 Your new post is loading...

|
Scooped by
mhryu@live.com
March 29, 5:51 PM
|
Motility often plays a decisive role in the survival of species. Five systems of motility have been studied in depth: those propelled by bacterial flagella, eukaryotic actin polymerization and the eukaryotic motor proteins myosin, kinesin and dynein. However, many organisms exhibit surprisingly diverse motilities, and advances in genomics, molecular biology and imaging have showed that those motilities have inherently independent mechanisms. This makes defining the breadth of motility nontrivial, because novel motilities may be driven by unknown mechanisms. Here, we classify the known motilities based on the unique classes of movement-producing protein architectures. Based on this criterion, the current total of independent motility systems stands at 18 types. In this perspective, we discuss these modes of motility relative to the latest phylogenetic Tree of Life and propose a history of motility. During the ~4 billion years since the emergence of life, motility arose in Bacteria with flagella and pili, and in Archaea with archaella. Newer modes of motility became possible in Eukarya with changes to the cell envelope. Presence or absence of a peptidoglycan layer, the acquisition of robust membrane dynamics, the enlargement of cells and environmental opportunities likely provided the context for the (co)evolution of novel types of motility.

|
Scooped by
mhryu@live.com
March 29, 5:44 PM
|
Precise control of gene expression in a cell-state-specific manner is essential for effective therapeutic interventions in complex and dynamic disease microenvironments. Traditional targeting strategies that rely on surface markers or cell type-specific promoters often assume static cellular identities, limiting effectiveness in context such as cancer and inflammation, where cell states are highly heterogeneous and dynamic. RNA sensors, such as RADAR (RNA sensing using Adenosine Deaminases Acting on RNA), provide a modular, programmable, and nonintegrating platform for classifying cell states. However, it is also characterized by low sensitivity and dynamic range, which limits its applications in detecting low-abundance transcripts. In this work, we integrate RADAR sensors with a signal amplification circuit to enhance sensitivity and dynamic range. We demonstrate that this combined RADAR-amplifier platform enables real-time monitoring of subtle changes in the abundance of endogenous transcripts under physiological conditions. Our results demonstrate the utility of this platform for fundamental biological studies and the development of precision therapeutic strategies.

|
Scooped by
mhryu@live.com
March 29, 5:32 PM
|
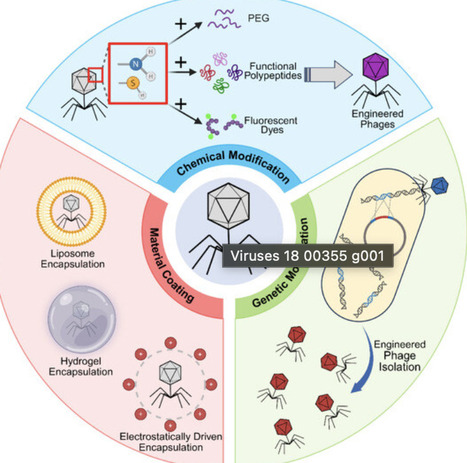
The escalating crisis of antimicrobial resistance (AMR) and the stagnating antibiotic pipeline have renewed interest in bacteriophage therapy. While natural phages offer specificity and low toxicity, their narrow host range, bacterial resistance, and safety concerns limit clinical use. To overcome these hurdles, phages are being engineered using biotechnology. This review outlines the history of phage therapy and systematically summarizes advances in engineered phage preparation, including genetic modification, chemical conjugation, and physical encapsulation. We highlight the application of engineered phages against multidrug-resistant infections, gastrointestinal diseases through gut microbiome modulation, and as targeted delivery vehicles or immune adjuvants in cancer therapy. While significant advances have been made, several critical challenges remain, particularly in regulatory approval, large-scale manufacturing, and ensuring long-term safety. We conclude that engineered phages, as customizable and precise biological tools, are poised to advance precision phage medicine, offering a transformative solution to AMR and fostering convergence across synthetic biology, medicine, and environmental science.

|
Scooped by
mhryu@live.com
March 29, 3:53 PM
|
T7 RNA polymerase is a foundational enzyme for biotechnology, but its utility for many potential applications is limited by low thermal stability of 43-44°C. While stabilized variants exist, the most stable commercial version has a proprietary sequence. In this work we developed a highly stable T7 RNAP using structure-based computational design. We combined mutations from previous stabilized variants (M5, M8, V7abcd) with new mutations identified by PROSS. These mutations were filtered using data-driven heuristics to preserve function. Our final design, T7T+, contains 30 point mutations from the original T7 RNAP and demonstrates a functional stability (T50) of 54.9°C in a thermal challenge assay, which is 2.4°C higher than the most stable, published open-source variant to date. Circular dichroism spectroscopy showed an apparent melting temperature of 53.8°C. T7T+ retains 59% of wild-type activity at 37°C. 16 of the 18 tested protein designs had higher stability against thermal challenge compared with the genetic background, attesting to the high success rates of existing non deep learning computational methods for the design of stable, functional proteins. A plasmid encoding T7T+ has been deposited in AddGene and is freely available for non-commercial use.

|
Scooped by
mhryu@live.com
March 29, 3:43 PM
|
Diatoms exhibit high competitive capacity in nitrogen assimilation, but the underlying mechanisms remain unclear. Here, we identify a non-ribosomal peptide synthase-like gene (PtNRPS1) with an atypical domain structure (A-T-R1-R2) in the marine diatom Phaeodactylum tricornutum, crucial for short-term nitrogen assimilation. In vitro enzyme assays show PtNRPS1 catalyzes conversion of L-tryptophan to tryptophanol, a tryptophan-derived indole compound that promotes diatom growth at concentrations far lower than indole-3-acetic acid. Transcriptomic, metabolomic analyses, and stable-isotope analyses indicate tryptophanol enhances short-term nitrogen assimilation. CRISPR-Cas9 knockout of PtNRPS1 abolishes tryptophanol biosynthesis and reduces nitrogen-assimilation enzymes activities, which are restored by exogenous tryptophanol. PtNRPS1 overexpression results in delayed but sustained enzyme elevation. Global distribution of PtNRPS1 homologues in stramenopiles positively correlates with nitrogen-assimilation gene abundance. Our findings suggest tryptophanol, synthesized by a diatom NRPS, accelerates nitrogen assimilation, providing a competitive edge in oceanic nitrogen acquisition. . Here the authors show that marine diatoms produce tryptophanol, a molecule that at extremely low concentrations rapidly activates nitrogen assimilation genes and enzymes. This facilitates acquisition of nitrogen, a vital ocean nutrient.

|
Scooped by
mhryu@live.com
March 29, 3:32 PM
|
In all kingdoms of life, the regulation of membrane-bound enzyme function via oligomerization is a fundamental aspect of cell physiology. Often, the mechanistic role of oligomerization is unclear, due to a lack of structure-function comparisons between constituent forms of the enzyme. Here, we elucidate the structural underpinnings of enzyme regulation and oligomerization in the quinol-dependent nitric oxide reductase (qNOR) from Neisseria meningitidis, by high-resolution structural analyses of the less active monomeric form (2.25 Å) and the highly active dimeric form (1.89 Å). The comparison revealed that broad helical flexibility near the dimer interface of the monomer causes a conformational change in a critical amino acid near the active site, located apart from the dimer interface. We demonstrate that the crosstalk between the dimer interface and catalytic site in qNOR allows enhanced activation of the enzyme via dimerization. Given Neisseria meningitidis’ dependence on qNOR to detoxify the host’s immune response of nitric oxide, our results pave a way for new strategies to combat bacterial infections, via the inactivation of qNOR by monomerization. More broadly, this provides new insights into the role of membrane protein oligomerization and its influence on regulating activity. Dimer-monomer structural and functional comparison of quinol-dependent nitric oxide reductase from Neisseria meningitidis reveals insights into the interlinking role of dimer stability and high catalytic activity.

|
Scooped by
mhryu@live.com
March 29, 3:14 PM
|
CRISPR-based epigenome editing represents a programmable strategy to precisely modulate gene expression, holding immense promise for therapeutic applications. However, the large size of the dCas proteins substantially impedes the delivery via adeno-associated virus (AAV) vectors. Here, through iterative bioinformatics analysis, structure-guided predictions, and functional assays, we identified and characterized a novel miniature subtype V-M CRISPR-Cas12m from Pelomicrobium methylotrophicum. PmCas12m exhibited flexible 5'-YTN-3' PAM-dependent recognition and robust double-stranded DNA binding properties, while lacking DNA cleavage activity, thus positioning it as an ideal tool for epigenome editing. Cryogenic electron microscopy (cryo-EM) structures of PmCas12m unveiled its unique molecular mechanism of DNA binding facilitating interference. Guided by these structural insights, we employed deep mutational scanning (DMS) and protein engineering to develop xCas12m, a hypercompact variant with highly potent and specific epigenome editing capabilities in human cells. We further constructed the xCas12m-CRISPRoff platform in a single AAV vector, which achieved durable epigenetic silencing and effective inhibition of hepatitis B virus (HBV) infection in a mouse model. Collectively, these findings establish xCas12m as a versatile epigenome editing platform with transformative potential for treating diseases, paving the way for clinical translation of epigenetic therapies.

|
Scooped by
mhryu@live.com
March 29, 2:45 PM
|
Cabbage black rot, caused by Xanthomonas campestris pv campestris (Xcc), is a severe disease worldwide. The predominant method for controlling this disease relies on agrichemicals, which increase antibiotic resistance. This study showed that the secondary metabolites of B. cereus BR3 elicited induced systemic resistance in cabbage and impaired the type III secretion system (T3SS) of Xcc. Furthermore, the extract of strain BR3 significantly enhanced the activities of plant-defense-related enzymes, suppressed the hypersensitive response (HR) induced by Xcc in tobacco, and attenuated the pathogenicity of Xcc on cabbage. Additionally, several genes of Xcc T3SS were repressed by the BR3 extract. Activity-guided purification identified 3-methylcinnamic acid as an effective inhibitor against the T3SS of Xcc. Overall, this study suggests that 3-methylcinnamic acid produced by strain BR3 disrupts a critical virulence mechanism in Xcc, providing a promising alternative for controlling the black rot disease in cabbage.

|
Scooped by
mhryu@live.com
March 29, 2:36 PM
|
The first and arguably most critical level of gene expression is transcription of the genetic information from DNA into RNA. Within this process, transcription initiation stands out as the key step that influences all downstream events. Central to initiation are promoters, DNA sequences that interact with the key enzyme of transcription, RNA polymerase. A single transcription unit may be controlled by one or multiple promoters, a strategy found across all domains of life. This review highlights how combinatorial promoter arrangements control gene expression in microorganisms, with brief comparisons to other organisms. We also explore how these promoter architectures function as sensors for stress-related molecules, such as antibiotics. We examine how these insights can be applied to predict previously unidentified mechanisms of antibiotic resistance. Finally, the use of dual-promoters in synthetic biology is outlined and discussed.

|
Scooped by
mhryu@live.com
March 29, 2:30 PM
|
Recent research has shown that the biosphere harbors an immense diversity of large and complex DNA viruses that infect a wide range of hosts. This is best exemplified by giant viruses of eukaryotes (phylum Nucleocytoviricota) and large phages, also referred to as jumbo phages, that infect bacteria (class Caudoviricetes). Both groups independently evolved large genomes and complex infection strategies, and recent research reveals striking convergences between these viral groups despite their distinct evolutionary origins. Virus factories and phage nuclei separate transcription from translation, mediate spatiotemporal regulation of infection, and involve elaborate interactions with host cytoskeletal systems. Moreover, both viral groups harbor an expanded repertoire of genes acquired from cellular hosts, including transcription and translation-related genes and metabolic enzymes, suggesting convergent strategies to optimize host takeover. Additionally, both lineages are embedded in networks of hyperparasitic interactions, including viral parasites and phage satellites, as well as other selfish genetic elements such as transposons and homing endonucleases, which may, in some cases, facilitate genome innovation and provide fitness benefits. The repeated emergence of genome gigantism across diverse viral clades within both the Nucleocytoviricota and Caudoviricetes suggests that selective pressures favoring gene gain, potentially through a genomic accordion mechanism, are widespread. Yet, genome expansions also harbor many features that are lineage-specific and highly context-dependent. This review explores the mechanistic, functional, and evolutionary parallels that shape the biology of genome gigantism in the virosphere.

|
Scooped by
mhryu@live.com
March 29, 1:59 PM
|
Effective collaboration between biotech start-ups and universities is critical for translating academic discoveries. This article examines intellectual property ownership models, licencing strategies, and publication timing, offering practical, experience-derived guidance for founders and technology transfer offices to strengthen commercialisation outcomes.

|
Scooped by
mhryu@live.com
March 29, 1:18 PM
|
Plants monitor their environment for microbial invaders using pattern-recognition receptors that detect microbe-associated molecular patterns (MAMPs). Flagellin, the main component of bacterial flagellum, contains the flg22 epitope recognized by the plant immune receptor FLS2. Immune recognition can create an evolutionary conflict, requiring bacteria to balance flagellar function and immune evasion. Here, we show that plant-associated Sphingomonas resolve this constraint by partitioning two flagellar functions, motility and colonization, across two divergent and independently acquired flagellin genes. Comparative genomics revealed widespread coexistence of FliC proteins expressing either an immunogenic variant (FliC-H) or a nonimmunogenic variant (FliC-L). The nonimmunogenic FliC-L is necessary and sufficient for full directional swimming, whereas FliC-H is dispensable for swimming, but sufficient for full attachment and colonization. Flagellin expression patterns mirror these functions. Thus, FLS2 recognizes the flagellar variant required for colonization rather than motility, potentially restricting colonizing bacteria from entering internal leaf and root tissues.

|
Scooped by
mhryu@live.com
March 29, 1:00 PM
|
Metabolic engineering calls for new bacterial chassis that combine robust metabolic and physiological traits with genetic tractability. We developed Pseudomonas vancouverensis DhA-51, a native one-carbon (C1)-trophic strain, into a platform that broadens chassis options beyond the usual Pseudomonas hosts while retaining native capacities relevant to bioproduction. We closed and annotated the chromosome to provide stable coordinates for genome editing and omics, established antibiotic susceptibility profiles, and explored replication of common broad-host-range plasmid toolsets. An expression toolkit, including three constitutive promoters and two chemically-inducible systems, was parameterized with fluorescent reporters. Clean markerless genome editing by homologous recombination was demonstrated through deletion of the benABCD cluster that mediates aromatic compound breakdown. Physiological tests showed growth on diverse carbon substrates and synthesis of medium-chain-length polyhydroxyalkanoates from glucose, octanoate, and C1 substrates, with polymer content and monomer composition tunable by the carbon-to-nitrogen ratio and feedstock type. Isotope tracing showed that methanol can be assimilated while formate is used as a reductant. The domestication sequence applied here shortens design-build-test-learn cycles and can be applied across the Pseudomonas genus to expand the portfolio of workable chassis for sustainable production of chemicals and materials from renewable feedstocks.
|

|
Scooped by
mhryu@live.com
March 29, 5:48 PM
|
Actinobacteria represent a prolific source of bioactive natural products. However, the complex transcriptional regulatory networks in these bacteria, particularly the interplay between transcription factors (TFs) and their regulatory ligands (TF-RLs), remain poorly characterized and lack dedicated resources. In this context, we introduce the Actinobacteria Transcription Factor Database (Actinobacteria TFDB), a comprehensive repository that systematically integrates TF-centric data across 25 representative species. The current version encompasses 629 TFs, classified into 69 families, documents 11,776 TF-target relationships and 28 TF posttranslational modification sites. Uniquely, it features a dedicated collection of 54 experimentally validated TF-RL interactions. Beyond providing standardized annotations, sequence and structural features, and regulatory networks, Actinobacteria TFDB incorporates a specialized TF-RL module that enables interactive exploration and visualization of allosteric regulatory mechanisms. By consolidating multi-dimensional TF data from diverse sources, this resource empowers systems-level analyses and facilitates the rational design of regulatory strategies to activate silent biosynthetic gene clusters and optimize metabolite production. The database is publicly available at http://mingleadgene.com:9315/#/home.

|
Scooped by
mhryu@live.com
March 29, 5:38 PM
|
In vivo multiple-enzyme cascades have attracted considerable interest for their ability to provide a native microenvironment that supports enzymatic activity and membrane protein function. This review outlined four pivotal strategies for their optimization, increasingly empowered by Artificial Intelligence (AI): (1) enhancing enzyme performance by enzyme discovery and engineering; (2) precisely modulating enzyme expression via rationally designed genetic regulatory elements; (3) implementing spatial and stoichiometric control using protein, nucleic acid, or synthetic scaffolds and compartments; and (4) employing multimodule systems including multiple cell modules and hybrid in vivo/in vitro cascades. Advances in AI accelerate these strategies, enabling novel approaches such as de novo protein design, directed evolution, and the computational design of genetic parts and supramolecular scaffolds. The integrated implementation of these methods substantially increased target compound titers. This lays a strong foundation for industrial implementation. However, several key challenges remain to be addressed.

|
Scooped by
mhryu@live.com
March 29, 5:24 PM
|
Exosomes, despite their promise as drug carriers for crossing biological barriers, remain underexplored for noninvasive posterior ocular delivery. Here, we demonstrate that semen-derived exosomes (SEVs) penetrate ocular barriers effectively, owing to their epidermal growth factor expression, which mediates reversible tight-junction disruption. SEVs reach the posterior segment via dual corneal and conjunctival routes. Using this, we engineered FA-SEVs@CMG eye drops, where SEVs are modified with folic acid (FA) and loaded with a nanozyme system (CMG) composed of carbon dots, manganese dioxide, and glucose oxidase. This eye drop leverages SEVs’ excellent penetration ability and FA’s targeting effect to enhance drug delivery to retinoblastoma (RB) cells. Internalized CMG induces intense oxidative stress, disrupts the autophagy-apoptosis balance, and triggers RB cell self-destruction. In vivo, FA-SEVs@CMG effectively inhibits RB growth while preserving retinal function. This work establishes the first SEV-based platform for noninvasive posterior segment delivery, offering a transformative strategy for treating posterior ocular diseases.

|
Scooped by
mhryu@live.com
March 29, 3:45 PM
|
Formic acid (FA), a key one-carbon liquid compound derived directly from CO2, can serve as a dual-purpose substrate in microbial metabolism, supplying both carbon and energy. Its potential for green biomanufacturing is immense, yet its inherent toxicity and poor metabolizability to most microbes pose a major hurdle in developing efficient microbial cell factories for value-added chemical production. Building on our prior discovery of Vibrio natriegens as a naturally proficient formic acid utilizer, we demonstrate here that formate supplementation as an auxiliary substrate can dramatically boost pyruvate production of V. natriegens from sodium gluconate, achieving a 1.9-fold increase in titer. Transcriptomic analysis revealed that formate presence induces global changes in gene expression. By subsequently downregulating the pyruvate consumption pathway, we engineered a strain that, when co-fed with formate and sodium gluconate, achieved a 49.0% improvement in pyruvate synthesis. Isotopic tracer analysis confirmed substantial formate assimilation, with approximately 9.43% incorporated into biomass. In a fed-batch fermentation, the engineered V. natriegens strain consumed 82.8 g/L sodium gluconate and 37.4 g/L formate (HCOONa·2H2O) within 51 h, producing 56.4 g/L pyruvate at a rate of 1.1 g/L/h. This work elucidates the stimulatory role of formate in the pyruvate biosynthesis of V. natriegens and establishes a novel strategy for leveraging this feedstock in microbial production.

|
Scooped by
mhryu@live.com
March 29, 3:38 PM
|
Bacteriophages are key drivers of microbial ecology, co-existing and co-evolving with bacteria across diverse environments. Limitations in culturing, alongside advances in sequencing and bioinformatics, have driven the use of metagenomics to explore viral diversity. Viral-specific analysis of >3000 food metagenomes from cFMD produced the FVGC, comprising ~3400 metagenome-assembled viruses, most of which belong to novel Caudoviricetes lineages (n = 91), with only ~15% represented in IMG/VR v4. Together, these findings reveal extensive uncharacterized viral diversity in food systems. Beyond serving as a reference, the FVGC facilitates detailed investigation of virus–host interactions. Viral sequences were pervasive across microbial genomes, with several bacterial families exhibiting near-universal associations with viral elements. Bacterial antiviral defence systems were abundant and taxonomically diverse, dominated by restriction–modification systems, while CRISPR–Cas systems showed pronounced lineage-specific distributions; in contrast, viral anti-defence genes were detected at low frequency (<10% of MAVs). Host prediction linked MAVs to clinically relevant taxa, including expanded ESKAPE pathogens such as Klebsiella pneumoniae, Acinetobacter baumannii, Staphylococcus aureus, and Enterobacter spp., highlighting the ecological connectivity between food-associated viruses and clinically important bacteria. Antimicrobial resistance signals were scarce, suggesting minimal phage-mediated AMR dissemination in food environments. This new publicly available viral database represents a valuable resource for further exploration of viral diversity.

|
Scooped by
mhryu@live.com
March 29, 3:21 PM
|
Microplastics (MPs) in wastewater treatment plants (WWTPs) represent novel ecological niches rather than passive contaminants. They rapidly acquire distinct microbial biofilms, forming the “Plastisphere”. This Perspective examines the underexplored role of quorum sensing (QS) within these complex synthetic microenvironments. We highlight how the intrinsic properties of MPs, such as surface hydrophobicity, chemical leachates, and aging states, create altered signaling landscapes. These distinct environments may decouple QS responses from classical population-density thresholds. Consequently, these dynamic interactions potentially enhance cooperative microbial metabolism and may indirectly facilitate the horizontal gene transfer of antibiotic resistance genes by fostering dense cellular proximity. Despite substantial methodological challenges in detecting active signaling in situ, MPs must be reconceptualized as quorum-modulating microscopes capable of reprogramming microbial communication. Advancing this field requires integrating high-resolution spatial imaging and functional genomics under operationally relevant conditions to bridge theoretical insights with empirical validation, ultimately informing future treatment designs and environmental risk assessments. Microplastics intensify quorum-sensing-mediated gene exchange in wastewater treatment plants, enabling their reconceptualization as quorum-modulating microscopes that reprogram microbial communication and function, based on their interplay with biofilms and quorum sensing within wastewater systems

|
Scooped by
mhryu@live.com
March 29, 3:10 PM
|
Direct Coupling Analysis has been instrumental over the past decade in leveraging evolutionary information and advancing our understanding of biomolecular structure and function. Here, we introduce sparse extensions of this method that explicitly incorporate structural information. StructureDCA focuses on physically relevant interactions by selectively retaining couplings between residues in spatial contact, and StructureDCA[RSA] additionally incorporates per-residue relative solvent accessibility. These models outperform state-of-the-art approaches in describing mutational landscapes, as they more effectively integrate structural context. Moreover, their sparse formulation enables orders-of-magnitude improvements in computational efficiency while preserving interpretability, providing a powerful framework for gaining mechanistic insights into mutation effects and advancing protein design. The StructureDCA models are available as a user-friendly Python package via the PyPI repository. The source code is freely accessible at https://github.com/3BioCompBio/StructureDCA, which also includes a Colab Notebook interface.

|
Scooped by
mhryu@live.com
March 29, 2:43 PM
|
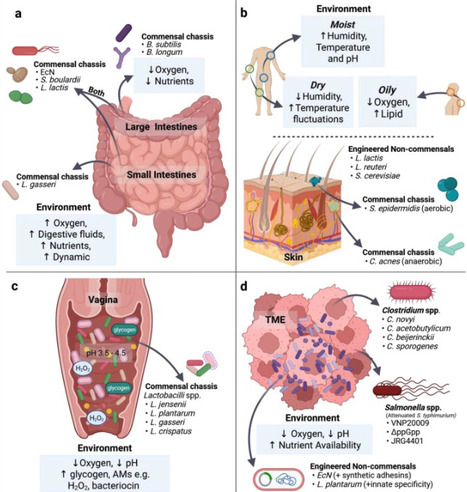
The human microbiota comprises a vast and diverse array of microorganisms that play critical roles in maintaining health and modulating diseases. Engineered live biotherapeutic products (eLBPs) harness genetically modified microbes to perform defined therapeutic functions within the host. A central challenge in developing effective eLBPs is the rational selection of an appropriate microbial chassis, which requires consideration of safety, genetic tractability, functional performance and the target host niche. As different body sites present distinct physiological and biochemical conditions that influence microbial survival, colonisation and activity, choosing a chassis well-adapted to the target niche is essential for therapeutic durability and sustained efficacy. This review summarises microbial chassis that have been widely employed for the gut, skin, vagina and tumour microenvironment, highlighting how their characteristics enable effective function within these niches. Strategies to improve eLBP colonisation within these niches are discussed, and emerging microbial candidates that hold promise as future eLBPs are also identified.

|
Scooped by
mhryu@live.com
March 29, 2:33 PM
|
Extracellular vesicles (EVs) are emerging as versatile therapeutic platforms, yet the mechanisms governing their biogenesis in yeast remain incompletely understood. Saccharomyces cerevisiae, a well-characterized and safe microbial chassis, naturally secretes abundant EVs and provides an attractive system for mechanistic dissection and engineering. Here, we establish S. cerevisiae as a tractable model for elucidating EV cargo loading. By combining multicopy expression of chicken interferon-λ (ChiIFN-λ) with cell wall perturbation, we achieved a tenfold increase in EV yield and efficient incorporation of ChiIFN-λ into EVs. Quantitative proteomics identified 1555 EV-associated proteins, including 501 predicted transmembrane proteins derived from multiple organelles. ChiIFN-λ overexpression and cell wall stress selectively reduced the abundance of key vesicle trafficking regulators, including SNARE, ESCRT and Rab proteins, indicating reprogramming of intracellular membrane trafficking pathways. Functional analyses further demonstrated that the SNARE proteins Sso2 and Nyv1 are enriched in the EV membrane and modulate EV size distribution and subpopulation composition. Together, these results reveal conserved protein-sorting machinery underlying yeast-derived extracellular vesicles (YDEVs) biogenesis and establish S. cerevisiae as a powerful platform for engineered EV production.

|
Scooped by
mhryu@live.com
March 29, 2:27 PM
|
Laboratory mice housed under specific pathogen-free conditions are the standard model in biomedical research. However, frequent germ-free rederivation and barrier housing restrict microbial exposure and interactions with commensal and pathogenic microorganisms. In contrast, wild mice encounter diverse microbial environments, undergo natural selection, and rely on robust immune responses for survival. Consequently, lifelong microbial exposure is a key driver in shaping mammalian physiology, establishing the need for naturalizing rodent models in biomedical research. By defining the concepts of ‘microbial self’ and ‘microbial nonself’, we propose a four-step guide for establishing a multigenerational wildling colony that accounts for both microbial self and microbial nonself. This reestablishes the common biological link shared by all free-living mammals, thereby improving the comparability between murine and human studies.

|
Scooped by
mhryu@live.com
March 29, 1:58 PM
|
One-third of food produced globally is lost or wasted, resulting in significant greenhouse gas emissions and economic losses. New strategies are needed to minimize food and agricultural loss or waste (FALW) and mitigate these negative planetary impacts. Filamentous fungi—a diverse group of microorganisms including molds and mushrooms—offer a unique solution to the FALW problem. These organisms are nature’s recyclers, capable of breaking down complex organic biomass, including food matter. Additionally, many fungi are edible and have long been used in food fermentation, suggesting they could be used to convert waste into food. In this review, we discuss emerging strategies across genetics, bioprocessing, and gastronomy that enable the production of sustainable foods from readily available FALW.

|
Scooped by
mhryu@live.com
March 29, 1:13 PM
|
Mammalian cells initiate antiviral signaling when cyclic GMP-AMP synthase (cGAS) detects cytoplasmic DNA and synthesizes 2′,3′-cyclic GMP-AMP (2′,3′-cGAMP), which activates stimulator of interferon genes (STING). Similarly, bacteria use cyclic oligonucleotide-based antiphage signaling systems (CBASS) to detect phage using ancestral cGAS/DncV-like nucleotidyltransferases (CD-NTases), but they are not known to use 2′,3′-cGAMP. Here, we discover a bacterial CD-NTase that produces 2′,3′-cGAMP to activate a Saf-2TM-SMODS-associated fused to various effector domains (SAVED) effector (CD-NTase-associated protein 14 [Cap14]), which initiates membrane disruption to restrict phage replication. Cryo-electron microscopy (cryo-EM) reveals that Cap14 binds 2′,3′-cGAMP to form a filament, while electrophysiology suggests that cGAMP activates membrane disruption. Swapping the Cap14 transmembrane domain with a nuclease domain yields a functional chimera that exclusively responds to 2′,3′-cGAMP. We hypothesize that other predicted transmembrane effectors in CBASS operons disrupt membranes, and we confirm this by showing that bacterial STING homologs with transmembrane domains restrict phage through membrane disruption. These findings expand our understanding of cGAS-STING-like pathways in bacterial immunity.
|
 Your new post is loading...
Your new post is loading...
































chang mw, microbiome, therapeutic, industry, clinical trial