Get Started for FREE
Sign up with Facebook Sign up with X
I don't have a Facebook or a X account
 Your new post is loading... Your new post is loading...
 Your new post is loading... Your new post is loading...
|

Sridhar Ranganathan's curator insight,
August 7, 2015 11:41 PM
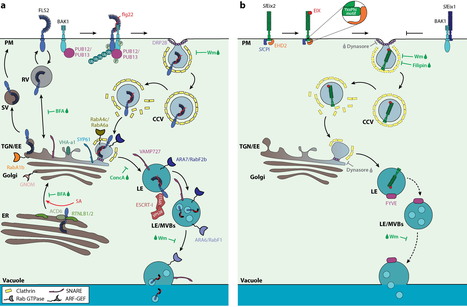
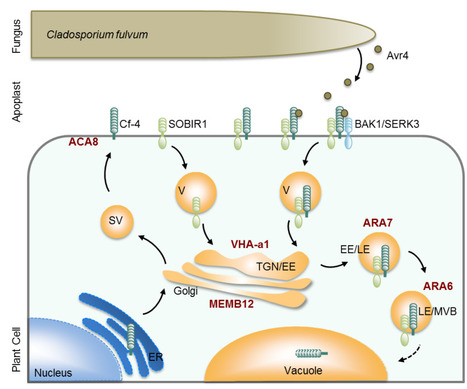
A significant challenge for plants is to induce localized defense responses at sites of pathogen attack. Therefore, host subcellular trafficking processes enable accumulation and exchange of defense compounds, which contributes to the plant on-site defenses in response to pathogen perception. This review summarizes our current understanding of the transport processes that facilitate immunity, the significance of which is highlighted by pathogens reprogramming membrane trafficking through host cell translocated effectors. Prominent immune-related cargos of plant trafficking pathways are the pattern recognition receptors (PRRs), which must be present at the plasma membrane to sense microbes in the apoplast. We focus on the dynamic localization of the FLS2 receptor and discuss the pathways that regulate receptor transport within the cell and their link to FLS2-mediated immunity. One emerging theme is that ligand-induced late endocytic trafficking is conserved across different PRR protein families as well as across different plant species.

Frank Menke's curator insight,
July 23, 2015 8:44 AM
The Pseudomonas syringae effector AvrB targets multiple host proteins during infection, including the plant immune regulator RPM1-INTERACTING PROTEIN4 (RIN4) and RPM1-INDUCED PROTEIN KINASE (RIPK). In the presence of AvrB, RIPK phosphorylates RIN4 at Thr-21, Ser-160, and Thr-166, leading to activation of the immune receptor RPM1. Here, we investigated the role of RIN4 phosphorylation in susceptible Arabidopsis thaliana genotypes. Using circular dichroism spectroscopy, we show that RIN4 is a disordered protein and phosphorylation affects protein flexibility. RIN4 T21D/S160D/ T166D phosphomimetic mutants exhibited enhanced disease susceptibility upon surface inoculation with P. syringae, wider stomatal apertures, and enhanced plasma membrane H+-ATPase activity. The plasma membrane H+-ATPase AHA1 is highly expressed in guard cells, and its activation can induce stomatal opening. The ripk knockout also exhibited a strong defect in pathogen-induced stomatal opening. The basal level of RIN4 Thr-166 phosphorylation decreased in response to immune perception of bacterial flagellin. RIN4 Thr166D lines exhibited reduced flagellin-triggered immune responses. Flagellin perception did not lower RIN4 Thr-166 phosphorylation in the presence of strong ectopic expression of AvrB. Taken together, these results indicate that the AvrB effector targets RIN4 in order to enhance pathogen entry on the leaf surface as well as dampen responses to conserved microbial features. |







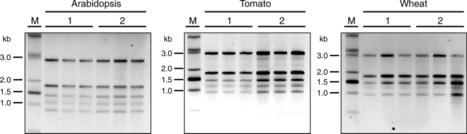
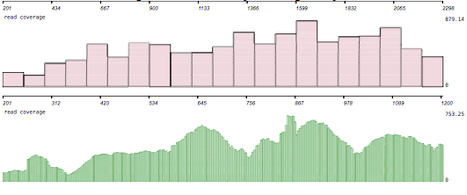



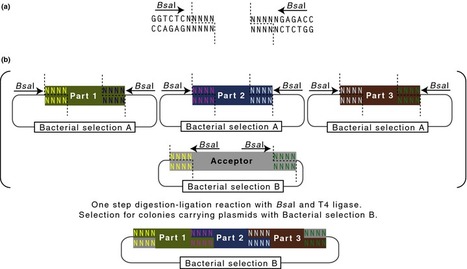








Barley (Hordeum vulgare L.) possesses a large and highly repetitive genome of 5.1 Gb that has hindered the development of a complete sequence. In 2012, the International Barley Sequencing Consortium released a resource integrating whole-genome shotgun sequences with a physical and genetic framework. However, because only 6,278 BACs in the physical map were sequenced, fine structure was limited. To gain access to the gene-containing portion of the barley genome at high resolution, we identified and sequenced 15,622 BACs representing the minimal tiling path of 72,052 physical-mapped gene-bearing BACs. This generated ~1.7 Gb of genomic sequence containing an estimated 2/3 of all Morex barley genes. Exploration of these sequenced BACs revealed that although distal ends of chromosomes contain most of the gene-enriched BACs and are characterized by high recombination rates, there are also gene-dense regions with suppressed recombination. We made use of published map-anchored sequence data from Aegilops tauschii to develop a synteny viewer between barley and the ancestor of the wheat D genome. Except for some notable inversions, there is a high level of collinearity between the two species. The software HarvEST:Barley provides facile access to BAC sequences and their annotations, along with the barley-Ae. tauschii synteny viewer. These BAC sequences constitute a resource to improve the efficiency of marker development, map-based cloning, and comparative genomics in barley and related crops. Additional knowledge about regions of the barley genome that are gene-dense but low-recombination is particularly relevant.