Your new post is loading...
 Your new post is loading...

|
Scooped by
Jean-Michel Ané
Today, 7:17 PM
|
Rooted in place, plants must continuously respond and adapt to their ever-changing environment to survive, especially as climate change intensifies. Calcium ions (Ca²⁺) play a central role in plant responses to both biotic and abiotic challenges. Ca²⁺ signaling involves the coordinated action of channels and transporters that generate specific “Ca²⁺ codes,” along with Ca²⁺-binding proteins that act as sensors to decode them. Studies over the past several decades have explored the molecular components that form the toolkit, pathways, and networks for the coding and decoding of Ca²⁺ signals in plants. This review focuses on the emerging mechanisms of calcium signaling in plants, beginning with an overview of the universal conceptual framework that governs the coding and decoding of Ca²⁺ signals, followed by examples of pathways in plant growth and reproduction, responses to abiotic stress and microbes, and systemic signaling in plants.

|
Scooped by
Jean-Michel Ané
February 21, 5:59 PM
|
NodABC-produced lipochitooligosaccharides in Frankia: defence suppression in actinorhizal symbioses?

|
Scooped by
Jean-Michel Ané
February 21, 5:36 PM
|
This protocol combines rhizobial spot inoculation with deep-tissue imaging to capture cellular processes during early nodule development in the legume Medicago truncatula. We describe steps to visualize DNA replication activity and cell geometry as indicators of cell proliferation and cell expansion processes. We detail steps for rhizobial spot inoculation of seedlings, incubation in 5-ethynyl-2′-deoxyuridine (EdU) medium, sample fixation and labeling of replicated DNA, clearing, staining of the cell walls, followed by confocal imaging.

|
Scooped by
Jean-Michel Ané
February 19, 6:06 PM
|
In this study, we expanded the understanding of cytokinin perception in legumes by generating and characterizing novel pea cytokinin receptor mutants carrying mutations for the four CHK genes in pea, CHK1, CHK2, CHK3 and CHK4. We constructed single, double, triple and quadruple mutants and analysed their shoot, root, and nodulation phenotypes. We evaluated their contributions to the activation of CK-responsive genes, TCSn promoter activity, and used RNAi knockdowns of CHK1 to explore its role in nodulation. We found key roles for CHK1 in promoting nodulation, CHK3 in delaying leaf senescence and CHK4 in promoting leaf size and axillary shoot branching. Traits such as stem elongation and width as well as shoot and root size were regulated redundantly by the CHK receptors. Overall, this work provides a genetic dissection of cytokinin receptor function in pea, advancing our understanding of hormone signalling in a crop legume and offering genetic insights with potential applications for improving both shoot architecture and symbiotic efficiency.

|
Scooped by
Jean-Michel Ané
February 18, 6:06 PM
|
Sinorhizobium meliloti BacASm and Escherichia coli SbmAEc are closely related proteins that function as homodimeric transporters to import peptides and other cargos through the cytoplasmic membrane into the cytoplasm. BacA is critical for S. meliloti to establish a nitrogen-fixing symbiosis with its legume hosts because of its ability to import Nodule Cysteine-Rich (NCR) plant peptides. This import protects the bacteria inside the nodule from the potentially lethal effects of these NCR peptides while also enabling NCRs to establish intracellular interactions that are essential for symbiosis. Our extensive multidisciplinary studies offer important insights into function of BacA/SbmA transporters and provide a molecular explanation for why BacA/SbmA orthologs from mammalian pathogens can replace BacASm but those from other rhizobia cannot.

|
Scooped by
Jean-Michel Ané
February 18, 5:37 PM
|
Plant-associated microbiomes are vital for sustainable agriculture, enhancing crop nutrient uptake, stress resistance, and yield while reducing agrochemical reliance. Microbial regulation, the targeted manipulation of these microbiomes, bridges microbial ecological potential and agricultural application. Currently, this regulation is categorized by whether microbial genomes are modified, encompassing two core strategies: external conditioning and internal engineering. External conditioning is a genome-unaltered approach that reshapes microbial communities via environmental cues, host traits, and agronomic practices, enabling rapid, low-regulatory tuning of microbiome function. In contrast, internal engineering alters microbial genomes via CRISPR-based tools, synthetic communities, and in situ editing to lock in stable beneficial traits for consistent field performance. Emerging technologies like AI-powered predictive models and advanced delivery systems support both strategies by boosting microbial regulation precision and scalability. Ultimately, microbial regulation optimizes plant–microbe interactions, expediting sustainable agricultural solutions to enhance climate resilience and global food security.

|
Scooped by
Jean-Michel Ané
February 16, 8:03 PM
|
Plants rely on specialized metabolites to defend against herbivores, yet how root-associated symbiotic microbes influence their regulation, coordination, and overall contribution to defense remains widely unknown.
We hypothesized that symbiotic microbes can reprogram specialized metabolic networks, thereby enhancing herbivore resistance, with a particular focus here on polyamine metabolic network. Using the fungal symbiotic endophyte Trichoderma harzianum, tomato, and the herbivore Spodoptera exigua, we combined greenhouse bioassays with molecular, genetic, and metabolomic analyses to investigate how Trichoderma symbiosis reshapes the plant polyamine metabolic network, and affects plant-herbivore interactions.
We found that in the absence of the root symbiosis, herbivory primarily activated polyamine uptake transport and catabolism, reflecting a stress-driven turnover response. Trichoderma symbiosis markedly reconfigured the polyamine network. Symbiosis primed uptake transport and catabolic responses, enhanced polyamine flux through activation of the ornithine decarboxylase pathway, and redirected polyamines into conjugated metabolites with anti-herbivore activity. Genetic analyses confirmed that this metabolic rewiring contributes to Trichoderma-induced resistance to herbivory, linking primary metabolic routes to the accumulation of specialized defense compounds.
Our study highlights root symbionts as key modulators of plant metabolism, showing that specialized metabolite diversity can be shaped by symbiotic interactions.

|
Scooped by
Jean-Michel Ané
February 15, 4:19 PM
|
The seed-to-seedling transition is a key step of plant microbiota assembly, where the seed microbiota meets the soil microbiota. Seed germination triggers profound physiological changes, including exudate release and oxidative bursts, which generate selective pressures shaping microbial survival and interactions. Here, we investigated how host selection and community composition influence bacterial colonization during this transition using a Synthetic Community (SynCom) approach on common bean (Phaseolus vulgaris) seeds. Specifically, we tested the importance of microbial interactions by applying 30 bacterial SynComs spawning a phylogenetic diversity gradient.
Seedling colonization success was primarily determined by strain identity but was strongly modulated by the SynCom context. Strains that were highly effective colonizers when inoculated alone often exhibited reduced transmission within SynComs, indicating that microbial interactions can either inhibit or facilitate colonization. Random forest models confirmed that colonization outcomes could not be predicted from single-strain performance alone. Instead, both phylogenetic relatedness and metabolic similarity among SynCom members emerged as key predictors of strain success, supporting the competition–relatedness hypothesis and highlighting the importance of community-dependent effects in shaping colonization success. Genomic analyses identified microbial traits linked to efficient seedling colonization, notably amino acid transport and metabolism, stress tolerance, ROS detoxification, and biofilm formation, highlighting the strong selective pressures acting during the seed-to-seedling transition. Besides key adaptative traits, our findings underscore the importance of microbial interactions to survive and colonize seedlings, bringing a novel perspective for the successful engineering of seed and seedling microbiota.

|
Scooped by
Jean-Michel Ané
February 12, 8:08 PM
|
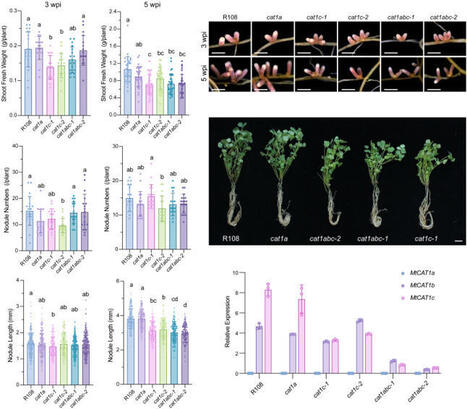
Legumes engage in nitrogen-fixing symbiosis with rhizobia, in which host legumes supply dicarboxylates as a carbon source to rhizobia, while rhizobia reciprocate by providing ammonium to the host plants. Beyond this classical model, accumulating evidence suggests that amino acid exchange is also essential for legume–rhizobium symbiosis. However, it remains unclear whether amino acid transporters are present on the symbiosome membrane (SM) to mediate amino acid exchange during symbiotic nitrogen fixation (SNF). In this study, we identified three amino acid transporters in Medicago truncatula—MtCAT1a, MtCAT1b, and MtCAT1c—which belong to a clade of the plant Cationic Amino acid Transporter (CAT) family known to transport a wide range of amino acids. Notably, MtCAT1b and MtCAT1c are predominantly expressed in infected nodule cells and localize to the SM. Genetic analyses further demonstrate that both MtCAT1b and MtCAT1c are required for amino acid exchange at the SM, with additional evidence indicating that bacteroid metabolism is disturbed in the mutants. Transport assays show that both MtCAT1b and MtCAT1c exhibit broad substrate specificity. Collectively, these findings identify MtCAT1b and MtCAT1c as key mediators of cross-kingdom amino acid exchange, which is essential for maintaining efficient SNF in root nodules.

|
Scooped by
Jean-Michel Ané
February 12, 5:53 PM
|
Symbiotic nutrient exchange between arbuscular mycorrhizal (AM) fungi and their host plants varies widely depending on their physical, chemical, and biological environment. Yet dissecting this context dependency remains challenging because we lack methods for tracking nutrients such as carbon (C) and phosphorus (P). Here, we developed an approach to quantitatively estimate C and P fluxes in the AM symbiosis from comprehensive network morphology quantification, achieved by robotic imaging and machine learning based on roughly 100 million hyphal shape measurements. We found that rates of C transfer from the plant and P transfer from the fungus were, on average, related proportionally to one another. This ratio was nearly invariant across AM fungal strains despite contrasting growth phenotypes but was strongly affected by plant host genotype. Fungal phenotype distributions were bounded by a Pareto front with a shape favoring specialization in an exploration–exploitation trade-off. This means AM fungi can be fast range expanders or fast resource extractors, but not both. Manipulating the C/P exchange rate by swapping the plant host genotype shifted this Pareto front, indicating that the exchange rate constrains possible AM fungal growth strategies. We show by mathematical modeling how AM fungal growth at fixed exchange rate leads to qualitatively different symbiotic outcomes depending on fungal traits and nutrient availability.

|
Scooped by
Jean-Michel Ané
February 12, 5:46 PM
|
Plants establish environmental connections through mycorrhizal symbiosis. These relationships enable them to obtain nutrients and cope with stress while simultaneously exchanging information through subterranean networks. A unified understanding of the molecular mechanisms underlying mycorrhizal interactions that drive adaptation and survival has not yet been achieved, in part because research on them stems from diverse fields of research, such as mycorrhizal ecology and plant epigenetics. This review presents recent studies demonstrating that epigenetic control serves as a central system enabling plants to adapt and maintain stable relationships with mycorrhizal fungi. We begin by describing different types of mycorrhizae. We then analyze mycorrhizal symbiosis by integrating plant and fungal genomic data with molecular evidence on DNA methylation, histone modification, chromatin remodeling, and small RNA pathways. We demonstrate that mycorrhizal symbiosis depends on changing chromatin states, which influence the regulation of the establishment, maintenance, and efficiency of symbiotic connections. They also regulate the balance between nutrient uptake and defense. They may underlie mycorrhizal stress and transgenerational “memory.” We review studies showing that RNA interference between different species enables reorganization of gene expression between plant and fungal cells. Finally, we identify key knowledge gaps and propose future research directions aimed at discovering reliable markers of mycorrhizal responses for epi-breeding and the development of climate-resilient agroecosystems.

|
Scooped by
Jean-Michel Ané
February 11, 4:53 PM
|
In this issue of Cell Host & Microbe, Hussain et al.1 demonstrate surprising microbiome resilience in oak trees under abiotic and biotic stress. In contrast to work on herbaceous plants or saplings, it was found that mature oak trees growing in situ experienced little microbiome change under stress, across both time and tissue type.

|
Scooped by
Jean-Michel Ané
February 10, 2:26 PM
|
Flavonoids, produced by the plant under nutrient stress, are required to initiate the legume-rhizobia symbiosis through the activation of rhizobial nod genes. Notwithstanding the central role of flavonoids in nodulation, their transcriptional regulation remains poorly understood. Here, we show that the nodulation signaling pathway 2 (NSP2) is required for transcriptional activation of flavonoid biosynthesis genes during nodulation in Medicago truncatula. Furthermore, MYB40, a legume-specific MYB transcription factor, is induced by rhizobia in the root epidermis. MYB40 directly binds to flavonoid biosynthetic gene promoters and is required for normal levels of nodulation. Biochemical and genetic evidence reveal that NSP2, not NSP1, interacts with MYB40 during rhizobial infection to strongly upregulate the symbiotic gene chalcone O-methyltransferase 1 in a manner dependent on MYB40 binding sites. Moreover, the overexpression of MYB40 and a microRNA-resistant NSP2 variant enhances nodulation under suboptimal rhizobial availability, suggesting this module fine-tunes symbiosis efficiency. Additionally, flavonoid regulation by NSP2 and MYB40 appears to facilitate arbuscular mycorrhizal colonization under nutrient starvation. Together, our findings establish an NSP2-MYB40 module that integrates symbiotic signaling with metabolic reprogramming, representing an evolutionary innovation for optimizing nitrogen acquisition in dynamic environments.
|

|
Scooped by
Jean-Michel Ané
February 21, 6:01 PM
|
Our study focuses on the metabolomic changes observed during the early stages of Alnus glutinosa and Frankia alni symbiosis. Amino acid metabolic profiling by HPLC-DAD-FLD and untargeted metabolomic analysis of secondary metabolites by UHPLC-QTOF were performed on the shoots and roots of A. glutinosa as well as on bacterial pellets of F. alni. Two culture conditions were compared: a single culture condition (where A. glutinosa or Frankia was grown alone) and a co-culture condition (where A. glutinosa and Frankia were grown together) at different culture times (D1, D2 and D3). Our results reveal a change in metabolism (primary and secondary) in both partners in the co-culture condition. For amino acids, this change was more important in the shoots than in the roots and in Frankia. A total of 16 amino acids (Asp, Asn, Ser, Gln, Gly, Cit, GABA, Ala, Arg, Tyr, Trp Val, Phe, Ile, Lys and Pro) were overproduced in the presence of Frankia in the shoots on the different sampling days. We hypothesised that the plant would modify its amino acid content in its shoots in anticipation of a transfer to Frankia for growth. At the same time, a drastic change in secondary metabolites occurs in the shoots, roots and Frankia at the three time points considered between the control condition and the co-culture condition. Statistical analyses enabled us to highlight the ions characterising the co-culture condition in the different biological compartments (i.e. shoots, roots and Frankia). The biomarkers identified in the shoots and Frankia varied greatly depending on the sampling day (i.e. D1, D2 and D3), revealing strong dynamics. The root biomarkers appear to be more stable over time, as several of them are common to all three sampling days.

|
Scooped by
Jean-Michel Ané
February 21, 5:44 PM
|
Ammonia (NH₃) is a critical chemical in global agriculture and industry with an annual production exceeding 170 million metric tons. Traditionally it is produced via the energy-intensive Haber-Bosch process, which requires high temperatures (400–500 °C), high pressures (150–300 bar), and fossil fuel-derived hydrogen, which accounts for roughly 1–2% of global energy consumption and contributes over 450 million metric tons of CO₂ emissions yearly. In contrast, biological ammonia production offers a sustainable and environmentally friendly alternative by harnessing microbial pathways for nitrogen fixation under ambient conditions. This review examines key microbial pathways for biological ammonia production, including nitrogenase-mediated nitrogen fixation by diazotrophic bacteria and archaea, which typically produce 1–5 mg NH₃ per litre of culture under laboratory conditions. It also highlights advancements in bioelectrochemical approaches, particularly microbial electrochemical systems (MES) that generate ammonia via dissimilatory nitrate reduction to ammonium (DNRA), achieving substantially higher yields of 10–50 mg NH₃–N per litre, corresponding to 0.012–0.061 g NH₃ L−1 day−1. Metabolic and enzymatic mechanisms are analysed, alongside challenges in optimising yields, improving efficiency, and achieving scalability. Integration with renewable energy sources could, optimistically, reduce energy inputs by 50–70% compared to traditional methods, supporting carbon-neutral nitrogen management. The review assesses future research directions, technological advancements, and industrial applications, emphasising synthetic biology and metabolic engineering, which have demonstrated yield increases of 30–50% in laboratory settings for nitrogen fixation systems.

|
Scooped by
Jean-Michel Ané
February 21, 5:32 PM
|
Arbuscular mycorrhizal fungi (AMF) are essential symbionts to most land plants. There is great interest in using AMF inoculation treatments to enhance restoration or agricultural efforts. However, little is known about AMF asymbiotic interactions that could shape soil ecosystems and inoculation outcomes. Studying these early traits is challenging given the obligate nature of AMF. Microfluidic devices enable high-resolution, single-plane, time-resolved observation of AMF hyphal traits in controlled, plant-less environments. The AMF-AnastomosisChip, introduced in this work, additionally separates spores and hyphae into lanes for detailed observations prior to entering a shared interaction zone. Using this device with Rhizophagus irregularis, we phenotyped key hyphal traits from germination to anastomosis. We compared the effect of a fatty acid (FA) treatment to other environmental factors in shaping AMF growth and fusion (anastomosis). Among our results, we confirm known effects of the FA treatment in increasing hyphal branching and longevity but interestingly show that multiple hyphae in a lane can negate FA effects. We also newly reveal that anastomosis occurs for 100% of interactions among hyphae from different lanes, but only for 50% of in-lane interactions, potentially linked to spore origin. Our findings validate the AMF-AnastomosisChip as a versatile platform for increasing discovery in trait-based microbial ecology, including for asymbiotic AMF interactions with environmental factors. This work sets the foundation for future studies with the AMF-AnastomosisChip on the effects of nutrient content, plant-derived molecules, or microbial community members on the AMF traits characterized in this study.

|
Scooped by
Jean-Michel Ané
February 19, 5:57 PM
|
WRI5a functions as a master regulator of symbiotic molecule exchange during mycorrhizal infection. In Medicago truncatula, MtWRI5a binds to AW-box motifs to directly activate key symbiotic genes (MtABCB1, MtSTR, and MtPT4), thereby facilitating the bidirectional exchange of lipids (C), phosphate (Pi), and IAA between the plant and the fungus. The figure was adapted from Wang et al. (2025)

|
Scooped by
Jean-Michel Ané
February 18, 5:38 PM
|
Arbuscular mycorrhizal fungi (AMF) and rhizobia are crucial for plant growth, acting as key components of ecosystem sustainability. However, it remains unclear how these two plant symbionts interact in legumes in response to the addition of nitrogen (N) or phosphorus (P) fertilizers to the soil. We conducted a global meta-analysis with 1644 independent observations to investigate this issue. The results indicated that inoculation with AMF and/or rhizobia significantly enhanced N and P uptake, biomass and yield in legumes. In contrast, their co-inoculation notably enhanced nitrogenase activity, soil total N content, and organic matter content without altering soil pH. Co-inoculation generally exerted an additive effect on legume growth and yield, although perennial trees derived synergistic benefits. Specifically, perennial leguminous trees demonstrated synergistic growth benefits under co-inoculation conditions, achieving biomass yields exceeding the sum of single symbiont effects; however, annual leguminous crops and forages displayed additive effects of co-inoculation. Critically, N and P fertilization disrupted these symbioses when application rates exceeded certain thresholds. Sole N addition inhibited rhizobial nodulation, with plant growth-promoting effects shifting from positive to negative beyond approximately 90 kg N ha⁻¹ . P addition reduced AMF colonization, with yield benefits diminishing above 58 mg P kg⁻¹ . These results reveal distinct threshold-dependent pathways through which soil nutrient availability interferes with legume-microbe mutualisms. Our study provides the global evidence of AMF-rhizobia synergy for legume productivity and soil C-N cycling, coupled with mechanistic insights showing how N and P fertilizations differentially undermine these partnerships. The results advocate for optimized nutrient management to harness microbial symbioses in sustainable agriculture.

|
Scooped by
Jean-Michel Ané
February 18, 1:24 PM
|
Root exudates are thought to play an important role in plant-microbial interactions. In return for nutrition, soil bacteria can increase the bioavailability of soil nutrients. However, root exudates typically decrease in situations such as drought, calling into question the efficacy of solvation and bacteria-dependent mineral uptake in such stress. Here, we tested the hypothesis of exudate-driven microbial priming on Cupressus saplings grown in forest soil in custom-made rhizotron boxes. A 1-month imposed drought and concomitant inoculations with a mix of Bacillus subtilis and Pseudomonas stutzeri, bacteria species isolated from the forest soil, were applied using factorial design. Direct bacteria counts and visualization by confocal microscopy showed that both bacteria associated with Cupressus roots. Interestingly, root exudation rates increased 2.3-fold with bacteria under drought, as well as irrigation. Forty-four metabolites in exudates were significantly different in concentration between irrigated and drought trees, including phenolic acid compounds and quinate. When adding these metabolites as carbon and nitrogen sources to bacterial cultures of both bacterial species, eight of nine metabolites stimulated bacterial growth. Importantly, soil phosphorous bioavailability was maintained only in inoculated trees, mitigating drought-induced decrease in leaf phosphorus and iron. Our observations of increased root exudation rate when drought and inoculation regimes were combined support the idea of root recruitment of beneficial bacteria, especially under water stress.

|
Scooped by
Jean-Michel Ané
February 16, 8:01 PM
|
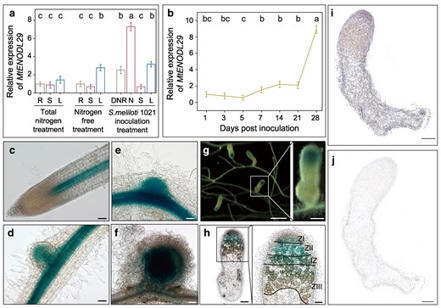
The early nodulin-like (ENODL) subfamily, part of the phytocyanin, arabinogalactan protein, and nodulin-like families, is involved in plant growth and stress resistance. However, its role in symbiotic nodulation remains poorly understood. In barrel medic (Medicago truncatula), we found MtENODL29 was strongly activated at the late stages of nodule development, particularly in the infection zone of nodules. Both RNA interference (RNAi) and mutation of MtENODL29 caused a considerable reduction in nodule numbers, an increase in cysteine protease activity, a dramatic decrease in leghemoglobin content, and signs of premature senescence in nodule cells, suggesting that disruption of MtENODL29 accelerates nodule aging. Transcriptome analysis of 7-dpi (day post inoculation) inoculated roots and 28-dpi nodules in enodl29 mutants showed significant downregulation of symbiotic genes, accompanied by differential expression of genes associated with lipid metabolism and transport. MtENODL29 mutation also negatively impacted plant growth and development. MtENODL29 bound to MtnsLTP (nonspecific lipid transfer protein), MtKCR (very-long-chain 3-oxoacyl-CoA reductase), and MtSec61γ (gamma subunit of the translocase complex Sec61) through its ALR (arabinogalactan protein-like region) domain. MtENODL29 co-localized with these proteins in the plasma membrane and endoplasmic reticulum. Notably, MtnsLTP showed high expression in the nodules, similar to MtENODL29, while MtKCR and MtSec61γ were also highly expressed in the leaves and stems. These results suggest that MtENODL29 participates in membrane lipid modification and transport by interacting with MtnsLTP, MtKCR, and MtSec61γ, facilitating the formation of symbiosome membranes as alfalfa rhizobium (Sinorhizobium meliloti) strain 1021 are released into nodule cells. Moreover, MtENODL29 influences plant growth, highlighting its role in coordinating plant development and symbiosis.

|
Scooped by
Jean-Michel Ané
February 12, 8:12 PM
|
Molybdenum (Mo) nitrogenase is a two-component enzyme complex that catalyzes the reduction of dinitrogen to ammonia and protons to hydrogen gas. We have shown that electrons for dinitrogen reduction can be delivered photochemically to the catalytic MoFe protein component by cadmium sulfide (CdS) nanocrystals. In this study, we used electron paramagnetic resonance spectroscopy to measure the transient populations of catalytic intermediates. We fit the populations with a pre-steady-state kinetic model, which allowed us to distinguish between productive and non-productive reaction pathways and extract the rate constants for the reaction. Our results demonstrated that the rate of catalytic electron delivery into MoFe protein increased with the concentration of the sacrificial electron donor. This enabled electron delivery to exceed the rate of hydride protonation, a relaxation pathway that competes with N2 binding. Thus, managing the balance between electron transfer and hole transfer reactions is required to achieve a kinetic regime that favors N2 reduction.

|
Scooped by
Jean-Michel Ané
February 12, 5:57 PM
|
Rhizobial type-Ⅲ effectors (T3Es) contribute to establishing symbiotic interactions with legume host plants, alongside Nod factors. However, the functions of most rhizobial T3Es, as well as the regulatory and molecular mechanisms underlying their symbiotic effects on hosts, particularly in soybean, are poorly documented. Here, we characterize the function of the T3E Nodulation Outer Protein C (NopC) in the broad-host-range rhizobium Sinorhizobium fredii HH103 for promoting symbiosis in soybean. NopC genotype influences root nodulation across diverse host germplasm and this is further influenced by GmRAC1, encoding a ROP/RAC family GTPase in soybean. GmRAC1 physically interacts with NopC to subsequently induce the expression of the essential symbiotic genes GmNIN2a/2b and GmENOD40. Knock-down of GmNIN2a/2b results in NopC failing to promote symbiosis, and Gmrac1 mutants have fewer nodules than the wild type. NopC facilitates multiple infection stages whereas the requirement for GmRAC1 is pronounced for infection-thread progression and nodule-primordia initiation. Natural variation in the GmRAC1 promoter largely dictates the symbiotic contribution of NopC during symbiotic establishment, and elite GmRAC1 haplotypes with strong expression were artificially selected in soybean breeding. Transgenic over-expression level and elite GmRAC1 haplotypes increase plant height, 100-seed weight and soybean yield. GmRAC1 serves as a key regulator of NopC-mediated symbiosis promotion and offers translational potential for enhanced symbiotic nitrogen fixation in molecular breeding of soybean.

|
Scooped by
Jean-Michel Ané
February 12, 5:48 PM
|
In the mutualism between leguminous plants and rhizobia bacteria, rhizobia live inside root nodules, creating potential for host genes to shape the rhizobial selective environment. Many host genes that affect symbiosis have been identified; however, the extent to which these genes affect selection acting on rhizobia is unknown. In this study, we inoculated 18 Medicago truncatula symbiotic mutants (including mutants that alter Nodule Cysteine-Rich (NCR) peptide production, plant defence, and nodule number regulation) with a mixture of 86 Sinorhizobium meliloti strains. Most mutations resulted in reduced host benefits, but the effects on rhizobial benefit (i.e., relative strain fitness) varied widely, revealing widespread host-by-strain fitness interactions. Genome-wide association analyses identified variants on rhizobial replicons pSymA and pSymB as important in mediating strain fitness responses to host mutations. Whereas most top variants affected rhizobial fitness with one host mutation (limited effect variants), nine affected fitness across six or more host mutations. These pervasive variants occurred primarily on pSymA, the symbiotic replicon, and include fixL and some metabolic genes. In contrast to the limited effect variants, variants with pervasive positive effects on strain fitness when host genes were mutated tended to adversely affect fitness in wild-type hosts. Competition assays across Medicago genotypes confirmed a pervasive role for one candidate (malonyl-CoA synthase), and AlphaFold multimer modelling suggests that many rhizobial top candidates could interact with host NCR peptides. Our results reveal how host genetic mutations alter strain fitness, setting the stage for improving rhizobial inoculants and breeding legume hosts better adapted to multi-strain environments.

|
Scooped by
Jean-Michel Ané
February 12, 5:32 PM
|
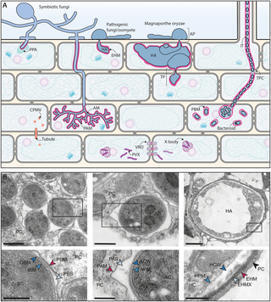
Plants encounter a myriad of microorganisms that can invade with detrimental or beneficial outcomes. Taking a membrane-centric point of view, we discuss the cellular events underlying microbe-induced cell signaling, the navigation of microbes within plant tissues, and the molecular exchanges occurring at plant–microbe interfaces. We discuss the implications of individual membrane lipids and the emerging role of membrane biophysics in non-self and modified-self sensing. We highlight the common themes underlying the active role of membranes during plant interactions with viruses, bacteria, oomycetes, and fungi, and define exciting directions for future research.

|
Scooped by
Jean-Michel Ané
February 11, 4:53 PM
|
Global forest biomes face increasing stressors and disease outbreaks that threaten ecosystem health. Tree-associated microbiota are vital for tree resilience, yet their responses to biotic and abiotic stressors in mature trees remain poorly understood. Using an experimental woodland plot of 144 Quercus petraea trees subjected to drought (rain exclusion), nutrient stress (ringbarking), and biotic treatments (bacterial pathogens and beetle larvae) to simulate acute oak decline, we tracked microbial communities in leaf, stem, and root/rhizosphere tissues across four time points over 2 years. Oak trees hosted distinct microbial communities across tissue types, which remained largely stable under stress. Rain exclusion significantly altered microbiota composition, though these changes explained less than 1% of total variance. Actinobacteriota, linked to drought tolerance, increased in the root/rhizosphere of rain-excluded trees. These findings reveal a surprising resilience of oak-associated microbial communities to environmental and biotic disturbances, highlighting their potential role in forest ecosystem stability.
|






 Your new post is loading...
Your new post is loading...