation learning, and monocular data modalities. In this paper, we propose a dual diffusion model-based representation learning framework for classifying AMPs, which effectively integrates both peptide sequence and structure information to address existing issues for the task. Specifically, our approach utilizes a multi-view feature construction module, which encodes peptide sequences and structures from distinctive perspectives, deriving initial feature representations with enriched biological semantics. To enhance representation learning, the proposed framework leverages both diffusion models for sequence and structure information respectively to effectively capture complex semantics from dual modalities. In addition, both single-modal and dual-modal contrastive learning are used to further advance the representation learning. Results of comprehensive experiments demonstrate that our model outperforms existing methods for the task of AMPs classification, providing a feasible solution to accelerating the discovery of novel antimicrobial agents.

|
Scooped by
mhryu@live.com
Today, 1:50 AM
|
Engineered living materials (ELMs) promise genetically programmable functions by coupling biological regulation to synthetic material responses. Here, we introduce genetically encoded, reversible shape-morphing in a peptide-crosslinked polyethylene glycol (PEG) hydrogel whose network density is modulated by opposing enzymatic pairs that induce crosslinking or hydrolysis. This molecular programmability alternates the hydrogel between deswelling and swelling/disintegration and produces 2 - 5-fold changes in mechanical properties. By fabricating a bilayer hydrogel with an inert layer, these molecular modulations are translated into a reversible and directional motion with angular bending motions exceeding 80 degrees. Further, by embedding genetically engineered bacteria or interfacing mammalian cells, producing the relevant enzymatic cues, the reversible shape-morphing of these ELMs is programmed at the genetic level. We further demonstrate genetically programmed, autonomous reversible bending in a bilayer hydrogel controlled by out-of-equilibrium counteracting biochemical reactions with dynamically changing respective reaction rates. This work establishes a concept where coordinated polymer/peptide material engineering and synthetic biology yield autonomous shape-morphing ELMs, opening avenues toward biohybrid soft robotics, adaptive microfluidic systems, and dynamic biomedical interfaces.

|
Scooped by
mhryu@live.com
Today, 1:21 AM
|
Cruz-Zaragoza et al. recently introduced a groundbreaking method to silence targeted mitochondrial gene expression in human cells. The approach uses peptide-morpholino chimeras delivered into the mitochondria to block specific mRNA translation. If adapted, this technique can revolutionize our ability to probe plant mitochondrial function and retrograde signaling with unprecedented precision.

|
Scooped by
mhryu@live.com
Today, 12:49 AM
|
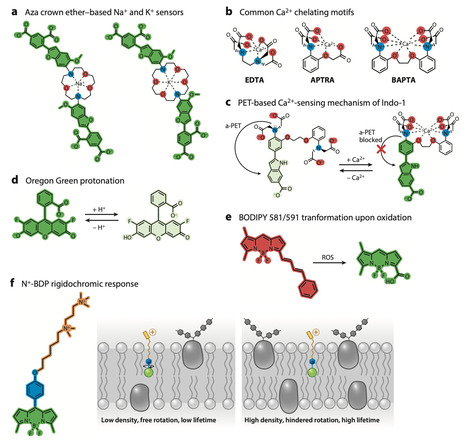
The advent of spatial and quantitative biology has led to immense advances in understanding the complex inner workings of plants, down to the molecular scale. Functional imaging of live plants, which enables the spatial and quantitative mapping of biochemical cues, physicochemical properties of cellular structures, and the dynamics of physical and chemical signals with unprecedented resolution, has become a key technology for advancing the mechanistic understanding of plant cell biology. In this review, we highlight progress in live functional imaging in plants through the use and development of chemical fluorescent probes, which enable plant functional imaging without requiring genetic manipulation of the study object. We explain how probes sense, target, and report on functional features within the plant cell; discuss their limitations, including toxicity; and provide case studies to exemplify how these tools can complement biological studies to unravel the complex machinery that makes plants work. We conclude by outlining the expected future development of this field and identifying key challenges that lie ahead.

|
Scooped by
mhryu@live.com
Today, 12:35 AM
|
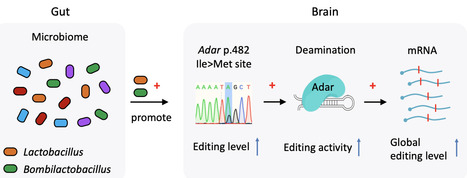
ADAR-mediated adenosine-to-inosine (A-to-I) mRNA editing contributes to the proteomic diversity and behavioral complexity of animals. While recent studies indicate that the gut microbiome influences various aspects of animal behavior, the potential involvement of RNA editing in gut-brain interactions remains unexplored. Comparative transcriptomic analyses were performed between heads of germ-free (GF) versus conventional (CV) honey bees. A total of 1,528 A-to-I editing sites are identified in honey bee heads, among which nonsynonymous editing sites are overrepresented compared to random expectation. Overall editing levels are significantly downregulated in GF compared with CV, but Adar gene is not differentially expressed. However, Adar p.482 Ile>Met auto-recoding site, which is speculated to modulate Adar activity, is identified as a high-confidence differential editing site (DES) with decreased editing level in GF. Quantification of gut microbiota across 12 CV individuals reveals a significant positive correlation between p.482 Ile>Met editing level and Lactobacillus and Bombilactobacillus abundance. Colonization of a single bacterium Lactobacillus or Bombilactobacillus instead of Gilliamella in GF bees successfully restores the Adar p.482 Ile>Met and the global editing level. Our work demonstrates the complex and dynamic transcriptomic diversity exerted by A-to-I RNA editing, and discovers the axis of gut-Lactobacillus/Bombilactobacillus-brain-Adar-global RNA editing, providing an exciting scenario that gut microbiomes could impact RNA editing which might further facilitate phenotypic plasticity of social insects.

|
Scooped by
mhryu@live.com
Today, 12:07 AM
|
Cyclic oligonucleotide-based antiphage signaling systems (CBASS) are immunity pathways in bacteria that use a cGAS/DncV-like nucleotidyltransferase (CD-NTase) enzyme to sense phage infection and initiate antiviral defense. Bacteria encode thousands of diverse CBASS operons, demonstrating a critical role for CD-NTase activation in controlling the prokaryotic response to viral infection. Here we discover proteolytic cleavage by phage prohead proteases as a mechanism of CD-NTase activation and demonstrate that diverse CBASS operons function as molecular sensors of protease activity. We reconstitute CBASS recognition of phage T4 infection in vitro and identify proteolytic cleavage of a surface exposed CD-NTase activation loop as a trigger of enzyme catalysis and nucleotide immune signal synthesis. Phage prohead proteases are sufficient to activate CBASS in vivo and explain how immune signaling is initiated during late stages of viral infection. Combining biochemical and structure-based phylogenetic analyses, we map activation loops in Clade A, D, and G CD-NTase enzymes and define critical residues that control CBASS recognition of distinct phage families. Our results define CBASS recognition of phage protease activity as a widespread mechanism of antiviral defense.

|
Scooped by
mhryu@live.com
Today, 12:00 AM
|
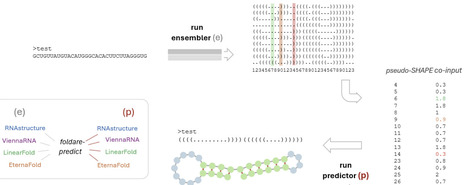
RNA secondary structure prediction remains limited by RNA conformational heterogeneity and scarcity of experimental data: many RNAs populate ensembles of near-isoenergetic folds, and structure-probing data such as SHAPE are often unavailable. Here we introduce FoldARE (Folding and Analysis of RNA Ensembles), a two-step framework that derives pseudo-SHAPE constraints from in silico structural ensembles and uses them to guide a downstream SHAPE-aware predictor. In the first ("ensembler") step, an ensemble is generated and parsed position-by-position to estimate single-strandedness frequencies, which are converted into a pseudo-SHAPE reactivity profile via a weight-and-threshold scheme. In the second ("predictor") step, this profile is supplied as a constraint to a SHAPE-compatible folding algorithm to produce an improved secondary structure. We systematically evaluated all combinations of four ensemble-capable predictors: ViennaRNA, RNAstructure, LinearFold, and EternaFold. We optimized parameters on a manually curated, structurally diverse 25-RNA training set and validated robustness using multiple scoring schemes, including identity-based measures and an exact base-pair-matching metric. The optimal configuration uses EternaFold as the ensembler and RNAstructure as the predictor, yielding consistent gains over all standalone methods and over other ensembler/predictor pairings. On external benchmark RNA structure collections (RNAstrand, ArchiveII, and bpRNA; total n = 1964 after filtering) and on the experimentally derived eFold dataset spanning human mRNAs, pre-miRNAs, and lncRNAs (n = 1024), FoldARE achieved the highest accuracy across datasets with highly significant improvements in paired comparisons. Beyond prediction, FoldARE provides modules for ensemble-level comparative analysis, including pairwise and multi-tool consensus assessment, per-nucleotide variability metrics, and interactive visualizations. It also supports the evaluation of m6A modification effects on folding ensembles using modification-aware engines. Together, our results show that mining ensemble statistics to generate pseudo-probing constraints is an effective, accessible strategy to improve RNA secondary structure prediction and to support ensemble-focused structural analysis. FoldARE is freely available on GitHub (https://github.com/TebaldiLab/FoldARE).

|
Scooped by
mhryu@live.com
March 5, 11:48 PM
|
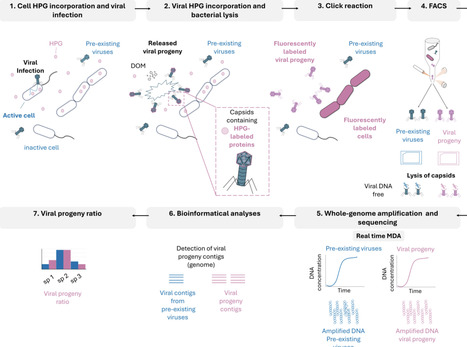
Marine viruses impact biogeochemical cycles through cell lysis, releasing organic matter and nutrients that fuel ocean productivity. Identifying and quantifying the specific viruses active in these processes remains a priority in the field. Here, we introduce a click-chemistry method to fluorescently label, sort, and sequence the genomes of newly produced viral particles (viral progeny) released from transcriptionally active host microbial cells, alongside the analysis of co-occurring inactive cells and pre-existing viruses in environmental samples. This approach, called viral BONCAT-FACS, combines biorthogonal non-canonical amino acid tagging (BONCAT) with environmental sample incubation, followed by single-virus and single-cell sorting by flow cytometry (FACS). Genomic analysis of translationally-active cells and new viral progeny in coastal seawater incubations confirmed BONCAT labelling and successful sorting of diverse marine bacteria, microeukaryotic cells, and virioplankton, with stark differences in the predicted turnover of specific groups of infecting viruses, including Pelagiphages, Methylophages, a Flavobacteriales-associated novel “Far-T4” clade, non-canonical DNA viruses of Naomiviridae using dU instead of dT, algae-infecting giant NCLDV viruses, and parasitic virophages. Sequenced BONCAT-active cells showed a strong enrichment in viral contigs relative to the inactive cell fraction, suggestive of a large proportion of translationally-active virocells. This study illustrates the effectiveness of viral BONCAT-FACS for uncovering genome-resolved virus-host dynamics. By providing a direct approach for tracking active viral infections in natural environments, this method enhances our ability to investigate behavior and interactions of these nanoscale predators, expanding our understanding of their role in ecosystem dynamics.

|
Scooped by
mhryu@live.com
March 5, 11:36 PM
|
Biotechnology holds great potential for sustainable industrial production, with methanol emerging as a promising alternative carbon source due to its availability, cost-effectiveness, and high degree of reduction. Natural methylotrophic microorganisms, such as Komagataella phaffii, are well-suited for methanol-based processes. However, the native xylulose monophosphate (XuMP) cycle in K. phaffii is less energy-efficient than the bacterial ribulose monophosphate (RuMP) cycle, which requires fewer ATP molecules for methanol assimilation. In this study, we introduced the bacterial RuMP cycle as the sole methanol assimilation pathway in K. phaffii. The resulting strain, RuMPi, grew on methanol as its sole carbon and energy source, achieving a specific growth rate of μ = 0.007 ± 0.001 h-1. Optimization and adaptive laboratory evolution (ALE) improved the strain’s performance, resulting in the final strain, RuMPi_mc_fba-ta_evo, with a growth rate of μ = 0.030 ± 0.001 h-1. The final strain exhibited biomass yields and methanol uptake rates comparable to the wild type at lower growth rates. This work demonstrates the feasibility of engineering K. phaffii with a heterologous RuMP cycle and provides insights for optimizing methanol utilization in methylotrophic yeasts for industrial applications.
|

|
Scooped by
mhryu@live.com
Today, 5:53 PM
|
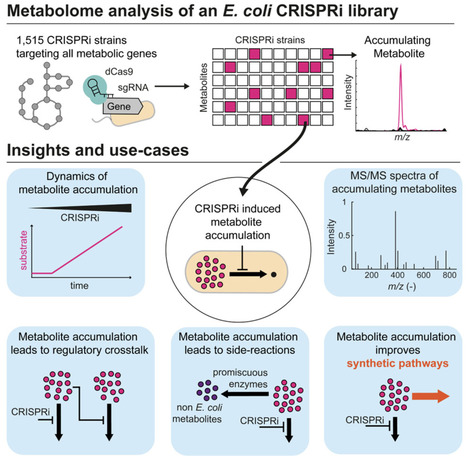
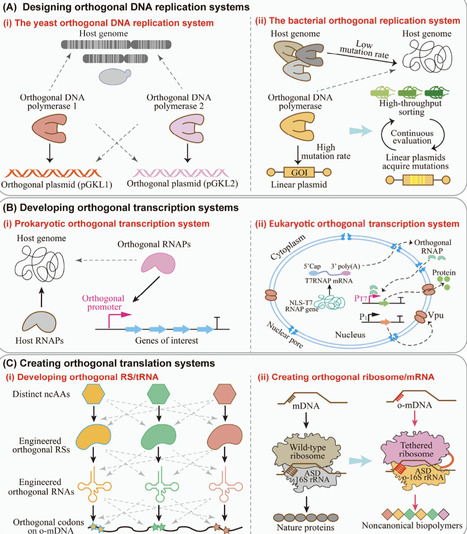
Microbial synthetic biology focuses on the application of rational engineering strategies to reprogram microbial cells, thereby providing them with novel functions to meet different requirements. However, this engineering process inherently disrupts natural metabolism, leading to increased complexity and unpredictability within the metabolic system. To address these challenges, a series of orthogonal strategies has been developed and implemented in the construction of orthogonal genetic systems, metabolic pathways, energy systems, and regulatory systems. This review summarizes recent advances and applications of orthogonal strategies in microbial synthetic biology. Finally, future research directions in orthogonal microbial synthetic biology are explored, aiming to provide new insights for future studies.

|
Scooped by
mhryu@live.com
Today, 5:02 PM
|
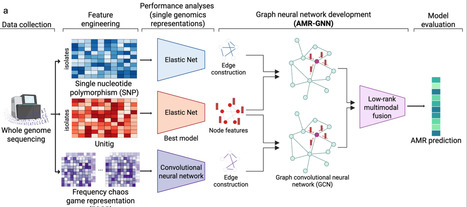
Whole-genome sequencing (WGS) data are an invaluable resource for understanding antimicrobial resistance (AMR) mechanisms. However, WGS data are high-dimensional and the lack of standardized genomic representations is a key barrier to AMR phenotype prediction. To fully explore these high-resolution data, we propose AMR-GNN, a graph deep learning-based framework that integrates multiple genomic representations with graph neural networks (GNN) to enable AMR phenotype prediction from genomic sequence data. We test AMR-GNN with Pseudomonas aeruginosa, a clinically relevant Gram-negative bacterial pathogen known for its complex AMR mechanisms. We present AMR-GNN as a proof-of-concept framework designed to address several key problems in AMR phenotype prediction with data-driven machine learning (ML) approaches, including using multiple genomic representations to enhance performance, to mitigate the influence of clonal relationships and to identify informative biomarkers to provide explainability. Follow-up validation on the largest publicly available dataset spanning both Gram-negative and Gram-positive pathogens highlights AMR-GNN’s broad applicability in detecting AMR in diverse and clinically relevant pathogen-drug combinations. Predicting antimicrobial resistance from bacterial genomic data is challenging. Here, the authors introduce AMR-GNN, a graph neural network that integrates multiple genomic representations to improve prediction, reduce clonal bias, and identify biomarkers.

|
Scooped by
mhryu@live.com
Today, 1:59 PM
|
Evolving dynamic, multi-state, and computational protein functionalities is challenging because it requires selection pressure on all the states of a protein of interest (POI) and the transitions between them. To create a continuous directed evolution paradigm for such properties, we genetically engineered budding yeast for optogenetic input to switch a POI “on” and “off,” which, in turn, controls a Cdk1 cyclin that is essential for one cell-cycle stage but detrimental for another. The method, “optovolution,” generates dynamic selection pressure on POI cycling at the timescale of tens of minutes. We used it to evolve 19 new variants of the LOV transcription factor El222, including in vivo green-light-responsive variants allowing LOV color-multiplexing. Evolving the PhyB-Pif3 optogenetic system, we discovered that loss of YOR1 makes supplementing phycocyanobilin (PCB) unnecessary. Finally, we demonstrated the generality of the method by evolving a non-light-responsive AND gate (PEST-rtTA). Optovolution makes difficult-to-engineer protein functionalities continuously evolvable.

|
Scooped by
mhryu@live.com
Today, 1:24 PM
|
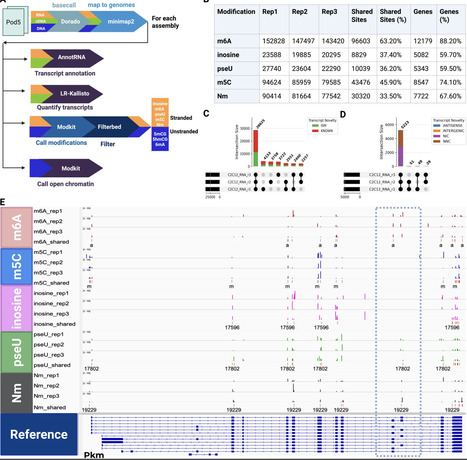
Oxford Nanopore (ONT) sequencing allows for the direct detection of RNA and DNA modifications from unamplified nucleic acids, which is a significant advantage over other platforms. However, the rapid updates to ONT basecalling models and the evolving landscape of computational tools for modification detection bring about challenges for reproducible and standardized analyses. To address these challenges, we developed Dogme to automate basecalling, alignment, modification detection, and transcript quantification. Dogme automates the reprocessing of ONT POD5 files by integrating basecalling using Dorado, read mapping using minimap2 and subsequent analysis steps such as running modkit. The pipeline supports three major types of sequencing data—direct RNA (dRNA), complementary DNA (cDNA), and genomic DNA (gDNA). Dogme facilitates detection of diverse RNA modifications supported by Dorado such as N6-methyladenosine (m6A), 5-methylcytosine (m5C), inosine, pseudouridine, 2’-O-methylation (Nm) and DNA methylation, while concurrently quantifying full-length transcript isoforms LR-Kallisto for transcript quantification for dRNA and cDNA. We applied Dogme to three separate mouse C2C12 myoblast replicates using direct RNA sequencing on MinION flow cells. We detected 96 603 m6A, 43 476 m5C, 8829 inosine, 10 055 pseudouridine, and 30 320 Nm sites in three biological replicates. The pipeline produced reproducible modification profiles and transcript expression levels across replicates, demonstrating its utility for integrative long-read transcriptomic and epigenomic analyses.

|
Scooped by
mhryu@live.com
Today, 2:00 AM
|
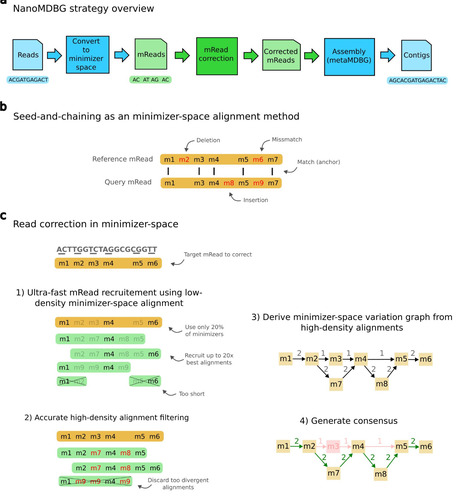
Third-generation long-read sequencing technologies, significantly improve metagenome assemblies. Highly accurate PacBio HiFi reads can yield hundreds of near-complete metagenome-assembled genomes (MAGs) from a single sample. Recently, the accuracy of the more cost-effective Oxford Nanopore Technologies (ONT) platform has increased to a per-base error rate of 1-2%. However, current metagenome assemblers are optimized for HiFi and do not scale to the large data sets that ONT enables. We present nanoMDBG, an evolution of metaMDBG, which supports the latest ONT reads through an error correction pre-processing step in minimizer-space. Across a range of ONT datasets, including a large 400 Gbp soil sample, nanoMDBG reconstructs up to twice as many high-quality MAGs as the next best ONT assembler, metaFlye, while requiring a third of the CPU time and memory. Critically, the latest ONT technology can now produce comparable MAG construction results as those obtained using PacBio HiFi at the same sequencing depth. Benoit et al. present nanoMDBG, a scalable metagenome assembler for the latest ONT long reads, developed by adding error correction to metaMDBG. They demonstrate superior performance to the state-of-the-art and equivalent results to PacBio HiFi reads.

|
Scooped by
mhryu@live.com
Today, 1:29 AM
|
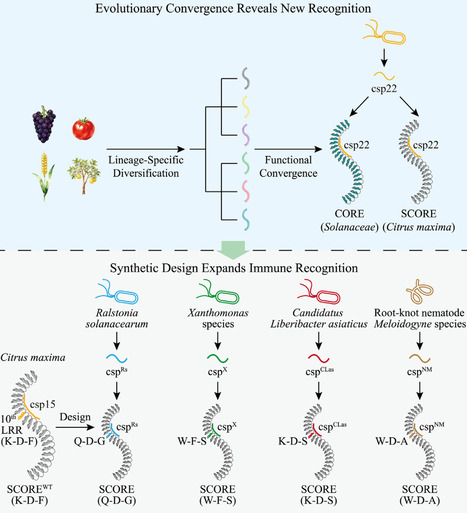
Plants rely on pattern recognition receptors (PRRs) to detect pathogens, yet PRRs differ across species. Ngou et al. show that convergent evolution yields a shared recognition logic in the selective cold shock protein receptor (SCORE) for bacterial cold shock protein peptides. Structure-guided design yields synthetic SCORE variants that broaden detection, offering robust, broad-spectrum crop resistance.

|
Scooped by
mhryu@live.com
Today, 1:15 AM
|
Plant hormones are essential small molecules that regulate plant growth, development, and systemic responses to environmental stimuli. These processes are mediated by complex signaling networks involving structurally diverse receptors, regulatory proteins, and dynamic protein–protein interactions. Advances in structural and functional biology over the past two decades have revealed how hormone receptors recognize their ligands and how they mediate responses from perception to signaling through transduction pathways and feedback regulation. In this review, we summarize the current knowledge of plant hormone receptors with experimentally determined structures and highlight their central roles in shaping plant biology. Finally, we discuss outstanding questions in the field and how emerging computational tools may help address these gaps.

|
Scooped by
mhryu@live.com
Today, 12:44 AM
|
Imbalances in the human gut microbiome, or dysbioses, are associated with multiple diseases but remain poorly understood. Existing biomarkers of dysbiosis fail to capture the ecological mechanisms that differentiate healthy from diseased microbiomes. We have developed a metric, the ecological network balance index (ENBI), that quantifies the balance between positive and negative microbial interactions. This metric was inspired by the phenomenology observed in a model for gut microbiome dynamics that we introduce in this work, which revealed alternative stable states with distinct emergent microbial communities: a healthy state dominated by negative interactions and a dysbiotic state dominated by positive interactions. The ENBI robustly differentiates these states in both simulated and empirical datasets spanning multiple diseases and correlates with disease progression in conditions such as colorectal cancer, which underscores its potential as a diagnostic tool.

|
Scooped by
mhryu@live.com
Today, 12:30 AM
|
The bacterial flagellar motor drives bacterial swimming and chemotaxis by rotating helical flagellar filaments. When E. coli navigates chemical gradients, the motor switches from counterclockwise (CCW) during forward swimming to clockwise (CW) during direction-changing tumbles. The motor responds indirectly to extracellular chemosensory input to membrane-bound chemoreceptors using an intervening intracellular signaling pathway. How the motor responds to its direct input signal—the diffusible messenger phosphorylated CheY (CheY-P)—remains poorly understood. Steady-state motor measurements have been modeled as an allosteric switch between CCW/CW states that depends on mean CheY-P levels. Allosteric models have suggested that as many as 20 CheY-P molecules can be bound to the motor when it switches rotational direction. But steady-state models cannot predict the sensitivity of the motor to dynamic changes in CheY-P that essentially modulate chemotactic behavior. We present an optogenetic reagent that precisely controls the direct dynamical input signal to the motor. We designed a “caged” molecule, Opto-CheY, that is transiently activated by photon absorption. We find that activation and binding of one to three additional CheY-P molecules is sufficient to switch the motor from the CCW to CW state. The sensitivity of the motor to small changes in CheY-P occupancy helps resolve a long-standing paradox about the high sensitivity of the chemotactic response to external sensory input. Optogenetic biochemistry by light-activated uncaging of signal molecules is a new strategy to dissect information-processing in the living cell.

|
Scooped by
mhryu@live.com
Today, 12:04 AM
|
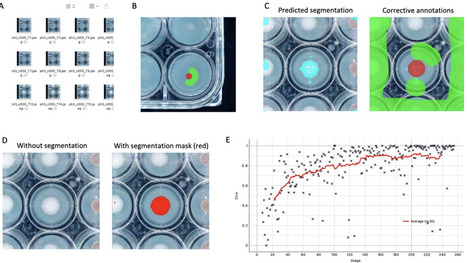
Measuring the growth rate of filamentous fungi is an essential phenotype assay in fungal biology, enabling the comparison of nutrient-related fitness metrics across various isolates, species and genera. Conventional methods are time consuming and labor intensive, which prohibits the adaptation and implementation of high-throughput phenotyping. Here, we suggest a high-throughput methodological pipeline to study fungal growth on solid media combining the use of 24-well plates, an automated image acquisition system, and human assisted deep learning analysis of acquired images. Training a deep learning model through an iterative process - with continuous feedback and corrective annotations - enabled the development of a satisfying model that automatically segments pixels belonging to either fungus or background within a few hours. We evaluated this deep learning model by applying it to two test sets: First, a set of 336 images was used to validate the results by comparison with manual measurements. We demonstrate that the automated segmentation approach provides robust estimation of fungal growth not significantly different to manually segmented data. Second, a larger test set consisting of 2,016 images was used to illustrate the scalability of the model. After training the model for less than two hours, the deep learning model segmented the entire image data set automatically within minutes. The presented method is easily scalable and adjustable to other fungi and growth morphologies, due to the interactive training. Moreover, by combining 24-well plates and automatic image acquisition, measurements can be sped up as growth is detected across a smaller surface area than a standard six or nine cm diameter petri dish. The proposed methodological pipeline thus offers a new tool for estimating fungal growth rates, which can accelerate measurements, reduce bias, and increase throughput.

|
Scooped by
mhryu@live.com
March 5, 11:54 PM
|
In their soluble forms, arsenic and barium are ubiquitous toxic elements. Mechanisms for their detoxification include reducing bioavailability by assimilation into organic forms or mineralization. It was previously found that Entotheonella sp., a bacterium common to the Red Sea sponge Theonella swinhoei (Demospongiae, Tetractinellida), accumulates these elements by mineralizing them intracellularly, thus acting as a detoxifying organ to the sponge host. Here, we utilize cryo-TEM and energy-dispersive spectroscopy (EDS) to investigate the accumulated minerals. Our results show that Entotheonella cells possess an internal membrane-enclosing sphere-like granules that contains barium, arsenic, sulfur, calcium, and phosphorus in high concentrations. Moreover, the bacterial cytoplasm contains many intracellular vesicles (ICVs) enriched with arsenic and sulfur. The coexistence of sulfur and arsenic may suggest the presence of cysteine-containing metal-binding proteins responsible for arsenic uptake and separation within the bacterial cell. To examine that hypothesis, we developed a protocol for vesicle isolation and performed proteomic profiling. Based on the proteins found, ICVs likely originate from the bacteria's outer membrane and contain proteins of known functions, including the transport and detoxification of toxic metals. These findings enhance our understanding of Entotheonella sp. and its host Tamiops swinhoei's unique strategies for hyper-accumulating and neutralizing toxic elements.

|
Scooped by
mhryu@live.com
March 5, 11:39 PM
|
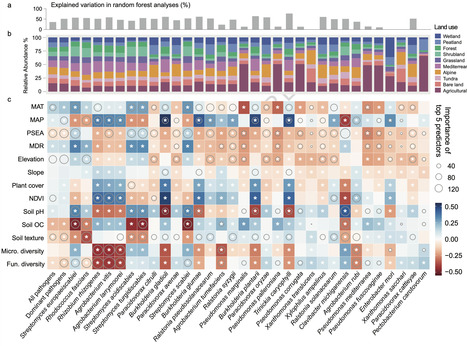
Soils are the primary environmental reservoir of plant pathogens impacting food production and ecosystem productivity worldwide. Yet, some soils can also suppress pathogens through environmental and microbial regulation. Here we integrate 1602 soil metagenomes from 59 countries with a greenhouse experiment to identify 32 dominant pathogens, including Ralstonia solanacearum, Clavibacter michiganensis, and Streptomyces europaeiscabiei. Pathogen hotspots occur primarily in warm ecosystems and agricultural soils, whereas higher soil microbial diversity, increased soil organic carbon and colder climatic conditions are associated with lower pathogen prevalence. Non-pathogenic Streptomyces spp., arbuscular mycorrhizal fungi AMF, and biosynthetic gene clusters BGC encoding terpenes and polyketides are associated with reduced pathogen prevalence. Predictive modelling suggests that several dominant bacterial pathogens are likely to increase in prevalence under future climate scenarios, particularly in tropical and subtropical regions. By identifying global drivers of dominant pathogens and their suppression, this study provides a foundation for improved surveillance and management of plant disease risks under climate change. A global study of 1602 soil samples identifies dominant bacterial plant pathogens and reveals microbial traits, soil carbon and climate that promote natural suppression, while climate change may increase disease risks in many regional hotspots.
|
 Your new post is loading...
Your new post is loading...





















2st,