 Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 12:24 AM
|
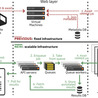
The HMMER web server, available at https://www.ebi.ac.uk/Tools/hmmer, provides online access to tools from the HMMER software suite (http://hmmer.org/) for protein analysis using profile hidden Markov models. Users can perform sequence similarity searches against a range of regularly updated protein sequence databases or annotate protein sequences with domains and families using profile HMM libraries from protein family databases. Since the 2018 update, the continued exponential growth of sequence databases has necessitated substantial infrastructural improvements to maintain search performance speed and service reliability. To achieve this, the web interface has been completely reengineered using modern web technologies (JavaScript and React), providing users with an enhanced experience, including session-based search history and streamlined results visualization. The web application programming interface has been rewritten to better support programmatic access with updated endpoints and JSON-based responses. The infrastructure has been redesigned to efficiently handle searches against much larger databases through horizontal scaling and asynchronous job processing. Target database offerings have been updated to reflect current usage patterns and data availability. The HMMER web server is free and open to all users, and there is no login requirement.

|
Scooped by
mhryu@live.com
Today, 12:10 AM
|
Plant-associated microorganisms can break down many contaminants, including naphthenic acid fraction compounds (NAFCs)—one of the toxic contaminants in oil sands process-affected water (OSPW). One way to take advantage of these processes is to use constructed wetland treatment systems (CWTSs), where aquatic plants work in coordination with microorganisms to remediate OSPW. However, the dynamics of the microbial communities and the potential contribution of distinct bacteria and fungi inhabiting sediments, rhizosphere, and water that comprise these systems remain poorly understood. To address this, we planted water sedge (Carex aquatilis) in mesocosm-scale wetland systems and compared it with unplanted controls. Naphthenic acid fraction compound (NAFC) concentrations and microbial community dynamics among various mesocosm compartments were monitored using mass spectrometry and high-throughput sequencing of the bacterial-archaeal 16S rRNA gene and the fungal ITS2 region. Carex-planted mesocosms reduced NAFC concentrations more rapidly than the unplanted controls, which coincided with shifts in abundance for both bacterial and fungal communities, especially in the Carex rhizosphere. The strongest relationships between microbial taxa and the decrease in NAFC concentrations were for anaerobic bacteria, suggesting a prominent role for these bacteria in NAFC degradation in CWTS. This study highlights potential microbial targets for improving CWTS efficiency for NAFC detoxification during OSPW remediation.

|
Scooped by
mhryu@live.com
April 27, 11:58 PM
|
The number of published isothermal amplification assays has increased substantially in recent years. Unfortunately, no harmonized guidelines, such as the Minimum Information for Publication of Quantitative Real-Time PCR Experiments guidelines, exist for publishing these methods, often resulting in incomplete reporting of assay composition and performance. In this study, we systematically evaluated nine published loop-mediated isothermal amplification (LAMP) assays for the detection of Pseudomonas aeruginosa. We aimed to assess whether publications provide (i) sufficient information on assay composition to allow implementation and reproduction and (ii) robust data on assay performance to evaluate their applicability. Assays were screened for basic functionality, analytical specificity, sensitivity, and limit of detection (LOD) in head-to-head experiments with qPCR. Four assays lacked essential composition details, and almost all did not report DNA concentrations or replicate numbers. Only six assays consistently amplified target DNA. Analytical specificity testing with 19 non-target strains contradicted previously reported 100% specificity, with only 3 maintaining specificity above 90% in our evaluation. Sensitivity testing with 13 P. aeruginosa strains confirmed 100% sensitivity for two assays. However, LOD experiments revealed significantly higher values than originally reported, with qPCR outperforming all LAMP assays. These findings highlight substantial discrepancies between published data and real-world assay performance. The absence of standardized formats, consistent units, and complete methodological details undermines replicability. Although this study focused on P. aeruginosa, the identified issues are widely relevant across different microbial targets. We advocate for increased publication standards and quality controls to ensure transparency, utility, and comparability of isothermal amplification assays and to support their translation into clinical and environmental applications. benchmark

|
Scooped by
mhryu@live.com
April 27, 11:12 PM
|
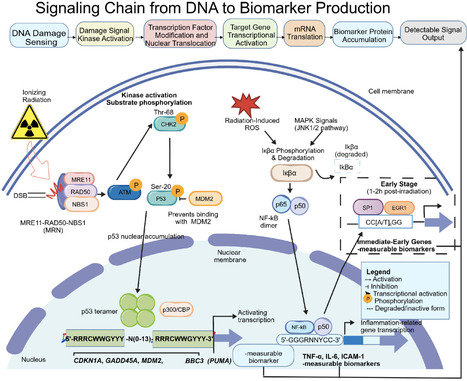
Radiation-responsive promoters represent a functionally distinct class of transcriptional regulatory elements that translate genotoxic stress signals into quantifiable gene expression outputs. These promoters occupy a unique mechanistic position within the broader radiation biomarker landscape: rather than directly measuring molecular damage products, they report the cellular interpretation of radiation-induced stress through coordinated gene regulatory networks. This review provides a systematic analysis of five major classes of radiation-responsive promoters—microRNA (miRNA) promoters, tRNA-derived small RNA (tsRNA) promoters, acute-phase protein gene promoters, DNA repair gene promoters, and long non-coding RNA (lncRNA) promoters—with emphasis on their regulatory logic, dose-response characteristics, and current evidence for clinical deployment. We further describe four complementary screening strategies: homology-based conservation analysis, functional genomics and transcriptomics, epigenetic modification profiling, and synthetic biology promoter engineering. Applications spanning biosensor development, biological dosimetry, treatment response prediction, and radiation-guided gene therapy are evaluated within a two-track framework that distinguishes biomarker-oriented applications (Track A) from tool-oriented reporter gene systems (Track B). Critical appraisal of current limitations—including insufficient clinical-grade validation, absence of standardized dose-response curves, and reproducibility deficits—is integrated throughout. Future priorities include multi-center prospective validation studies, FAIR-compliant data infrastructure, AI-driven multi-omics integration, and point-of-care detection platforms. Radiation-responsive promoter biology holds significant potential for advancing precision radiotherapy and nuclear emergency medical response, contingent upon systematic closure of the current evidence gap relative to established gold-standard cytogenetic methods.

|
Scooped by
mhryu@live.com
April 27, 11:00 PM
|
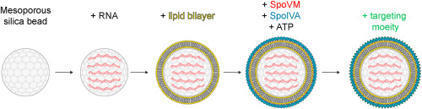
RNA therapy, which includes delivery of mRNA or siRNA, shows promise for treating various diseases, but difficulties in targeting specific cell types and low efficiency of loading RNA into nanoparticles remain hurdles to achieving widespread use. Previously, we reported the assembly of biocompatible synthetic bacterial spore-like particles, termed “SSHELs”, which are built atop a porous silica core encased in a lipid bilayer and two bacterial proteins that form a stable proteinaceous surface that may be covalently modified with targeting proteins of interest. Here, we employ micron-scale SSHELs constructed using a fusogenic lipid and decorated with affibodies targeting cell surface HER2 to specifically deliver model mRNA and siRNA molecules specifically to HER2-positive ovarian and breast cancer cells, with high RNA loading efficiency and cargo capacity. SSHEL particles therefore represent a versatile vehicle for the delivery of not only small molecules, but also therapeutic RNA to specific cell types.

|
Scooped by
mhryu@live.com
April 27, 7:40 PM
|
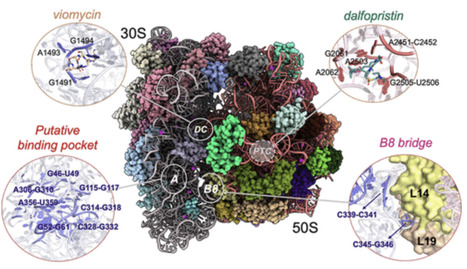
The bacterial ribosome is a key antibiotic target, yet peptide-based modulation from its functional and allosteric sites is underexplored. We developed a computational pipeline combining SiteMap-derived binding-site detection, consensus docking with Glide and rDock, all-atom truncated molecular dynamics (MD), and coarse-grained MD (CGMD) simulations to identify peptide candidates against four E. coli ribosomal sites: the decoding center, peptidyl transferase center, a putative binding pocket on 30S, and the intersubunit bridge B8. Consensus-selected peptides recapitulated hallmark contacts of the native inhibitors viomycin and dalfopristin, and their interaction fingerprints delineate site-specific scaffolds that enable prioritization of inhibitor candidates with enhanced ribosomal affinity, thereby guiding the rational design of novel and effective peptide-based therapeutics. Notably, the peptide CycPeptMPDB_2508 exhibited binding affinity across all investigated sites, nominating it as a versatile lead core for antimicrobial peptide design. Dynamic cross-correlation matrices derived from CGMD simulations captured coupled motions between distal regions of the ribosome, while residue interaction network analysis identified hub residues enriched near the putative binding pocket and B8 bridge, outlining putative allosteric pathways linking local pockets to global motions relevant to decoding and domain closure. This work provides a concise, testable framework for ribosome-targeted peptide discovery and, to the best of our knowledge, constitutes the first ribosome–peptide virtual screening study to employ the viparr module for truncated ribosome–peptide complexes, suggesting the potential applicability of this approach to complex systems and broadening its scope.

|
Scooped by
mhryu@live.com
April 27, 7:23 PM
|
Melanin, a multifunctional biopolymer, holds significant potential in biomedicine, agriculture, and food industries. However, challenges in low solubility and inefficient extraction hinder its industrial-scale production. This review systematically categorizes melanin types (eumelanin, pheomelanin, allomelanin, neuromelanin, pyomelanin) based on their biosynthetic pathways and biological sources. We critically analyze state-of-the-art strategies for enhancing melanin yield, including genetic engineering, fermentation optimization, and physicochemical interventions. Furthermore, we highlight emerging applications in biopesticides, food packaging, and cancer therapeutics. By bridging biosynthesis mechanisms with scalable production technologies, this work provides a roadmap for advancing melanin utilization in sustainable industries.

|
Scooped by
mhryu@live.com
April 27, 1:01 AM
|
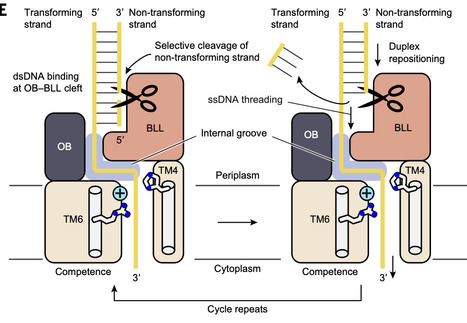
Natural transformation is one of the major pathways of horizontal gene transfer in bacteria, enabling the acquisition of extracellular DNA and its integration into the host genome. ComEC is a membrane protein responsible for DNA translocation in this process, yet its precise function and structure have remained elusive. Here, we report cryo–electron microscopy structures of ComEC in DNA-free, single-stranded DNA (ssDNA)–bound, and double-stranded DNA (dsDNA)–bound forms, together with biochemical analyses. These structures reveal that ComEC cleaves one strand of dsDNA at its extracellular domain and guides the remaining strand into a positively charged pore formed within the membrane domain. These findings provide a structural basis for the long-hypothesized roles of ComEC in both DNA processing and translocation across the inner membrane during natural transformation.

|
Scooped by
mhryu@live.com
April 27, 12:51 AM
|
Bacteria often survive viral attack and environmental stress by sharing genes that enhance their defenses. The cholera pathogen Vibrio cholerae carries a sedentary chromosomal integron (SCI), a genetic element containing hundreds of mostly promoterless gene cassettes, about 10% of which encode antiviral systems. Cassettes are thought to reshuffle under stress to the favorable first array position, yet the SCI in pandemic V. cholerae has remained static for more than 60 years. In this study, we show that SCI diversification efficiently occurs by horizontal transfer linked to the genus’s aquatic lifestyle: DNA released from lysed cells is taken up by naturally competent vibrios and integrated into the first position of the SCI array, the primary site of strong expression, where it confers resistance to phage and potentially other threats.

|
Scooped by
mhryu@live.com
April 26, 11:40 PM
|
This study investigates the relationship between the gut microbiota and specific diseases. Data was collected from the Human Gut Microbiome Atlas, which examines regional variations across 20 countries on five continents, categorizing microbial species by taxonomy, from genus to species. The Atlas provides color-coded phylum classifications, numerical species counts within the same genus, and an analysis of dysbiosis-related associations with 23 diseases, as well as region-enriched species. The data stratified samples into distinct categories such as westernized, non-westernized, cancerous, and non-cancerous. The findings demonstrate that tree-based ensemble methods, such as Bagging and Boosting prediction methods, achieved the highest accuracies across all categories due to their robustness in handling the complex, high-dimensional data. The XGBoost model yielded the strongest predictive performance, achieving 91% accuracy for westernized cancer-associated samples, 84% accuracy for non-westernized cancer-associated samples, 92% accuracy for westernized samples, and 78% for non-westernized samples. Additionally, advanced topological data analysis was used to assess the global structure and underlying patterns within the dataset.

|
Scooped by
mhryu@live.com
April 26, 12:39 PM
|
The site-specific incorporation of noncanonical amino acids (ncAAs) has revolutionized protein research. However, synthesizing multifunctional biopolymers demands moving beyond single-type ncAA incorporation to simultaneously encoding multiple distinct ncAAs. This review summarizes transformative progress in the genetic encoding of multiple distinct ncAAs, including the development of mutually orthogonal translation systems and the expansion of codon repertoires. We highlight recent breakthroughs that have achieved the incorporation of multiple distinct ncAAs. Notably, integrating rare codon recoding with stop codon suppression in mammalian systems has recently enabled the genetic encoding of five distinct ncAAs. These innovations establish a robust platform for advanced biosensors, next-generation therapeutics, and synthetic biology with expanded chemical repertoires.

|
Scooped by
mhryu@live.com
April 26, 11:19 AM
|
Recent advances in Machine Learning have transformed antibody development through in silico models, accelerating therapeutic candidate identification. However, challenges persist: rapid adaptation of property predictors to laboratory-specific assays with incomplete datasets; batch effects introducing systematic bias; assay costs necessitating efficient unseen property prediction. We introduce a novel multimodal architecture featuring specialized tokenization and embedding projection that integrates text and protein language models (pLM) and a learning strategy to enable context-conditioned multi-property prediction without learning shortcuts. Our framework enables prompting without dictionary merging across modalities, creating a compact model capable of context-conditioned learning for multi-property prediction. The orchestrating model avoids pLM-to-text projection while enabling inference-time adaptation without retraining. Using 876,898 antibody heavy chain sequences with batch effect simulation, our architecture achieved Spearman’s ρ > 0.8 across multiple developability properties, significantly outperforming fine-tuned multimodal LLMs and showed the ability to leverage correlation between properties for prediction. This approach has the potential to address critical antibody development challenges.

|
Scooped by
mhryu@live.com
April 25, 11:56 PM
|
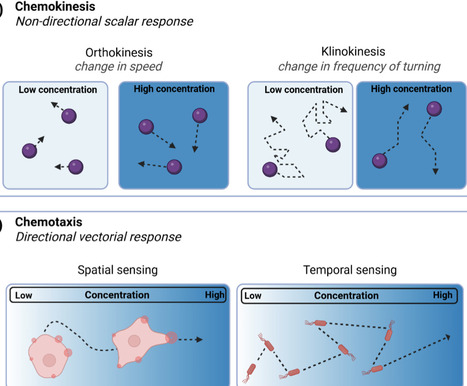
Chemotaxis is the directional motion of objects in response to chemical gradients, a process that drives navigation in micro/nanomotors, enabling a bio-inspired transition toward autonomous direction control. However, current studies lack consistent mechanistic justification, definition, and standardized experimental validation. This review summarizes key physical principles governing active chemotaxis, surveys experimental strategies for generating chemical gradients and quantifying responses, and examines how individual chemotactic mechanisms and long-range, anisotropic chemical interactions give rise to emergent collective behaviors, while outlining prospective applications. Together, these efforts establish a principled engineering framework for advancing chemotaxis as a robust functional navigation modality in synthetic micro/nanomotors. This Review Article examines artificial chemotaxis in micro/nanomotors, detailing mechanisms, experimental strategies, and collective behaviors. It highlights the need for standardized validation and explores applications in biomedical and industrial fields, emphasizing the challenges and potential of synthetic chemotaxis.
|

|
Scooped by
mhryu@live.com
Today, 12:16 AM
|
Although valued for their nutritional content and flavor, peanuts are ranked among the nine major food allergens in the United States, capable of inducing life-threatening reactions mediated by immunoglobulin E (IgE). Ara h 1, Ara h 2, Ara h 3, and Ara h 6 are recognized as major allergens due to the prevalence and structural stability of their epitopes, which make them resistant to digestion and processing. To address these challenges, processing treatments of different natures have been tested (e.g., physical, chemical, and biological) to reduce the allergenic content and create safer and more nutritious nuts. Allergen degradation studies discussed in this paper highlight the roles of endogenous peanut enzymes and microbial metabolites, as well as computational tools useful to model theoretical protein transformation under degradative treatments. Fermentation is emphasized for its multifactorial potential, combining the synthesis of bioactive metabolites by generally recognized as safe (GRAS) microbes with their proteolytic and redox activity to reduce allergenicity in peanuts while improving nutritional and sensory quality. Despite significant progress of these approaches, the development of a viable hypoallergenic peanut product remains elusive. Therefore, potential fermentation enhancers are discussed, including the addition of naturally extracted polyphenols that can alter the structure of peanut epitopes while increasing their proteolytic susceptibility. This review integrates structural insights, physical, chemical, and biological interventions, and computational approaches to provide a comprehensive perspective on peanut allergen reduction. By emphasizing epitope-targeted fermentation and synergistic treatments, we outline directions for developing safer peanut-based foods that balance efficacy, feasibility, and consumer acceptance.

|
Scooped by
mhryu@live.com
Today, 12:06 AM
|
The exceptional pressure resistance of bacterial spores limits the application of high hydrostatic pressure (HHP) in food sterilization, yet the underlying mechanisms remain poorly understood. Here, we examined how structural modifications (coat, cortex, inner membrane [IM], α/β-type small acid-soluble proteins [α/β-type SASPs]) in Bacillus subtilis spores affect HHP-induced germination, superdormant (SD) subpopulation, and inactivation, which together define HHP resistance. Key findings reveal that (i) coat defects likely alleviate cortex compression on IM, thereby influencing germination-associated proteins. This alteration reduces germination efficiency at 200 MPa but enhances it at 500 MPa, leading to an expanded SD subpopulation and increased resistance at 200 MPa, while a diminished SD ratio and reduced resistance at 500 MPa; (ii) increased cortex cross-linking combined with core expansion potentially modifies IM properties, strongly affecting germination-related proteins. This change markedly promotes germination under both 200 and 500 MPa, resulting in decreased SD ratios and thus lower spore resistance under both conditions; (iii) reduced cardiolipin content in IM likely alters its biophysical properties, specifically increasing germination efficiency at 500 MPa. This contributes to a lower SD ratio and reduced resistance at 500 MPa; (iv) the absence of α/β-type SASPs has minimal effect on resistance at 500 MPa but reduces resistance at 200 MPa by decreasing the SD ratio, potentially due to spontaneous germination after treatment. Collectively, HHP resistance is modulated by multiple structural components through distinct mechanisms. Structural modifications that alter IM biophysical properties or the function of germination-related proteins appear to be critical regulators of pressure-induced germination.

|
Scooped by
mhryu@live.com
April 27, 11:16 PM
|
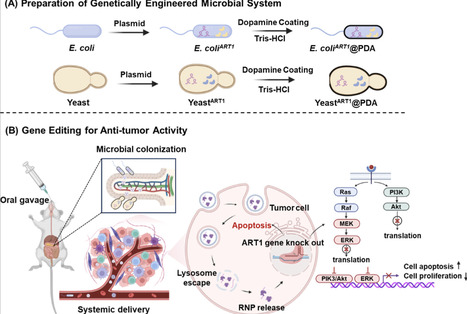
Colorectal cancer (CRC) poses a serious threat to human health. CRISPR/Cas9 technology offers new therapeutic strategies for the management of this disease, but its oral application is severely hindered by the limitations of suitable delivery systems. Herein, we develop and compare two separate orally delivered, genetically and chemically modified CRISPR/Cas9 delivery platforms based on E. coli BL21 and Pichia pastoris X33, which upon colonization in the intestine, secreted extracellular vesicles carrying the Cas9 protein and ART1-targeting sgRNA for tumor-specific gene disruption. Arginine ADP-ribosyltransferase 1 (ART1) plays a crucial role in the biological regulation of colon cancer, which was for the first time to the best of our knowledge, employed in vivo as a target gene in this study. Furthermore, we employed polydopamine (PDA) coating and gastrointestinal synthetic epithelial lining systems to facilitate microbial viability and intestinal retention, establishing on site cell factories for sustained CRISPR secretion. In subcutaneous tumor-bearing murine models, both delivery systems demonstrated comparable antitumor efficacy with significant tumor suppression. Taken together, the genetically modified microbial platform using bacterial and yeast strategies shows great potential and broad therapeutic versatility, offering a promising CRISPR-based solution for CRC treatment.

|
Scooped by
mhryu@live.com
April 27, 11:06 PM
|
Engineering of microbial cells, including E. coli, is essential in prototyping genetic designs used in numerous applications throughout synthetic biology. While many advanced genome editing tools, such as CRISPR-based tools, offer new capabilities with genetically recalcitrant organisms, these tools often do not offer an immediate advantage in readily manipulated microbes, such as E. coli, especially when scarless modifications are not critical. We describe a comprehensive recombineering tutorial that we commonly use for multiplex engineering of E. coli using antibiotic markers. We leverage a group of 15 antibiotic resistance cassettes, most of which can be readily included when designing double-stranded DNA donors intended for recombineering and purchased from several vendors. Using these methods, 10–15 defined modifications to a single host strain can be achieved in less than three weeks, using two-day editing cycles. We discuss sequences and protocols as well as the optimal design of genetic modifications and the associated DNA.

|
Scooped by
mhryu@live.com
April 27, 10:56 PM
|
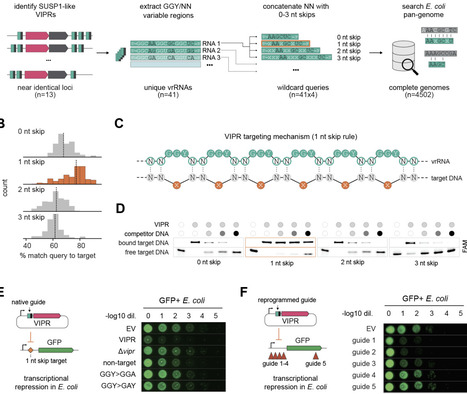
CRISPR-Cas systems use RNA-guided proteins for adaptive immunity through a mechanism whose origin is unknown. Here we report the discovery of Viral Interference Programmable Repeat (VIPR) systems consisting of a Vipr protein more ancient than CRISPR-Cas and vrRNAs comprising alternating GGY/NN motifs. Unlike canonical guide RNAs that base pair with nucleic acid targets using an uninterrupted sequence, vrRNAs recognize double-stranded DNA through a noncontiguous code in which the variable NNs of each repeat collectively specify a target that itself contains a gapped recognition sequence. Analysis of natural vrRNA targets suggests VIPR acts against competing phages. We demonstrate programmable phage defense by redirecting the complex for transcriptional repression. These results suggest that the roots of adaptive immunity lie in ancient warfare between viruses, and reveal a new logic for programmable genetic control.

|
Scooped by
mhryu@live.com
April 27, 7:37 PM
|
To overcome protein aggregation and poor secretion during the production of the thermophilic glycogen branching enzyme from Aquifex aeolicus (AaGBE) in Bacillus subtilis, this study developed a two-stage fermentation strategy that decouples cell growth from protein folding. This physiological approach was combined with multilevel engineering, including a protease-deficient host, a signal peptide-independent secretion pathway, a high-copy plasmid with tandem promoters, and N-terminal coding sequence optimization. The integrated strategy effectively alleviated aggregation, achieving an extracellular AaGBE titer of 0.42 g/L and an activity of 1,012 U/mL─representing a 55-fold improvement over the unoptimized strain. The purified AaGBE exhibited a specific activity of 2389 U/mg and retained robust thermostability. These results demonstrate that decoupling cell growth from protein folding provides an effective strategy for the secretion of aggregation-prone industrial enzymes.

|
Scooped by
mhryu@live.com
April 27, 1:11 AM
|
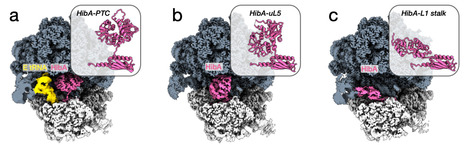
Ribosome hibernation preserves translation machinery during stress, yet its mechanisms in Archaea remain poorly defined. Using cryo-EM analysis, we studied hibernation pathways in Pyrococcus abyssi stressed cells. We identified HibA, a previously unrecognized family of hibernation factors widespread in Archaea. HibA consists of a bacterial-like HPF/RaiA domain fused to a Cystathionine Beta Synthase module. Unexpectedly, HibA binds to the ribosome in three different conformations, occupying the A, P and E sites of tRNAs, as well as that of mRNA, enhancing its ability to protect the ribosome from degradation. Idle ribosomes also frequently accumulate the archaeal homolog of eukaryotic ribosome maturation protein SBDS (aSBDS), suggesting that stressed archaeal cells may engage parallel hibernation routes in which aSBDS can complement HibA. Deletion of hibA in Thermococcus barophilus delays recovery from stationary phase and reduces 70S ribosome pools, establishing its role in ribosome preservation. Taxonomic profiling shows that many archaeal lineages encode distinct repertoires of ribosome-associated protection factors, underscoring the modular and multi-layered nature of archaeal hibernation systems. In addition, a comprehensive phylogenetic analysis highlights the evolutionary relationships between prevalent ribosome hibernation factors across Bacteria and Archaea. Ribosome hibernation preserves translation machinery during stress. Here, the authors identify a ribosome hibernation factor that is widespread in archaea and combines bacterial HPF/RaiA-like and cystathionine beta-synthase domains.

|
Scooped by
mhryu@live.com
April 27, 12:55 AM
|
Monitoring molecular activities within single live cells is vital for understanding cellular differentiation, senescence, heterogeneity, and disease progression. However, conventional single-cell analyses often rely on micromanipulation or extraction followed by downstream measurements, which cannot capture in situ real-time dynamics. Fluorescent labeling and electrochemical methods provide temporal resolution but face limitations in labeling, substrate scope, and multiplexing. Here, we present a nanopore probe that enables real-time, multiplexed monitoring of intracellular activities in single live cells. The device integrates an aluminum oxide nanostraw membrane for molecular extraction and a glass nanopore membrane for single-channel electrical detection. Using a hippocampal neuron model of ischemia-hypoxia, we simultaneously tracked dynamic changes in intracellular glutamate, ascorbic acid, and adenosine triphosphate—three key molecules involved in oxygen-glucose deprivation-induced neuronal edema. Our findings establish this nanopore probe as a powerful platform for real-time, label-free molecular profiling at the single-cell level, opening opportunities for studying disease mechanisms and therapeutic responses.

|
Scooped by
mhryu@live.com
April 27, 12:43 AM
|
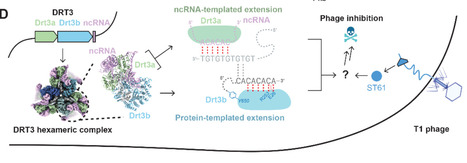
Defense-associated reverse transcriptases (DRTs) are widespread bacterial antiphage systems that use unconventional mechanisms of polynucleotide synthesis. We show that DRT3, which comprises two distinct RTs (Drt3a and Drt3b) and a noncoding RNA (ncRNA), synthesizes alternating poly(GT/AC) double-stranded DNA. Cryo–electron microscopy structures at 2.6 Å resolution reveal a D3-symmetric 6:6:6 complex of Drt3a, Drt3b, and ncRNA. Drt3a produces the poly(GT) strand using a conserved ACACAC template within the ncRNA. Notably, Drt3b synthesizes a complementary, protein-primed poly(AC) strand in the complete absence of a nucleic acid template, using conserved active site residues specific to Drt3b to enforce precise base alternation. These findings expand the functional landscape of nucleic acid polymerases, revealing a protein-templated mechanism for sequence-specific DNA synthesis.

|
Scooped by
mhryu@live.com
April 26, 12:41 PM
|
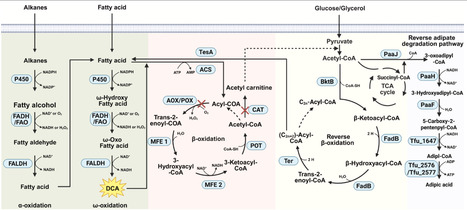
Medium- and long-chain dicarboxylic acids (M/LCDAs) are key monomers for the synthesis of nylons and high-performance engineering plastics. Compared to traditional chemical methods, microbial synthesis offers advantages such as environmental friendliness and high regioselectivity. However, its industrial application remains limited by bottlenecks, including low mass transfer efficiency on hydrophobic substrates, instability of key oxidase systems, and cellular metabolic imbalances. This review summarizes recent strategies leveraging enzyme engineering, systems metabolic engineering, and diverse synthetic biology approaches to overcome current limitations in the biosynthesis of M/LCDAs. We specifically highlight mechanisms for enhancing the transmembrane transport of hydrophobic substrates and the mining of novel transporters. Furthermore, we elaborate on protein engineering efforts targeting key enzymes (e.g. cytochrome P450s), covering rational design, fusion expression, and novel dimerization techniques. At the systems level, we discuss metabolic network regulation achieved through the construction of the reverse β-oxidation cycle (r-BOX) and the reprogramming of cofactor regeneration and energy metabolism. Finally, future perspectives on integrating AI-aided design and waste valorization are proposed to provide theoretical guidance for the efficient and sustainable biomanufacturing of M/LCDAs.

|
Scooped by
mhryu@live.com
April 26, 11:22 AM
|
Nitrate (NO3−) serves as a pivotal molecule with dual functions in nutrient supply and signaling during plant growth and development. Precise monitoring of its spatiotemporal dynamics in planta is therefore essential for dissecting the regulatory mechanisms underlying plant nitrogen metabolism. However, conventional nitrate detection methods suffer from inherent limitations, including destructive sampling, insufficient spatiotemporal resolution, and an inability to achieve real-time whole-plant monitoring. Here, we report a genetically encoded nitrate biosensor, designated NitNRCL1, constructed using a split firefly luciferase complementation system. Functional validation in both prokaryotic and eukaryotic systems demonstrates that NitNRCL1 responds to changes in nitrate availability and generates stable chemiluminescent signals in bacteria and diverse plant species. Importantly, NitNRCL1 enables non-invasive, real-time, and whole-plant monitoring of nitrate levels in living plants. Using NitNRCL1, we successfully imaged the spatiotemporal dynamics of nitrate signaling in Arabidopsis thaliana. Collectively, our findings establish NitNRCL1 as a robust and novel tool for investigating nitrate transport, signaling, and metabolic pathways in plants. This biosensor advances our mechanistic understanding of plant nitrate biology and provides a technical foundation for breeding nitrogen-use-efficient crops and developing precision fertilization strategies.

|
Scooped by
mhryu@live.com
April 26, 11:16 AM
|
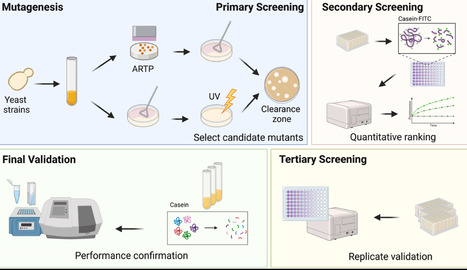
Efficient secretion of heterologous proteases in S. cerevisiae remains a major bottleneck due to host cytotoxicity and secretory stress. In this study, we developed a yeast chassis strain through three rounds of iterative evolution, combining ultraviolet (UV) and atmospheric and room temperature plasma (ARTP) mutagenesis with a fluorescence-based screening pipeline. The final strain, MA10Ah121, exhibited protease production nearly three times that of its parental strain LB184M and maintained genetic stability across passages. Importantly, MA10Ah121 supported enhanced expression of two structurally distinct collagenases, ColG (Class I) and ColH (Class II), demonstrating its broad utility for recombinant protease production. Whole-genome sequencing revealed substantial chromosomal remodeling, including reversion of a 151 kb duplication enriched in genes related to energy metabolism and protein synthesis, along with widespread structural variations. These changes reflect a genome-wide adaptation that rebalances biosynthetic capacity and stress tolerance. This work provides a robust and versatile S. cerevisiae chassis for the biosynthesis of proteases and other challenging recombinant enzymes in industrial biotechnology applications.
|
 Your new post is loading...
Your new post is loading...






















tn-seq