Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 1:38 AM
|
Plant disease control is trapped in a constant evolutionary competition between plants and pathogens. Pathogens evolve faster than we can breed resistance genes or register new fungicides. Notably, plants employ multiple immune strategies, including both disease resistance and tolerance, to protect their health, enriching the contextual understanding of plant defense mechanisms. Transgenic crops, while powerful, continue to face regulatory and public acceptance challenges in many regions, with regulatory timelines varying across global frameworks. Meanwhile, chemical control faces an accelerating loss of efficacy due to resistance and is increasingly incompatible with climate-smart and biodiversity-friendly farming mandates. We argue that it is time to decouple crop immunity from plant genetics altogether and outsource it to an editable, transient, and regulation-resilient microbial layer: the endophyte microbiome.

|
Scooped by
mhryu@live.com
Today, 1:15 AM
|
The intracellular redox balance, which is determined by the relative levels of oxidants and reductants, plays a crucial role in regulating various cellular processes. Genetically encoded redox-sensitive green fluorescent proteins (roGFPs) are invaluable tools for spatiotemporally monitoring intracellular redox potential. However, the relatively low fluorescence intensity and weak response of existing sensors have restricted their applications in subcellular organelles, such as the Golgi apparatus, and in living animals. In this study, we rationally engineered an enhanced roGFP variant (eroGFP1.2), which exhibited a more than fivefold increase in fluorescence brightness and a 50% greater dynamic range compared to the wild-type roGFP1 in both E. coli and HeLa cells. Using the eroGFP1.2 sensor, we calibrated redox potential across different subcellular compartments within HeLa cells. Furthermore, when fused to human glutaredoxin-1 (Grx1), the chimeric sensor enabled specific and sensitive detection of glutathione redox dynamics in living cells. We also employed eroGFP1.2 to monitor a reactive oxygen species (ROS) burst at the wound site following tailfin transection in larval zebrafish. This work advances our understanding of GFP-based biosensor optimization and provides a powerful, high-sensitivity tool for investigating thiol-disulfide redox regulation in live cells and intact organisms.

|
Scooped by
mhryu@live.com
Today, 1:08 AM
|
Most bacteria are capable of living and surviving under a wide range of conditions, including exposures to chemical and physical stressors. A key to survival under adverse conditions is their ability to induce specific stress response systems that mitigate harmful effects. Examples are the well-studied SOS system induced under replication stress caused by DNA damage and the OxyR response induced by oxidative stress (e.g., hydroxyl radicals). In the present study, we have investigated interactions between these two systems under the assumption that harmful conditions may generally induce more than one bacterial response, and in such a case, their combined effect might feature additional factors critical for survival. We used E. coli strains that were constitutively induced for the SOS system (using the recA730 allele) in combination with the OxyR oxidative response (using the oxyR2c allele). The main finding was that joint expression of these systems strongly diminished the mutator effect associated with SOS induction. This so called SOS mutator effect is due to the error-prone DNA polymerase Pol V (product of the umuDC operon, UmuD′2UmuC), which is derepressed upon SOS induction. Gene expression studies revealed an overall reduction of genes of the SOS regulon in the double mutant, with the largest reduction for the genes of the umuDC operon. In parallel, Western blots showed the UmuD and UmuD′ proteins to be severely depleted from extracts of such strains. The data suggest an additional layer of control of SOS mutagenesis under certain stress conditions.

|
Scooped by
mhryu@live.com
Today, 12:46 AM
|
Marine algal-microbial symbioses constitute essential functional units that drive ocean biogeochemical cycles and trigger harmful algal blooms. Yet, a long-standing controversy persists regarding the mechanisms of algal-microbial symbiose assembly, specifically whether phycosphere microbiota are predominantly shaped by deterministic algal-driven selection or by stochastic environmental processes, with no definitive resolution to date. Here, we examined phycosphere communities associated with a series of Skeletonema strains, tracking their taxonomic and functional dynamics across successive growth stages. Despite pronounced taxonomic diversity, reflected in distinct community compositions, successional trajectories, and microbial networks, shotgun metagenomic analyses revealed highly conserved functional repertoires across samples, with consistently abundant core pathways, including amino acid biosynthesis, secondary metabolite and antibiotic production, and ABC transport systems. Statistical analyses further revealed a marked decoupling of taxonomy and function, with functional redundancy enabling taxonomically distinct lineages to perform equivalent metabolic roles. Based on these findings, we propose a dual assembly model in which deterministic algal host-driven selection constrains functional composition, while stochastic processes govern species-level membership. This “function-first, taxonomy-stochastic” paradigm reconciles opposing assembly theories, underscores functional resilience in the face of taxonomic turnover, and provides a conceptual foundation for the rational design of synthetic algal-microbial consortia in marine biotechnological applications.

|
Scooped by
mhryu@live.com
Today, 12:24 AM
|
The genetic code is highly redundant, with many synonymous codons encoding the same amino acid. Codon usage influences RNA structure, signaling, and translation rates. Differences in tRNA availability modulate elongation, with rare codons slowing translation and affecting co-translational folding and gene expression. Despite their functional importance and non-random distribution, rare codons are underrepresented in natural datasets, restricting the development of predictive models. We developed a transformer-based model that predicts codon sequences from amino acids, substantially improving rare codon prediction. The model learns codon signatures encoding species identity, RNA thermodynamic properties, and elongation constraints without explicit labels. Attention analysis shows that codon choice depends on both short and long-range sequence contexts, recovering dicodon effects and highlighting additional motifs. Finally, predictions correlate with experimental measurements of the impact of synonymous mutations on protein fitness, linking gene sequence to fitness and functional consequences, providing a framework to connect sequence variation, translation, and protein function.

|
Scooped by
mhryu@live.com
March 31, 11:55 PM
|
Plant immunity should be reconsidered beyond the boundaries of the plant genome. We propose that the rhizosphere microbiome may function analogously to a decentralized immune system, contributing adaptive defenselike properties and memory effects. In this forum article, we discuss how this perspective reframes immunity as an emergent property of plant–microbiome interactions, shifting the focus from a solitary host toward an integrated holobiont.

|
Scooped by
mhryu@live.com
March 31, 11:03 PM
|
Obligately lytic (virulent) phages always lyse host cells to release progeny viruses, while temperate phages can either lyse their hosts or integrate into host genomes as prophages, forming lysogens. There is a rich history of work studying the relative advantages and disadvantages of these two phage life history strategies, but little of this work has addressed the spatial constraints common to biofilm environments. We developed a live imaging system to track lytic infections, lysogenic infections, and uninfected cells at single-cell resolution within three-dimensional E. coli biofilms. We find that biofilm structure substantially impacts the ecological success of different phage infection strategies. Temperate phages have the unique capacity to release phages from lysogens that have undergone lytic induction from within the interior of mature biofilms. When this occurs in biofilm contexts that do not limit phage diffusion, lytic infections expand rapidly, but lysogenic infections are favored as phage mobility declines in densely packed biofilm architectures. In matrix-replete biofilms that do limit phage mobility, lytic phage infection is more limited, favoring lysogenic growth. Direct competition assays between lysogenized host bacteria and obligately lytic phages—with or without the ability to superinfect lysogens—revealed that spatial structure and superinfection potential together greatly impact phage competition outcomes during co-invasion into pre-existing, phage-susceptible biofilm populations. Highly packed, phage diffusion-impeding biofilms disproportionately favored temperate phages in the lysogenic cycle over obligate lytic phages, highlighting how biofilm architecture can constrain lytic phage infection and promote vertical phage genome transmission strategies.

|
Scooped by
mhryu@live.com
March 31, 10:27 PM
|
Proteomic studies have suggested that E. coli isocitrate lyase (ICL) undergoes multiple acetylation events, partially inhibiting its activity. However, the molecular basis of this regulation and the contribution of individual lysine residues had not been defined. This study demonstrates that acetylation of ICL in E. coli is acetyl-phosphate–dependent and reversible by the CobB deacetylase, establishing a key post-translational regulatory mechanism within the glyoxylate shunt. Site-specific acetylation at K13 and K308 inhibits ICL activity by destabilising the tetrameric assembly and rendering the protein more prone to degradation, whereas lysine-to-arginine substitutions at these positions alleviate this inhibition, enhancing carbon flux distribution, metabolic flexibility and biomass yield without the burden of plasmid-based overexpression. Leveraging this regulatory insight, a KR mutant bearing lysine-to-arginine substitutions at residues 13 and 308, engineered directly into the chromosomal aceA gene, maintained wild-type growth rates while reducing acetate overflow and improving metabolic balance during glucose depletion and acetate assimilation, leading to a 61% increase in lycopene production. These findings highlight regulatory-based metabolic engineering as a powerful strategy to optimise bioproduction and pave the way for extending this approach to other central metabolic enzymes to develop robust microbial cell factories for the sustainable synthesis of biofuels, biochemicals and high-value compounds.

|
Scooped by
mhryu@live.com
March 31, 9:31 PM
|
Disruption of organelle interactions due to metabolic stress is a crucial factor in the pathological processes of many degenerative diseases. Compared with animal cells, the participation of chloroplasts enables plant cells to show defensive adaptation under stress. Therefore, delivering plant-derived photosynthetic systems into animal cells may help to establish a more stable organelle interaction network. Here, we show that plant-derived photosynthetic systems can effectively restore homeostasis of the interaction network of animal organelles. Specifically, plant-derived nanothylakoid units provide energy to animal cells, regulate intracellular Ca2+ homeostasis, increase endoplasmic reticulum (ER) lipid unsaturation and global membrane fluidity, reduce abnormal contact between mitochondria and ER, and alleviate mitochondrial dysfunction. By combining implantable light-emitting diodes with wireless charging, we expand photosynthesis therapy, enabling treatments for deeper tissues. This study provides a proof-of-concept for disease treatment based on the regulation of organelle interaction networks by natural photosynthetic systems and establishes a therapeutic approach for treating deep tissues. Unlike animals, plant possess mitochondria and chloroplasts and exhibit a unique defensive adaptation to stress. Here, the authors demonstrate the feasibility of restoring organelle interaction network homeostasis in animal cells by introducing plant-derived photosynthetic systems.

|
Scooped by
mhryu@live.com
March 31, 1:43 PM
|
Rhizosphere microbiomes are essential for plant growth and stress tolerance, yet how microbial diversity shapes drought resilience in soybean remains unclear. Here, we demonstrate that high rhizosphere microbial diversity, generated via dilution-to-extinction manipulation of soil microbiome diversity, improves soybean performance under drought. Integrated metabolomic and transcriptomic analyses identify 2-naphthoic acid as a diversity-induced root exudate that accumulates exclusively under drought. This metabolite selectively recruits Sinorhizobium CS204 via chemotaxis and ATP-binding cassette (ABC) transporter-mediated uptake, as confirmed by in vitro substrate utilization assays and targeted mutant construction. Molecular docking and microscale thermophoresis reveal direct interactions between this metabolite and nitrogen-cycling proteins, enhancing denitrification and nitrogen fixation of S.CS204. Co-application of 2-naphthoic acid and S.CS204 significantly improves plant nutrient acquisition and photosynthesis under drought. Collectively, our study underscores the pivotal role of rhizosphere microbial diversity in triggering the exudation of root metabolites to recruit keystone taxa, establishing microbe-plant synergies that bolster drought tolerance.

|
Scooped by
mhryu@live.com
March 31, 1:25 AM
|
E. coli is a major workhorse for industrial biotechnology, yet reliance on plasmids is hindered by intrinsic instability. While genomic integration offers superior stability, it remains challenging for integrating large DNA fragments, particularly in difficult-to-engineer B strains (e.g., BL21) compared to K12 strains like MG1655. Moreover, a systematic study comparing genomic expression across B and K12 strains and different loci is absent. To address these issues, we firstly refined a CRISPR methodology that achieved unprecedented integration efficiencies in both strains: 40% for single-step insertion of a 17.5-kbp fragment in BL21 and 100% for a 9.5-kbp fragment in MG1655. Using this updated workflow, we rationally screened and compared 10 genomic loci in BL21 and MG1655 across varying media. The results demonstrated that host strain and genomic locus selection are more critical for expression than carbon source or medium composition. Although BL21 generally exhibits higher gene expression than MG1655, expression at certain loci can be very low. Therefore, we highlight the need for greater caution in locus selection for BL21 due to its higher variability. We leverage our validated loci for the scarless and marker-less integration of the astaxanthin synthesis pathway (23 genes, including multi-copy key genes, crtY, crtZ, crtW, crtE, crtB, and crtI) into the BL21 chromosome. The resulting plasmid-free strain produced 426 mg/L astaxanthin in glucose defined medium, showcasing a robust and generalizable strategy for complex pathway integration and optimization in challenging host strains.

|
Scooped by
mhryu@live.com
March 31, 1:18 AM
|
5-Aminolevulinic acid (5-ALA) is a basic precursor in the bioproduction of tetrapyrrole compounds, such as heme, chlorophyll, and vitamin B12. Its extensive applications in agriculture, medicine, cosmetics and animal feed industry have garnered substantial attention. In agriculture, 5-ALA works as an effective plant growth regulator, improving crop yield, stress tolerance, and product quality. Nonetheless, old chemical production of 5-ALA faces challenges because of complex reaction pathways, low productivity, and environmental concerns. This has directed towards a rising interest in sustainable microbial production approaches. Latest developments in metabolic engineering and synthetic biology have assisted the development of effective bacterial cell factories for 5-ALA bioproduction. Key approaches comprise optimizing precursor supply and cofactor regeneration, reworking inborn C4 and C5 biosynthetic pathways, classifying and alleviating rate-limiting enzymes, engineering transport systems for 5-ALA secretion, decreasing competing pathways, and reducing by-product formation. Furthermore, the addition of biosensor-based vigorous regulation and systems-level metabolic control has considerably boosted production robustness and yield. These technical novelties have accelerated the evolution of microbial 5-ALA production from the laboratory to industrial uses. This review offers a comprehensive overview of recent advancements in bacterial 5-ALA bioproduction, focusing on host selection, pathway engineering strategies, regulatory mechanisms, and evolving biotechnological applications. Lastly, it discusses present challenges and future perspectives to guide the development of next-generation microbial platforms for competent and sustainable 5-ALA synthesis.

|
Scooped by
mhryu@live.com
March 31, 1:08 AM
|
Understanding protein–ligand interactions in plants and microbes is essential for advancing agricultural biotechnology and developing antimicrobial strategies. In plants, these interactions govern critical physiological processes such as growth, immunity, and stress response, while in microbes, they influence pathogenicity, virulence, and survival. Experimental determination of bioactive compound–protein interactions is labor-intensive and limited in scope, and existing computational methods often suffer from rigid modeling assumptions. In this study, we collected and curated experimentally validated active and inactive compounds against plant and microbial target proteins from publicly available databases. Using this curated dataset, we developed machine learning-based classification models that predict compound bioactivity with an accuracy of 86% for plant proteins and over 90% for microbial proteins. These models form the core of PM-BioPred, a web-accessible prediction server designed to assist researchers in identifying potentially bioactive compounds against plant and microbial targets. PM-BioPred provides a user-friendly interface for submitting compound queries and retrieving bioactivity predictions. PM-BioPred is freely accessible to the academic community at https://pmbiopred.streamlit.app/ and serves as a valuable resource for plant-pathogen interaction research and compound repurposing efforts. The platform aims to facilitate the early-stage screening of agrochemicals and antimicrobial candidates.
|

|
Scooped by
mhryu@live.com
Today, 1:32 AM
|
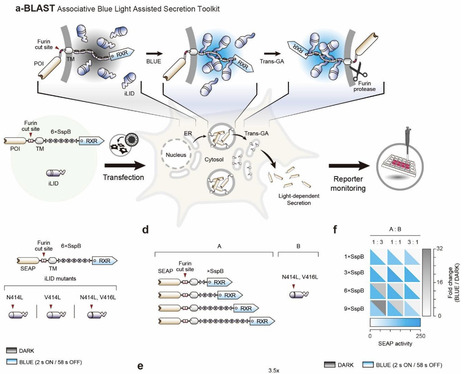
Precise control over protein secretion is essential for programming intercellular communication and coordinating complex physiological responses. However, conventional methods relying on transcriptional regulation or chemical induction often lack the spatiotemporal precision and reversibility required to mimic endogenous signaling dynamics. Here, we present the Blue Light-Assisted Secretion Toolkit (BLAST), a genetically encoded system that orchestrates protein release from the endoplasmic reticulum via light-tunable protein-protein interactions. BLAST comprises two complementary modules utilizing both light-induced iLID/SspB association (a-BLAST) and LOV2/Zdk1 dissociation (d-BLAST). Both modules harness the highly conserved RXR motif to enforce strict ER confinement in the dark state. Most importantly, by utilizing non-destructive steric masking rather than enzymatic cleavage, BLAST achieves unprecedented temporal resolution with strict reversibility. We demonstrate that both systems can be repeatedly toggled ON and OFF, instantaneously arresting cargo release upon light withdrawal to generate highly controlled, pulsatile secretion profiles. Leveraging this dynamic control, we successfully achieved the rapid, robust, and light-triggered secretion of complex therapeutic proteins, including insulin and interleukin-12. By bypassing transcriptional delays and irreversible activation steps, BLAST provides a generalized, plug-and-play platform for the on-demand delivery of therapeutic proteins, significantly expanding the optogenetic toolbox for synthetic biology and cell-based therapies.

|
Scooped by
mhryu@live.com
Today, 1:12 AM
|
In biotechnological processes, cell density and physiology are critical parameters for controlling the feed rate, harvest time, and process performance. We developed an automated flow cytometry approach that enables continuous, real-time (fully automated, hourly) monitoring of bacterial populations in continuous bioreactors. The method employed a double-staining protocol that combined DAPI to assess total DNA content and Alexa Fluor 488-EdU via Click-iT technology to identify the proportions of cells undergoing active DNA replication through EdU incorporation. The integrated workflow included fixation, permeabilization, staining, and measurement steps and was applied to three Gram-negative strains: Bradyrhizobium sp., Escherichia coli, and Stenotrophomonas rhizophila. Automated analysis captured growth dynamics and cell cycle progression, providing insights into population behavior under different dilution rates. In this study, automated on-line sampling enabled hourly flow cytometry measurements of cell concentration and physiological indicators during continuous cultivation, supporting real-time monitoring and control in industrial biotechnology.

|
Scooped by
mhryu@live.com
Today, 1:01 AM
|
Recent advances in single-cell technologies have revealed the dynamic and heterogeneous nature of host-pathogen interactions at the single-cell level. This review explores how cellular variability—both within clonal bacterial populations and among genetically identical host cells—gives rise to distinct infection outcomes, from pathogen clearance to persistence across multiple biological scales, from single cells to tissues and the whole organism. We highlight the conceptual and technological progress that has enabled the dissection of these interactions at single-cell resolution, including microscopy, single-cell transcriptomics, proteomics, and emerging dual RNA-seq and spatial approaches. Drawing on examples from well-characterized bacterial pathogens like Listeria monocytogenes, Salmonella enterica, and Mycobacterium tuberculosis, we discuss how stochastic gene expression, intrinsic and extrinsic factors, as well as tissue context shape the variable activation of the immune responses and ultimately determine the outcomes of host-pathogen interactions. We argue that the outcome of single-cell interactions is shaped by a combination of host states, bacterial-intrinsic features, and the local microenvironment. We further discuss how computational and mathematical modeling can integrate these heterogeneous single-cell events across spatial scales, linking intracellular variability with tissue-level pathogenesis and progression of infection. Gaining insight into and controlling these layers of variability holds promise for the development of more precise, context-dependent antimicrobial strategies.

|
Scooped by
mhryu@live.com
Today, 12:40 AM
|
How functional protein sequences are distributed in sequence space is fundamentally important for evolutionary theory and protein design, particularly if a large diversity of protein functions are hidden in evolutionarily unexplored areas of the sequence space. However, this question is understudied in part because experimental and computational studies use extant sequences as a starting point to study sequence space. Here, we study whether extant sequences are representative of the entire functional sequence space. Across thousands of protein families from vertebrates and bacteria we calculate the dimensionality and the volume of sequence space occupied by extant homologs. We find that the observed dimensionality and volume of extant sequence space are minuscule, many orders of magnitude smaller than what we estimated using a model of protein evolution. Simulating sequence evolution we then quantify the impact of phylogeny, selection, and epistasis on restricting the evolutionary exploration of sequence space. We find that sequence evolution from a single common ancestor, or a single point of origin in sequence space, is by far the largest limiting factor that reduces the dimensionality and volume of extant sequence space. These results indicate that there are vast areas of functional sequence space that have not been explored in evolution because of the excessive restrictions on natural exploration of the protein sequence space imposed by the point of origin effect. We suggest that protein design methods that rely on extant sequences may be limited in their ability to discover truly novel functions.

|
Scooped by
mhryu@live.com
Today, 12:07 AM
|
Overflow metabolism is a highly undesirable phenomenon that occurs in virtually all cell factories. Although several schemes have been applied to avoid overflow metabolism, this problem remains far from being solved. In this study, a genetic sensor for detecting overflow metabolism was designed, and its substrate specificity and reversibility were demonstrated. The circuit was then used to express the small protein SgrT as an output. SgrT inhibits the main glucose transport system in E. coli, thereby reducing the glucose uptake rate. This strategy eliminated overflow metabolism in batch cultures. Furthermore, overflow metabolism was reduced by 80% when the cells were transiently exposed to excess glucose during fed-batch cultures compared to the control cells. The proposed scheme can be useful for optimizing cellular performance and dealing with heterogeneous conditions typical of large-scale bioreactors.

|
Scooped by
mhryu@live.com
March 31, 11:52 PM
|
CRISPR‒Cas systems represent powerful tools for genome regulation. However, the large size of Cas proteins limits their efficient delivery via an adeno-associated virus (AAV), thereby restricting their clinical translation. Here, we engineer the IS200/IS605 transposon-encoded nuclease TnpB, along with its ωRNA scaffold, to create an enhanced TnpB system, which serves as a compact toolkit for gene activation, genome editing, and base editing. The gene activator enTnpBa increases expression by 2889-fold with a minimized 93 nt ωRNA and robustly activates endogenous genes in mammalian cells. We develop a single-AAV-based regimen for immune activation (AAV-ImmunAct) that delivers enTnpBa to activate CXCL9, IL-15, and IFN-γ. AAV-ImmunAct effectively enhances T cell migration and activation, increases killing of cancer cell lines and patient-derived organoids, and synergizes with anti-PD-1 therapy in humanized mice. Here, we establish enTnpB as a compact and versatile platform for genome regulation and a promising tool for cancer immunotherapy. CRISPR–Cas tools enable genome regulation but are often too large for efficient AAV delivery. Here, authors engineer a compact enhanced TnpB–ωRNA system (enTnpB) as a versatile genome regulation platform and develop a single-AAV regimen, ImmunAct, to activate endogenous cytokines and enhance cancer immunotherapy.

|
Scooped by
mhryu@live.com
March 31, 10:52 PM
|
Controlled plant growth in laboratories can be achieved by cultivating plants under sterile or axenic conditions on predefined synthetic growth media, typically supplemented with sugar. In nature, plants do not receive exogenous sugar supplies, form symbiosis with microbes, and plant growth is influenced by soil edaphic factors. Thus, physiological and multi-omic analyses of plants grown on synthetic media will differ from those of soil-grown plants due to the influence of sucrose, and the absence of microbiota and soil edaphic factors on plant growth. The rapid advances in spatial omics call for accurate characterization of plants grown under conditions similar to soil. To address the issue, we developed Artificial Soil (ArtSoil), a growth medium containing essential nutrients for plant growth, and aqueous soil extract (ASE) to maintain soil microbiomes and edaphic factors, simultaneously eliminating the need for sugar supplementation in the medium. We compared Arabidopsis thaliana grown on conventional media and on ArtSoil under various growth conditions. We showed that complex soil microbiota in ArtSoil promote plant growth without physiological side effects induced by sucrose. We demonstrate an application for ArtSoil in single-cell transcriptomics and report microbiota-induced cell-type-specificity in immune and nitrogen signaling. We tested ArtSoil with six types of ASEs to demonstrate its potential to decouple nutrient effects from microbiota in plant growth. We conclude that ArtSoil offers a more physiologically relevant alternative to conventional media for studying plant growth within a soil-like context.

|
Scooped by
mhryu@live.com
March 31, 9:45 PM
|
Rapid identification of viral infections and specific variants in patient samples requires a simple and multiplexed RNA detection method that does not rely on DNA sequencing. Although recent direct detection assays based on CRISPR–Cas13a offer rapid RNA detection by avoiding reverse transcription and DNA amplification required of gold-standard PCR assays, these assays are not easily multiplexed to detect multiple viruses or variants without dividing the sample into separate reactions. Here we show that Cas13a acting on single-target RNAs exhibits variable nuclease activity that depends on the interaction between the target RNA and crRNA. To exploit this feature for multiplexed detection, we devised a crRNA modification strategy that enables programmable tuning of Cas13a’s nuclease enzymatic rates. Using a droplet-based Cas13a assay, we demonstrate that kinetic signatures can be harnessed to differentiate among respiratory viruses and SARS-CoV-2 variants in contrived and clinical samples. This kinetic barcoding strategy can be extended to additional RNA targets through simple modification of crRNAs. A multiplexed RNA detection method exploits crRNA-dependent variability in Cas13a activity on RNA targets for kinetic barcoding and can be used to distinguish among SARS-CoV-2 variants in clinical samples.

|
Scooped by
mhryu@live.com
March 31, 9:02 PM
|
Although considerable research has been conducted on how bacteria respond to temperature, there are still few studies that use simple protein allocation models to characterize these responses. To bridge this gap, we utilize a protein allocation framework to delineate quantitative correlations among temperature, growth rate, and the fraction of functional protein sectors. By calibrating the model with experimental temperature-dependent growth curves of Escherichia coli, we predict how proteomic resources are reallocated across various temperatures. This approach enables us to elucidate, from a proteomic perspective, the temperature dependency of constitutive β-galactosidase expression and the variations in E. coli cell size. Our research deepens the understanding of bacterial physiological adaptation to temperature changes.

|
Scooped by
mhryu@live.com
March 31, 12:17 PM
|
Protein malnutrition remains a major global health challenge, particularly in regions where cereal grains dominate daily diets and access to diverse protein sources is limited. Cereals such as rice, wheat and maize provide most of the world’s calories, yet their grain proteins are often low in essential amino acids and poorly balanced for human nutrition. Improving both the quantity and quality of cereal protein therefore represents a critical opportunity to enhance human health while reducing reliance on environmentally intensive animal-based foods. In this Review, we synthesize recent advances in understanding how grain protein content and composition are regulated in cereals, and why protein enhancement has historically been constrained by trade-offs with starch accumulation and yield. We discuss how domestication and modern breeding reshaped carbon and nitrogen allocation in cereal grains, creating a starch-dominant optimum that limits protein concentration. Drawing on genetic studies from rice, maize and wheat, we highlight emerging strategies that improve nitrogen acquisition, amino acid transport, storage protein composition and endosperm buffering capacity, enabling partial decoupling of protein accumulation from yield penalties. Finally, we place cereal protein biofortification within a broader nutritional and environmental context. Enhancing protein density and amino acid balance in staple cereals can improve dietary adequacy for vulnerable populations while lowering greenhouse gas emissions per unit of nutrition. Together, these insights position cereal protein biofortification as a scalable and equitable pathway towards healthier diets and more sustainable food systems under global climate and population pressures. Cereal protein biofortification can improve nutrition while maintaining yield and lowering environmental impact. This Review shows how genetics and breeding can enhance protein quality in staple cereals to support healthier and more sustainable diets.

|
Scooped by
mhryu@live.com
March 31, 1:20 AM
|
Artificial intelligence (AI) is advancing synthetic biology biosensors (SBBs), driving a fundamental shift from rational design to AI-driven prediction. This review establishes a systematic framework linking AI algorithms to the Design-Build-Test-Learn (DBTL) cycle. We explicitly analyze the engineering paradigms of AI-enabled cell-based SBBs and AI-optimized cell-free SBBs, highlighting how computational intelligence addresses platform-specific bottlenecks. Crucially, we synthesize the AI-driven workflow into three core frontiers: AI-guided robust sensor element design, AI-assisted signal processing for accurate performance characterization, and AI-driven closed-loop optimization to accelerate autonomous evolution. Furthermore, representative applications of SBBs are investigated, including multi-pollutant environmental detection, continuous biomarker monitoring, food safety tracking, and intelligent biomanufacturing. Beyond achievements, we critically evaluate unresolved obstacles, notably the “reality gap” and the “small-data dilemma”. Finally, we propose a roadmap centered on bio-digital hybrid interfaces, explainable AI, and data standardization to accelerate the transformation of SBBs into robust, field-deployable intelligent sensing systems.

|
Scooped by
mhryu@live.com
March 31, 1:12 AM
|
As an excellent industrial chassis, Bacillus subtilis plays a pivotal role in promoting the efficient and sustainable manufacturing of high-value biological products. However, the intrinsic resource trade-off between growth and production remains a formidable barrier for biosynthetic efficiency. Therefore, from the perspective of cellular resource allocation, this review provides a comprehensive analysis through a holistic framework encompassing synthetic toolkits, mechanisms, engineering strategies, and industrial applications. We analyze the fundamental limitations of resource partitioning, focusing on the proteome allocation competition driven by heterologous synthesis and the global physiological feedback responses triggered by metabolic burden. To address resource constraints, the paper discusses fundamental engineering modules to enhance strain robustness and metabolic capacity, as well as system-level strategies for rational resource allocation. Finally, by showcasing diverse industrial applications, this work provides profound insights into overcoming metabolic trade-offs and lays the foundation for the rational design of high-performance B. subtilis cell factories.
|
 Your new post is loading...
Your new post is loading...