Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 1:51 AM
|
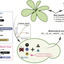
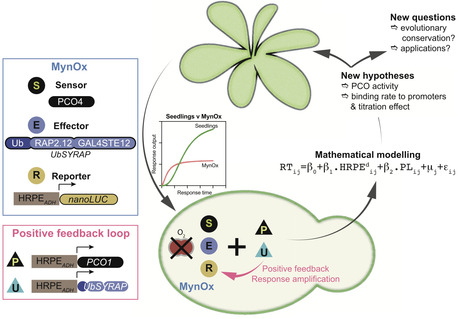
Plants face hypoxic conditions either chronically, as particular tissues are characterized by fluctuating or stable low oxygen levels, or acutely, when flooded. In vascular plants, transcriptional adaptive responses to hypoxia are rapidly mounted by Ethylene Response Factors VII (ERFVIIs), regulated by Plant Cysteine Oxidases (PCOs) through the cysteine branch of the N-degron pathway (Cys-NDP) for oxygen sensing. However, this relatively simple regulatory circuit, consisting of both constitutively expressed as well as hypoxia-inducible ERFVIIs and PCOs, interacts with diverse signaling cues and pathways invoked by hypoxia. To understand the share of the PCO-mediated oxygen sensing mechanism in the production of hypoxia responses, we insulated the PCO/ERFVII circuit from Arabidopsis thaliana and adapted it to Saccharomyces cerevisiae. Using a reporter gene to monitor the output of the circuit allowed us to compare the speed and amplitude of response to hypoxia in the engineered yeast and the source organism. Hypoxia triggered ERFVII stabilization both in Arabidopsis and yeast, leading to a similarly fast transcriptional response that was however larger in plants. A simple hypoxia-inducible feedback loop improved the amplitude of response in yeast, demonstrating the importance of this regulation in the endogenous PCO/ERFVII circuit. Finally, computational modeling of the yeast circuit enabled us to identify promoter competition and presence of hypoxia-inducible PCOs as key parameters that shape early hypoxia responses in plant cells.

|
Scooped by
mhryu@live.com
Today, 1:39 AM
|
Abiotic stress is a major problem which threatens agricultural productivity and global food security. Drought, extreme temperatures, salinity, and heavy metal contaminations cause disturbances in plant cellular homeostasis, impair metabolic processes, and reduce crop yields. Plants possess innate defense mechanisms against abiotic stresses by expressing stress-responsive genes, accumulating osmo-protectants, and the activation of antioxidant enzymes. Failure of these defense mechanisms under prolonged stress conditions leads to homeostatic imbalances. Recently, nanotechnological approaches like nanoparticle-mediated delivery systems have been developed to enhance the plants' resilience against stress conditions. Engineered nanoparticles (ENPs) have unique physicochemical properties such as a high surface area, high reactivity, and a tunable surface chemistry. These properties enable the nanoparticles to interact with plant systems at molecular and cellular levels, modulate stress signaling pathways, trigger the upregulation of stress-responsive genes, and reduce oxidative stress by increasing antioxidant enzymes' activity. However, under certain environmental conditions, ENPs may also induce oxidative damages and exhibit phytotoxicity. This review encompasses a comprehensive overview on the role of nanoparticles in abiotic stress management, along with detailed insights into the biosafety and environmental toxicity of ENPs. Overall, the review highlights the novel insights of ENPs-plant interactions and identifies existing knowledge gaps through a systematic literature review to guide future research towards sustainable agriculture.

|
Scooped by
mhryu@live.com
Today, 1:22 AM
|
As ubiquitous features of every natural environment, microbes have profoundly shaped eukaryotic biology throughout evolution. Circadian clocks evolved in all domains of life as central regulators that align physiology with environmental cycles, yet whether they respond directly to microbial signals remains unknown. Here, we demonstrate that evolutionarily diverse microbes potently reset mammalian cellular clocks and can drive phase shifts in plants and algae, indicating cross-kingdom effects of microbes on circadian rhythms. In mammals, exposure to soluble bacterial components distinct from canonical innate immune ligands induces acute PER2 upregulation independently of Bmal1 or nascent transcription. A targeted inhibitor screen and biochemical assays implicate p38 MAPK as a modulator of this response. Taken together, this positions bacterial exposure as a previously unrecognized circadian clock input, revealing a new axis of host-microbe interaction that modulates biological timing at the cellular level.

|
Scooped by
mhryu@live.com
Today, 12:55 AM
|
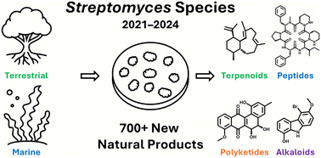
Streptomyces species have long been recognized as prolific sources of novel natural products, generating a wide array of clinically-relevant antimicrobial and cytotoxic compounds. In this review, we examine 174 publications from 2021 to 2024 that describe 716 previously unreported secondary metabolites isolated from Streptomyces strains sourced from a range of marine and terrestrial environments. Natural product researchers used a variety of chemical and biological techniques, including genome mining, molecular networking, and heterologous expression, to uncover novel compounds with a diverse array of structures, including cyclic and linear peptides, macrocyclic, polyaromatic, and other polyketides, terpenoids, alkaloids, and azoxy compounds, among others. The variety of bioactivities exhibited by these secondary metabolites, including antibacterial, anticancer, antifungal, antiviral, and anti-inflammatory effects, illustrates the enduring potential of the Streptomyces genus to produce promising new bioactive natural products.

|
Scooped by
mhryu@live.com
Today, 12:27 AM
|
The Gram-negative outer membrane (OM) is an asymmetric bilayer that protects cells from environmental stress and antibiotics. This asymmetry, with lipopolysaccharide (LPS) in the outer leaflet and glycerophospholipids (GPLs) in the inner leaflet, requires coordinated synthesis of both lipid classes. The committed step of LPS biosynthesis is catalyzed by LpxC, a prime antibiotic target. Here, we show that lysophospholipids (LPLs), considered byproducts of membrane turnover, act as signaling molecules restoring OM homeostasis when LPS synthesis is limited. In the presence of the LpxC inhibitor PF-5081090 (PF), loss of the LPL recycling system increased growth, suppressed envelope stress responses, improved OM asymmetry, and lowered GPL levels to maintain GPL-to-LPS balance. This recycling system includes the transporter LplT, which moves LPLs across the inner membrane, and the acyltransferase/acyl-ACP synthetase (Aas), which acylates them to regenerate GPLs. These protective effects required the OM phospholipase PldA that degrades mislocalized GPLs into LPLs and free fatty acids. Although previous work showed that PldA-generated fatty acids stabilize LpxC and promote LPS synthesis, our findings reveal a complementary role for LPLs in signaling reduced GPL synthesis when LPS is limiting. Genetic and chemical manipulation of fatty-acid flux altered PF resistance, confirming that decreased GPLs drives protection. The two PldA-derived signals, fatty acids that promote LPS synthesis and LPLs that suppress GPL synthesis, likely operate under different metabolic conditions to interpret membrane stress and restore OM balance. This lipid-feedback mechanism establishes the first signaling function for bacterial LPLs and reveals a new layer of regulation in envelope homeostasis.

|
Scooped by
mhryu@live.com
April 14, 4:39 PM
|
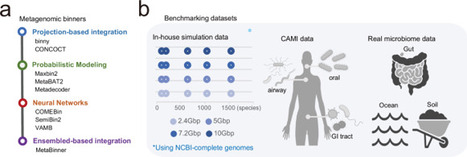
Metagenomic binning is essential for reconstructing prokaryotic genomes from metagenomic samples. We benchmarked various binning tools using Critical Assessment of Metagenome Interpretation (CAMI)-simulated, custom-simulated, and real metagenomic datasets, primarily focusing on short-read sequencing data. Our analysis highlights critical factors influencing binning efficacy: (i) Sequencing depth and taxonomic complexity strongly impact binning performance, while CAMI-simulated benchmarking datasets exhibit substantially lower complexity than human gut and environmental metagenomes, (ii) Chimeric genome rates vary widely across tools, (iii) Multi-sample binning is most effective with about 20 samples, as using too few or too many samples can reduce its benefits, and (iv) Binning efficacy was lower for single-end sequencing samples due to reduced contig quality and assembly fragmentation. Neural network-based tools consistently outperformed others in genome recovery from both real samples and simulated samples with realistic taxonomic complexity, though at higher computational cost. By integrating and refining genome bins from the top three binning tools, we recovered >30% more high-quality genomes than previous methods. This study provides practical guidance for improving metagenomic binning to facilitate the reconstruction of prokaryotic genomes. Benchmarking metagenomic binning tools with simulated and real datasets reveals factors affecting genome recovery and provides practical guidelines for improving metagenome assembled genome reconstruction.

|
Scooped by
mhryu@live.com
April 14, 4:25 PM
|
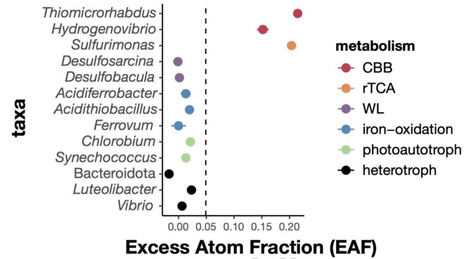
Aquatic environments absorb ~2.5 gigatonnes of atmospheric carbon each year, more than the carbon stored in the atmosphere, soils, and all biomass combined. Primary producers transform this dissolved inorganic carbon into biomass that can subsequently flow into other trophic levels, or be released back into the environment through viral lysis. While there is substantial knowledge about the diversity and activity of viruses infecting photoautotrophic primary producers and the ecosystem impact, little is known about viruses infecting chemoautotrophs, representing a gap in our understanding of key processes driving microbial carbon cycling. Here, we combine metagenomics with quantitative 12/13C stable isotopic probing (qSIP) mesocosm experiments in a marine-derived meromictic pond to quantify population-specific isotopic enrichment, identify key chemoautotrophic primary producers, and virus-host dynamics. Isotopically enriched carbon is tracked from the genomes of chemoautotrophs to putative viruses, showing that active populations of hydrogen/sulfur-oxidizing chemoautotrophs (Thiomicrorhabdus, Hydrogenovibrio, Sulfurimonas, Sulfurovum) are targeted by viruses. This work provides the foundation for revealing the diversity and activity of viruses infecting globally-widespread chemoautotrophs. Our study sheds light on trophic interactions that impact microbial carbon cycling in aphotic environments and builds toward biogeochemical models that incorporate viral impacts on chemoautotrophic microbial communities. In this study, Luo and colleagues identify previously unknown viruses that actively infect highly productive chemoautotrophs. These findings provide new insights into key trophic interactions and virus-host dynamics that impact microbial carbon cycling in aphotic environments.

|
Scooped by
mhryu@live.com
April 14, 4:02 PM
|
Lignocellulosic biomass (LCB) is a plentiful resource, and its effective utilization is essential for mitigating resource scarcity. Xylose, the second most abundant sugar in LCB after glucose, is present in significant quantities. Nevertheless, its current utilization is markedly lower than that of glucose. Although microbial conversion of LCB into high-value products is promising, inefficient xylose metabolism remains a bottleneck. Advances in metabolic engineering and synthetic biology techniques offer powerful tools to improve microbial xylose utilization. In this review, we summarize recent research progress in microbial xylose metabolism, emphasizing xylose metabolic pathways, metabolic engineering strategies, and the production of high-value chemicals derived from xylose. We also discuss future opportunities to overcome key challenges, including efficient xylose extraction from LCB, coutilization of glucose and xylose, and enzyme and pathway optimization. These insights aim to support the development of more efficient bioconversion processes for xylose and contribute to the broader utilization of lignocellulosic feedstocks.

|
Scooped by
mhryu@live.com
April 14, 3:48 PM
|
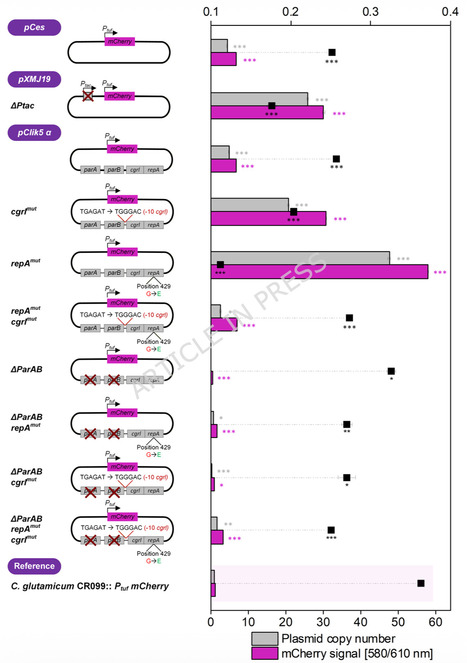
We rationally redesigned the replication control region of the widely used pClik 5α (pCG1-family) backbone by introducing targeted mutations in the repA gene, an antisense RNA (cgrI) promoter, and putative partitioning genes parAB, and constructed a systematic panel of high-copy variants. Using a Ptuf-driven mCherry reporter as a quantitative readout, we identified plasmids that supported several-fold higher fluorescence than the parental backbone while maintaining robust growth. Fluorescence-based gene-dosage estimation indicated a strong increase in apparent plasmid copy number. Independent qPCR-based plasmid copy number determination using two plasmid loci confirmed that the lead variant pClik 5α repAmut reached approximately 28–30 copies per chromosome equivalent, compared to approximately 2–3 copies for the parental plasmid, corresponding to an approximately 10-fold increase. Genome-wide transcriptome analysis revealed a defined and adaptive transcriptional response to elevated plasmid copy number and expression burden, characterized by adjustments in membrane-associated transport, respiratory functions, and amino acid-related metabolism, without evidence of collapse of core biosynthetic functions. When the best-performing replicon was applied to episomal expression of a codon-optimized pedACDCgl operon, pediocin PA-1 titers increased by 2.5-fold compared to the best pXMJ19-based reference under identical, previously optimized process conditions, placing the system, under comparable cultivation formats, within the upper range of reported microbial pediocin production processes. This work demonstrates that rational engineering of pCG1-family replication modules in C. glutamicum can unlock markedly higher plasmid copy numbers and expression capacities while preserving physiological robustness. The resulting high-copy pClik 5α derivatives, exemplified by pClik 5α repAmut, provide a versatile high-copy expression platform with demonstrated utility for recombinant reporter protein and antimicrobial peptide production in C. glutamicum and offer a foundation for further integration with folding, secretion, and process engineering strategies to advance industrial AMP production.

|
Scooped by
mhryu@live.com
April 14, 3:39 PM
|
Anthropogenic activity, driven by industrialization, agricultural practices, and waste disposal, has emerged as a predominant contributing factor to environmental pollution. These activities release substantial amounts of toxic pollutants into the environment, such as heavy metals, organic pollutants, microplastics, and nanomaterials, adversely affecting various ecosystems. These toxic substances can exert considerable stress on various microorganisms, including bacteria, fungi, and microalgae. The impact of anthropogenic pollutants on microorganisms is a nascent area of study, particularly as environmental stressors continue to increase in both quantity and complexity. This review aims to enhance our understanding of how microorganisms (bacteria, microalgae, and fungi) respond to the anthropogenic pollutants including heavy metals, organic pollutants such as polycyclic aromatic hydrocarbons (PAHs), nanomaterials and microplastics. It explores the toxic effects of these pollutants on diverse microbial species. Furthermore, the review covers studies that examine the molecular mechanisms underlying microbial resistance both through natural resistance processes and adaptive laboratory evolution or evolutionary engineering strategies. The review also highlights how omics technologies such as genomics, transcriptomics, proteomics and metabolomics reveal conserved and unique molecular mechanisms to gain insight into the pollutant-specific and organism-specific adaptation strategies. Nevertheless, limitations in community-level multi-omics studies, the relatively limited data on fungi, and the challenges associated with studying mixed cultures hinder a comprehensive understanding of microbial response and resistance mechanisms to anthropogenic pollutants. Addressing these gaps will be pivotal in leveraging the molecular mechanisms to guide the development of novel strategies to obtain pollutant-tolerant strains for bioremediation, bio-monitoring, and synthetic biology applications.

|
Scooped by
mhryu@live.com
April 14, 1:50 PM
|
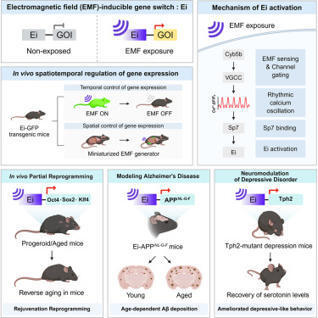
Gaining precise control of gene expression is crucial in biomedical applications. However, spatiotemporal precision remains challenging. Here, we present a remotely controlled in vivo gene switch responsive to electromagnetic fields (EMFs) that enables precise spatiotemporal activation of target genes. We uncovered the EMF-inducible gene switch activation mechanism via a CRISPR-Cas9 screen, identifying cytochrome b5 type B (Cyb5b) as an essential mediator likely acting as an EMF sensor. The EMF-inducible gene switch was activated by rhythmic oscillatory calcium dynamics rather than generic calcium influx, defining a precisely tuned and bio-orthogonal induction mechanism. Functionally, EMF activation of the Oct4-Sox2-Klf4 (OSK) cassette induced in vivo partial reprogramming in aged mice, conditional expression of human mutant amyloid precursor protein (APP) for Alzheimer’s disease (AD) modeling recapitulated pathological features, and EMF-mediated Tph2 expression restored serotonergic activity and ameliorated depressive-like behaviors in Tph2-mutant depression mice. Overall, a remotely controlled EMF-inducible gene switch represents a versatile and effective biomedical platform.

|
Scooped by
mhryu@live.com
April 14, 1:30 AM
|
Ancestral state reconstruction (ASR) is a foundational tool in comparative biology, offering insights into the evolutionary history of lineages. With each new evolutionary model, our ability to estimate ancestral states with increased biological realism has improved. However, the field has primarily relied on marginal reconstructions, which focus on individual nodes. This framework is analytically tractable and appropriate for node-specific hypotheses, but it is not designed to identify the most probable sequence of evolutionary events across a tree. We argue that for researchers interested in evolutionary trajectories, joint reconstructions provide a more effective way to characterize the full history of transitions. Traditionally, joint reconstruction algorithms focused only on the single most likely sequence, but here we use conditional probabilities derived from stochastic mapping to sample the distribution of plausible ancestral histories efficiently. Furthermore, we provide tools to quantify and summarize this joint uncertainty. Through simulations and an empirical case study, we demonstrate that joint reconstructions more effectively recover simulated trait histories than node-wise marginal estimates and that the uncertainty surrounding these histories can be biologically meaningful. We apply our methods to epidemic multidrug-resistant Klebsiella pneumoniae and find that the evolution of antibiotic resistance is not a single narrative but a series of competing histories. Each of these histories exhibits distinct phenotype–genotype transitions that node-wise approaches would struggle to identify, yet have critical implications for predicting and understanding resistance evolution.

|
Scooped by
mhryu@live.com
April 14, 1:08 AM
|
The gut microbiome governs aspects of human growth and development. While human milk's primary purpose is metabolism, it also provides nonnutritious biologics and macromolecules. This mixture includes the human milk oligosaccharides (HMOs), which are indigestible and survive the low pH of the stomach and small intestine, reaching the large intestine intact. Here, HMOs serve as prebiotics for beneficial bacteria, providing a competitive growth advantage over potential pathogens. Upon metabolizing HMOs, commensals generate short-chain fatty acids and metabolites that enhance the gut community. Therefore, HMOs work to develop and sustain the gut microbial community as a living therapeutic that prevents illness from potential microbial pathogens and modulates development of the infant gut. The goal of this targeted review is to characterize the roles HMOs play in governing bacterial and viral members of the infant gut microbiome, describing how HMOs both define a healthy microbiota and prevent microbial dysbiosis.
|

|
Scooped by
mhryu@live.com
Today, 1:43 AM
|
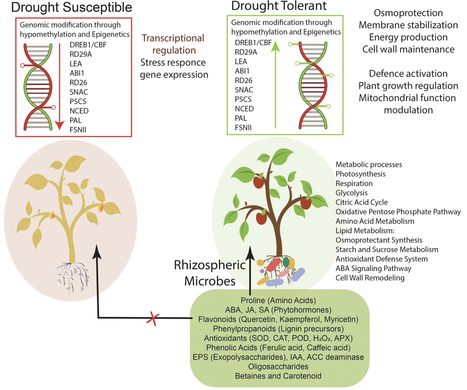
The frequent occurrence of drought, salinity and heat disasters due to global climate change has become a problem that cannot be ignored and seriously restricts food security and sustainable agricultural development. The role of rhizobacteria in the response of plants to abiotic stress has an important guiding significance in improving plant growth. This review summarizes the response of plant rhizosphere microbial communities to abiotic stress, analyzes the mechanism by which rhizosphere-related bacteria assist plants to resist abiotic stress, and expounds on the interaction between soil physical and chemical properties, the plant root metabolome, and the rhizosphere microbiome under abiotic stress. This review systematically summarizes the core roles and mechanisms of rhizobacteria in plants' defense against abiotic stress. Stress reshapes the rhizosphere microecology, with drought enriching Firmicutes and Actinobacteria, salt stress increasing Bacteroidetes abundance, and heat stress expanding the dominance of thermotolerant bacteria. Microbial diversity and network structure undergo adaptive reorganization. Streptomyces and Bacillus, as the twin stars aiding plants in enhancing stress resistance, provide medium- to long-term protection through rich secondary metabolites and mycelial networks, while Bacillus achieves acute responses via rapid spore germination, signal induction, and nutrient competition. Rhizobacteria improve soil nutrient availability by regulating carbon, nitrogen, and phosphorus cycles, secreting organic acids and enzymes, and induce plant osmotic adjustment, antioxidant, and anti-ethylene signaling networks through extracellular polysaccharides, volatile organic compounds, plant hormones, and 1-aminocyclopropane-1-carboxylic acid (ACC) deaminase pathways, thereby systematically enhancing the host's water use efficiency and membrane stability. Future research should integrate multi-omics and field validation to precisely construct rhizosphere bacterial communities, providing theoretical basis and technical routes for green agriculture.

|
Scooped by
mhryu@live.com
Today, 1:34 AM
|
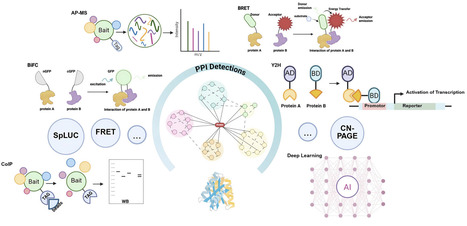
Protein–Protein interactions (PPI) wire plant cells, assembling metabolons, routing signals, and coordinating organelle crosstalk. We review experimental platforms and the computational signals long used to predict PPIs. While experimental platforms and traditional computational approaches have long been employed for PPIs prediction, recent advances in artificial intelligence offer unprecedented opportunities to map plant interactomes comprehensively. To provide a systematic overview, we categorize current methodologies into four thematic families: (i) sequence-centric predictors utilizing protein language models to extract evolutionary features; (ii) structure-based predictors integrating coevolutionary signals to reconstruct 3D complex arrangements; (iii) network-level learners employing graph architectures to capture global interactome topology; and (iv) geometric and generative methods leveraging symmetry-aware networks for specific site identification and de novo design. Despite rapid gains, plant applications are constrained by paralog expansion, compartmentalization, dynamic microenvironments, and the sparse availability of gold standards in the field. Next-generation plant AI PPI models should be organelle-aware, multimodal, rigorously benchmarked, structure-gated, and condition-validated.

|
Scooped by
mhryu@live.com
Today, 1:06 AM
|
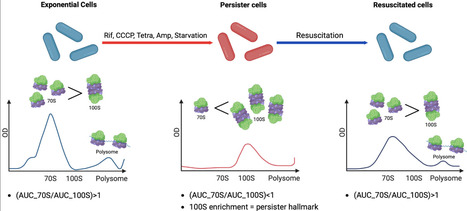
Persister cells survive antibiotic exposure and contribute to infection relapse, yet the molecular features that distinguish them from actively growing cells remain incompletely defined. Here, we used sucrose gradient-based ribosome sedimentation profiling to characterize ribosome complex distributions in E. coli persister cells and monitored their dynamics during resuscitation. Rifampicin-induced persister cells were characterised by pronounced enrichment of translationally inactive 90–100S ribosome complexes and a concomitant reduction in 70S ribosomes relative to exponentially growing cells. Upon nutrient replenishment, ribosome distributions progressively shifted toward higher 70S and polysome (complexes of multiple ribosomes simultaneously translating a single mRNA) levels, coinciding with growth recovery, indicating that resuscitation involves gradual remodelling of ribosome states rather than abrupt restoration of active translation. Functional analysis of ribosome-associated factors demonstrated that RMF, HPF and RaiA promote 100S ribosome accumulation and enhance persister formation, whereas deletion of rmf severely impaired both 100S formation and persistence. In contrast, loss of HflX did not measurably affect persister formation, consistent with a role downstream of persister establishment. In multiple stress-induced persister models including rifampicin, tetracycline, CCCP and starvation, as well as in a clinically relevant E. coli O157:H7 (EHEC) strain, ribosome distributions consistently exhibited a quantitative reversal of the AUC_70S/AUC_100S ratio (Ratio < 1.0) relative to exponentially growing cells (Ratio > 1.0). Collectively, these findings demonstrate that this shift in the 70S-to-100S balance is a consistent and shared feature of E. coli persister physiology and that ribosome state distributions link persister formation to resuscitation dynamics. These findings provide a quantitative ribosome-state framework that may inform the development of anti-persistence strategies targeting ribosome hibernation factors.

|
Scooped by
mhryu@live.com
Today, 12:32 AM
|
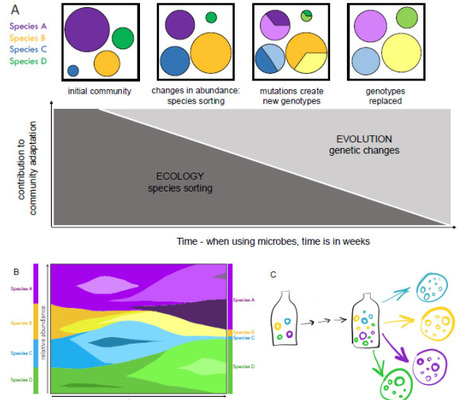
Experimental approaches with a defined microbial community extend concepts from classical experimental evolution of single species to study the complexities of community level changes. Successfully assembling a microbial community, with traceable members, in a defined and manipulatable environment, would provide an effective opportunity to experimentally study general processes and principles in evolutionary ecology. The dynamics between species sorting and genotypic changes is a core, yet not well understood, topic across biological researc. The use of a synthetic microbial community inspired on natural communities reproduced in laboratory environments (for example, fermented foods), enables for experimental evolution of diverse communities. Appreciating the capabilities of communities to respond to changing environments and consequent selection pressures beyond genetic changes within single species must be understood in the face of rapid global environmental changes. Investigating the dynamics between intra- and interspecies processes is significant for understanding community responses to change in natural and managed systems across short and longer temporal scales

|
Scooped by
mhryu@live.com
April 14, 11:52 PM
|
Cell-free protein synthesis (CFPS) is an in vitro platform that enables rapid protein production using cell extracts, energy sources, and genetic templates. Owing to its fast response, elimination of cell culture, open reaction environment, lyophilization compatibility, and high programmability, CFPS has emerged as a versatile engine for diagnostic sensing. Recent advances have integrated CFPS with modular genetic circuits, CRISPR-based detection, isothermal amplification, and portable formats such as paper-based devices and microfluidic chips, enabling sensitive and specific detection of viral nucleic acids, pathogen antigens, and small-molecule targets. These platforms further support multiplexed and point-of-care testing, substantially reducing assay time, cost, and infrastructure requirements. Despite remaining challenges in biosensor design for novel targets, analytical sensitivity in complex samples, batch-to-batch reproducibility, and clinical translation, continued engineering optimization is rapidly improving CFPS performance and robustness. This review summarizes the fundamental principles of CFPS, its major technological platforms, recent progress in diagnostic applications, and key challenges and opportunities for future development.

|
Scooped by
mhryu@live.com
April 14, 4:31 PM
|
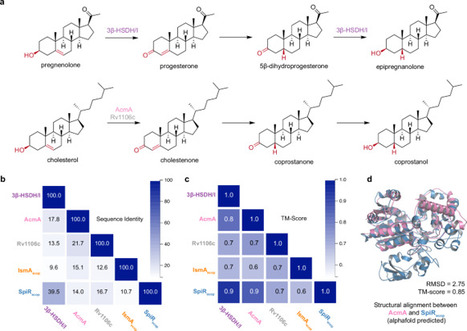
The gut microbiota contributes to cholesterol homeostasis by converting cholesterol into coprostanol, a non-absorbable sterol excreted in the feces. However, the enzymes mediating this process remain poorly defined. Here, we identify spiR, a steroid Δ5-4 isomerase/3-keto reductase from Eubacterium coprostanoligenes that catalyzes the initial oxidation of cholesterol to cholestenone, a requisite step in coprostanol production. We confirm that SpiR oxidizes both cholesterol and pregnenolone, and stereospecifically reduces 3-keto-steroids to 3β-hydroxylated forms. We show that SpiR preferentially binds to cholesterol over related steroids and functions as an NAD(H)-dependent homodimer. Through phylogenetic analysis, we show that spiR clusters with known Δ5-4 isomerases and is restricted to an uncultured clade within Acutalibacteraceae, where it frequently co-occurs with species encoding ismA, a gene previously implicated in cholesterol conversion. We analyze a multi-omic dataset from three human cohorts and find that spiR homologs were strongly enriched in individuals exhibiting cholesterol conversion. We also show that spiR homologs have a greater predictive power for cholesterol conversion than ismA homologs, establishing them as superior markers of microbial cholesterol metabolism. Our findings refine the enzymatic model of cholesterol metabolism in the gut and establish spiR as a critical biomarker and mechanistic driver for microbiome-mediated cholesterol reduction. Here, the authors identify SpiR, a gut bacterial enzyme that converts cholesterol, exclusively in a clade of uncultured gut bacteria, and show it is a superior predictor of microbial cholesterol metabolism in humans.

|
Scooped by
mhryu@live.com
April 14, 4:06 PM
|
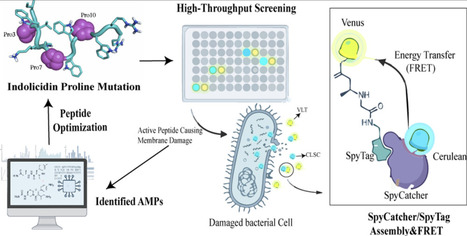
Antimicrobial peptides (AMPs) are key to innate immunity but face challenges in rational design due to their diversity and complex mechanisms. The lack of sensitive and full function-based high-throughput screening methods for AMP variants has further impeded systematic exploration and optimization. To address this, we developed a high-throughput screening method for AMPs integrating SpyTag/SpyCatcher click biology and fluorescence resonance energy transfer (FRET). Attributing to the ultrahigh reaction rate and specificity of the SpyTag/SpyCatcher pair, this screening method allows real-time fluorometric detection of the cell lytic activity of AMPs against their natural targets, i.e., bacterial cells. Applied to an indolicidin proline-scanning library, this approach identified variants with enhanced activity against E. coli and B. subtilis. Collectively, this work establishes a function-based, high-throughput screening platform for AMPs that can fully reflect their functional complexity, thereby providing an efficient and scalable tool for screening AMP libraries and facilitating data-driven optimization and functional analysis.

|
Scooped by
mhryu@live.com
April 14, 4:00 PM
|
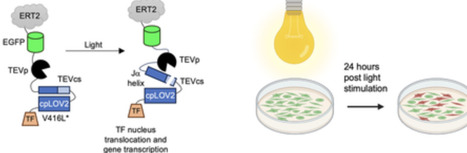
Optogenetic tools have revolutionized the control of gene expression with high spatial and temporal resolution. Here we present a Single-chain Light-Activatable Transcriptional Reporter (SLATR), a system capable of fluorescently tagging target cells with minutes of white light stimulation. In its inactive, or dark state, a transcriptional factor is cytosolically bound, preventing nuclear translocation. White light irradiation triggers its release through the protease cleavage of a site that is sterically caged by the circularly permuted Avena sativa LOV2 (cpAsLOV2) domain. We discovered that cpAsLOV2 cages the cleavage site more efficiently than AsLOV2, achieving low background in the SLATR design. We demonstrate that SLATR exhibits a signal-to-background ratio between 3.4 and 36 and achieves reporter activation within 60 min of light stimulation. Furthermore, SLATR outperforms the only other single-chain light-activatable transcriptional reporter, LAUNCHER, with faster kinetics, greater light sensitivity, and markedly lower background under identical stimulation conditions. Our single-chain light-activatable transcriptional system expands the optogenetic toolkit though providing a simpler system for regulating gene expression with precise spatiotemporal control.

|
Scooped by
mhryu@live.com
April 14, 3:44 PM
|
Engineered probiotics are emerging as versatile biological platforms capable of delivering therapeutic functions, modulating host–microbiota interactions, and enabling innovative strategies for preventing or treating metabolic, infectious, and inflammatory conditions. Advances in synthetic biology have expanded microbial engineering along a continuum ranging from self-cloned or intragenic modifications—based on deletions or recombination events that recapitulate naturally plausible genomic changes—to fully transgenic constructs expressing heterologous bacterial, viral, or human genes. This technological diversity demands proportionate and mechanistically informed safety evaluation, with particular emphasis on genetic stability, ecological compatibility, and the potential for horizontal gene transfer (HGT). This review examines the principal applications of engineered probiotics in human health, including strains designed to enhance endogenous functions, eliminate detrimental activities, neutralize toxins, interfere with pathogen signaling, degrade biofilms, express therapeutic proteins, act as mucosal vaccine platforms, serve as tumor-targeted immunotherapeutic vectors, or enable emerging systemic and brain-directed delivery strategies. We also highlight the current regulatory heterogeneity across international frameworks and discuss the relevance of recent EFSA guidance, which clarifies that modifications involving only deletions or the reinsertion of native sequences may entail markedly different regulatory obligations compared with constructs carrying truly novel genetic traits. To promote regulatory convergence, we propose a unified safety-assessment framework that integrates classical toxicological testing with a construct-specific evaluation of HGT potential. This approach combines whole-genome sequencing to define the engineered locus, validated qPCR assays for highly specific detection, and controlled exposure experiments using competent microbiota and environmental recipient strains to quantify the extremely low probability of gene transfer under worst-case conditions. Such a structured methodology provides a scalable, evidence-driven basis for evaluating engineered probiotics according to the biological nature of the modification rather than a one-size-fits-all model. Engineered probiotics hold substantial translational promise, provided that safety assessments remain adaptive, risk-proportionate, and aligned with mechanistic understanding of microbial genetics and ecology.

|
Scooped by
mhryu@live.com
April 14, 2:54 PM
|
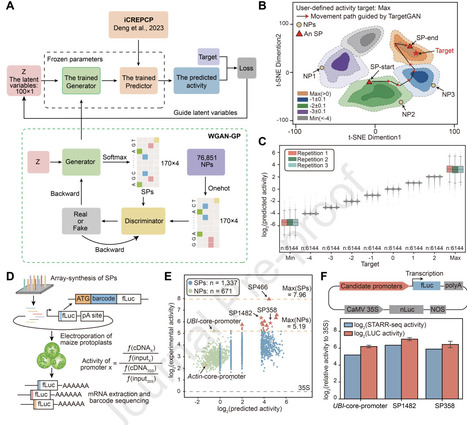
Plant core promoters (PCPs) are key genetic elements controlling gene expression, holding significant value for crop breeding and plant synthetic biology. However, natural promoters (NPs) are constrained by limited diversity and a narrow activity range, and it remains unclear whether synthetic promoters (SPs) can transcend these natural constraints. Here, we present TargetGAN, a deep learning framework trained on 76,851 NPs that integrates generative adversarial networks (GANs) with a pre-trained activity predictor to enable the de novo design of PCPs with user-defined activity. We used TargetGAN to generate 55,296 SPs, selecting 5,250 for high-throughput functional validation using STARR-seq. Of these, 2,909 were successfully characterized, exhibiting a moderate correlation (PCC = 0.6435) between predicted and experimental activity. Surprisingly, 29 of these SPs exhibited ultra-high activity, exceeding the maximum activity of tested NPs. Further orthogonal validation via luciferase (LUC) reporter assays demonstrated a strong positive correlation with STARR-seq measurements across a broad dynamic range. Notably, the most active synthetic candidate, SP1482, significantly outperformed the strongest tested NP, the UBI-core-promoter, achieving a 128-fold expression increase relative to the 35S minimal promoter. Interpretable motif analysis suggests that ultra-high-activity promoter design can be achieved through the precise arrangement of strong activating motifs. These results demonstrate that TargetGAN is a robust and generalizable framework for the targeted generation of PCPs tailored to user-defined targets, and will be a powerful tool both for precise gene regulation in plant systems and for overexpression analysis in genetic engineering and synthetic biology.

|
Scooped by
mhryu@live.com
April 14, 1:31 PM
|
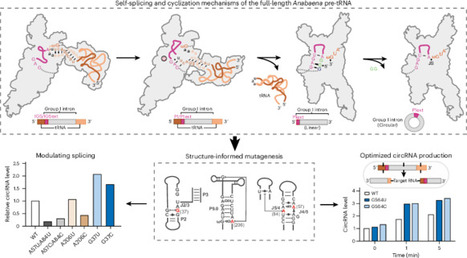
Group I introns are catalytic RNAs capable of self-splicing and generating circular RNAs, processes central to RNA metabolism and biotechnology. Yet, full-length ribozyme structures containing entire exon sequences and the structural basis of postsplicing circularization have remained limited. Using cryo-electron microscopy, we resolved multiple conformational states of the full-length Anabaena tRNA(Leu) precursor, capturing key intermediates of splicing and cyclization. In the apo state, the exons preassemble into a mature tRNA-like conformation that promotes P1 helix formation. Transitions through the splicing states involve substantial rearrangements essential for catalysis. Unlike other group I introns, the Anabaena intron circularizes without sequence loss, using its guanosine-binding site as the catalytic center. Mutational analyses confirm that G37 reorientation and a conserved wobble receptor motif precisely position the circularization site, driving efficient cyclization even in engineered PIE systems. These findings uncover unique mechanisms of RNA catalysis and establish structure-based optimization for advancing RNA circularization technologies. This study reveals how a group I intron from an Anabaena pre-tRNA undergoes structural changes to splice RNA and form circular RNA. The structures define key catalytic steps and offer a blueprint for engineering more efficient circular RNA systems.

|
Scooped by
mhryu@live.com
April 14, 1:13 AM
|
Chromalveolate algae such as diatoms, haptophytes, and dinoflagellates are main contributors to oceanic primary production, sustaining marine ecosystems and global carbon cycles while synthesizing a striking array of acetylated carotenoids like fucoxanthin and peridinin. These pigments optimize photosynthetic light harvesting in the algae and offer nutritional benefits for humans, yet knowledge of their biosynthetic pathways is still incomplete, particularly the shared acetylation step. By screening 39 candidate genes in the diatom Phaeodactylum tricornutum, we identified an enzyme with xanthophyll acetyltransferase (XACT) activity that is indispensable for this modification. Disrupting XACT in Phaeodactylum and the eustigmatophyte Nannochloropsis oceanica abolished xanthophyll acetylation. Phylogenetic analyses revealed that XACT is exclusively present in chromalveolates synthesizing acetylated xanthophylls. In vitro assays with recombinant XACT enzymes from Phaeodactylum, Nannochloropsis, the brown alga Ectocarpus siliculosus, the dinoflagellate Symbiodinium tridacnidorum, and a haptophyte confirmed their general activity toward allenic precursor carotenoids but exhibited lineage-specific substrate preferences, explaining the diversified carotenoid structures across lineages. The broad substrate specificity of XACT from Phaeodactylum led us to reinvestigate the substrate specificities of other enzymes involved in fucoxanthin formation, indicating that fucoxanthin biosynthesis in diatoms proceeds via a multibranched rather than a linear pathway. XACT from Ectocarpus showed a distinctly narrow substrate spectrum, providing key evidence for the order of the two previously proposed steps in brown algal fucoxanthin biosynthesis. Our work resolves a long-standing gap in marine carotenoid biosynthesis and identifies the relaxed substrate specificities of the enzymes involved as an important driver for the multitude of algal carotenoid structures.
|
 Your new post is loading...
Your new post is loading...