Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 12:50 AM
|
Bacterial communities and the bacteriophages infecting them are the basis of every ecosystem, including holobionts. The various ways in which these microorganisms interact with each other in complex communities over the life of the host affects the holobiont fitness. Despite being ubiquitous and environmentally relevant, plant-associated microbial communities remain understudied, especially in the phyllosphere, mainly because of the low abundance of microbes and the complexity of the system. In this work we followed bacteria and phage community dynamics in the phyllosphere over a growing cycle of Arabidopsis thaliana, to understand the ecology and relevance of bacteriophages in complex bacterial communities. We focused on Pseudomonas, a common plant pathogen and commensal, and the phages infecting them, in three setups of increasing complexity: in vitro, controlled experiments in planta and in wild populations of A. thaliana. We found that bacterial communities are resilient to phage infection, and more dynamic than the phages infecting them over the growing season, suggesting that although ubiquitous and abundant, bacteriophages exert selective pressures on leaf bacterial communities only intermittently.

|
Scooped by
mhryu@live.com
Today, 12:37 AM
|
Bile acids and their derivatives are key regulators of host metabolism, immunity, and microbiome interactions, with growing therapeutic potential. Conventional production methods of bile acids relying on animal extraction or chemical synthesis are inefficient, costly, and environmentally unfriendly. Microbial synthesis offers a sustainable alternative but is limited by the absence of genetic tools for native anaerobic producers. Here, we reconstructed the bile acid 7α-dehydroxylation pathway in Bacillus subtilis, a safe and tractable host. We characterized the key Bai enzymes and performed comprehensive analyses of the bile acids generated through in vitro assays and whole-cell catalysis. Multiple intermediates and end products, including cholyl-CoA, 3-oxo-cholic acid, 3-oxo-4,5-6,7-didehydro-deoxycholic acid, 3-oxo-4,5-dehydro-deoxycholic acid, 3-oxo-deoxycholic acid, and deoxycholic acid, were identified. This work establishes a functional heterologous platform for bile acid biosynthesis, enabling sustainable production and future applications in therapeutics and microbiome engineering.

|
Scooped by
mhryu@live.com
Today, 12:10 AM
|
In light of climate change and growing resource scarcity, microbial production of organic acids offers a sustainable alternative to fossil-based chemical synthesis. In this study, malic acid production by Aspergillus oryzae was optimized through cultivation temperature adjustment and biotin supplementation, while organic acid formation from various carbon sources was systematically characterized. The process conditions applied during substrate screening, including pH control using Na2CO3 and NaOH, Zn2+ supplementation and hypoxia, were based on previously established strategies to stimulate malic acid production. Cultivation at 35°C increased respiratory activity compared to 32°C, resulting in an average productivity of 0.17 g L−1 h−1. Biotin supplementation enhanced productivity by 20% and increased the carbon yield, defined as the proportion of consumed carbon recovered in malic acid, by 5%. Under optimized cultivation conditions, the highest malic acid productivity was achieved in cultivation with glucose as substrate and Na2CO3 as pH-neutralizing agent, reaching 57.57 g L−1 malic acid with a yield of 0.66 g g−1 and an overall productivity of 0.24 g L−1 h−1, while fructose and glycerol resulted in substantially lower productivities. Furthermore, we demonstrate the ability to perform carbon balancing even in the presence of carbonate-based neutralizing agents. This is achieved by quantifying and subtracting the CO2 generated during neutralization reactions from the total emissions, enabling precise determination of microbial CO2 production and calculation of carbon yields. By systematically combining optimization strategies reported in previous studies, this work achieves productivity and carbon efficiency exceeding those of the individual approaches reported so far.

|
Scooped by
mhryu@live.com
April 15, 11:25 PM
|
Postbiotics, defined as preparations of inanimate microorganisms and their components that confer health benefits, are emerging as stable and safe alternatives to probiotics. This review summarizes recent advances in the biochemical composition, mechanisms of action, and applications of postbiotics and paraprobiotics in human and animal nutrition. It also highlights innovations in omics technologies, artificial intelligence, and sustainable production strategies that are shaping next-generation microbiome-based functional foods and therapeutic interventions.

|
Scooped by
mhryu@live.com
April 15, 10:55 PM
|
Investigating ecological interactions within microbial ecosystems is essential for enhancing our comprehension of key ecological issues, such as community stability, keystone species identification, and the manipulation of community structures. However, exploring these interactions proves challenging within complex natural ecosystems. With advances in synthetic biology, the design of synthetic microbial ecosystems has received increasing attention due to their reduced complexity and enhanced controllability. Various ecological relationships, including commensalism, amensalism, mutualism, competition, and predation have been established within synthetic ecosystems. These relationships are often context-dependent and shaped by physical and chemical environmental factors, as well as by interacting populations and surrounding species. This review consolidates current knowledge of synthetic microbial ecosystems and factors influencing their ecological dynamics. A deeper understanding of how these ecosystems function and respond to different variables will advance our understanding of microbial-community interactions.

|
Scooped by
mhryu@live.com
April 15, 5:19 PM
|
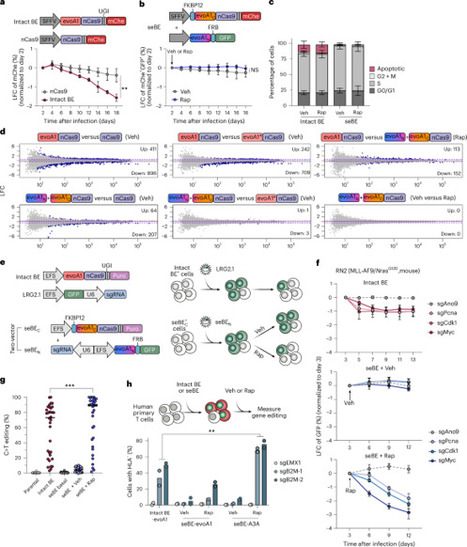
Cancer functional genomics using CRISPR base editors (BEs) holds great promise for molecular characterization and new target discovery. However, traditional BEs, using intact DNA deaminases as mutators, are often constrained by limited control and nonspecific toxicities. Here we developed a small-molecule-controllable system using split-engineered BEs (seBEs). By placing deaminase activity under small-molecule control, seBEs significantly reduced cellular toxicity and enabled robust and inducible in vivo functional genomics screens. High-density seBE genetic screens using ~11,000 single guide RNAs in vitro and ~3,700 single guide RNAs in vivo reveal known and previously unknown loss-of-function and dominant-negative mutations in cancer therapeutic targets. A deeper tiling seBE screen against Adar1, a key mediator in cancer immunotherapy, reveals critical residues within functional domains that show no phenotype in vitro but distinctively elicit non-cell-autonomous cancer dependencies in vivo. Overall, our seBE system offers a generalizable, controllable and highly efficient method to systematically identify key residues in cancer functional genomics. Inducible base editors enhance cancer genomic screens in vivo.

|
Scooped by
mhryu@live.com
April 15, 4:58 PM
|
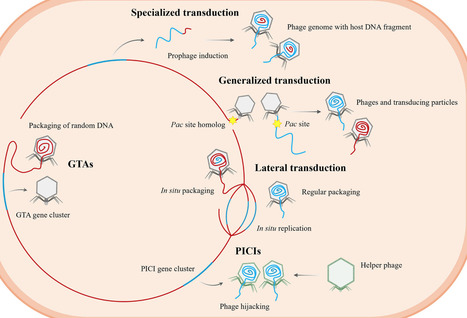
Fecal microbiota transplantation (FMT) is an effective therapy for recurrent Clostridioides difficile infection and is increasingly being explored for other microbiota-associated diseases. However, general research has largely focused on bacterial engraftment, overlooking the contribution of the gut virome. In this perspective, we highlight phage-mediated horizontal gene transfer (HGT) as a potentially influential process occurring following FMT. Donor-derived phages may potentially influence community structure, engraft in resident bacteria, and modulate microbial functions or host physiology. In addition, temperate phages are well-equipped to mobilize bacterial genes, such as metabolic functions, stress-response traits, and antibiotic resistance determinants, raising the possibility that gene flow could well contribute to FMT outcomes. We propose a conceptual model in which phages act as bidirectional mediators of adaptation, not only accompanying bacterial communities but also influencing gut ecosystems in subtle, yet potentially consequential, ways.

|
Scooped by
mhryu@live.com
April 15, 3:45 PM
|
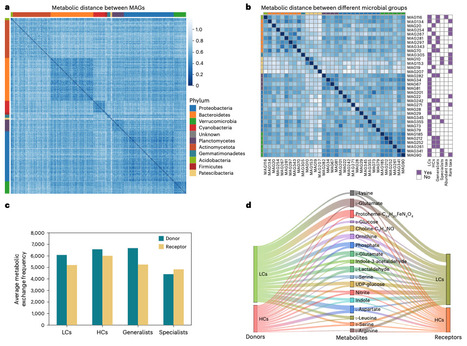
Microorganisms secrete extracellular vesicles (EVs) that transport bioactive molecules, including proteins and metabolites. While their functions are well studied in model microbes, their ecological contributions to natural ecosystems remain largely unexplored. Here we performed an integrative study investigating the role of environmental EVs in shaping microbial community assembly in the Xinglinwan Reservoir. By combining genome-scale metabolic models and multi-omics of field EVs, we found that EVs mediated metabolite exchanges mainly through carrying amino acids, disaccharides, carbohydrate-active enzymes (CAZymes) and signals. EVs can facilitate the growth of amino acid auxotrophic strains. Moreover, EVs act as an external reservoir of functional traits, potentially reinforcing stochastic assembly processes and conferring functional redundancy to the ecosystem. Collectively, our integrative data demonstrate that EV-mediated metabolic exchange is an auxiliary mechanism supplementing classical nutrient transport in aquatic environments. EVs emerge here as a significant, distinct vector in biogeochemical cycling, offering a critical layer for resolving complex natural microbial interactions. Microorganisms release extracellular vesicles, but their ecological roles in natural environments remain unclear. A year-long multi-omics study reveals that environmental extracellular vesicles mediate metabolite exchange central to carbon and nitrogen cycling while stabilizing microbial communities.

|
Scooped by
mhryu@live.com
April 15, 1:30 PM
|
The root nodules formed by rhizobia and leguminous plants are specialized structures for nitrogen fixation. However, a large number of non-rhizobial endophytes (NREs) also coexist within the nodules, and their contribution to nitrogen fixation under abiotic stress conditions remains unclear. Here, using the wild leguminous shrub Sophora davidii as model system, we identified an important NRE (Bacillus siamensis BT-9-1) by analyzing keystone taxa within the bacterial cooccurrence network of root nodules. This strain could improve the survival of Mesorhizobium metallidurans YC-39 under saline-alkali stress. A mechanistic investigation revealed that the expression of ilvA, ilvH, and ilvD was downregulated, and the contents of (2S)-isopropylmalate and succinic acid decreased in M. metallidurans YC-39 under saline–alkali conditions, whereas B. siamensis BT-9-1 presented increased accumulation of these metabolites. These findings indicate that B. siamensis BT-9-1 cross-feeds M. metallidurans YC-39 with these metabolites, rescuing the compromised branched-chain amino acid synthesis pathway and the TCA cycle in saline–alkali environments. Eventually, coinoculation with B. siamensis BT-9-1 and M. metallidurans YC-39, along with (2S)-isopropylmalate and succinic acid supplementation, increased nitrogenase activity of the symbionts. Our study reveals a novel mechanism by which non-rhizobial endophyte Bacillus species enhances the growth and nitrogen fixation efficiency of M. metallidurans under saline-alkali stress through the delivery of key metabolites.

|
Scooped by
mhryu@live.com
April 15, 1:51 AM
|
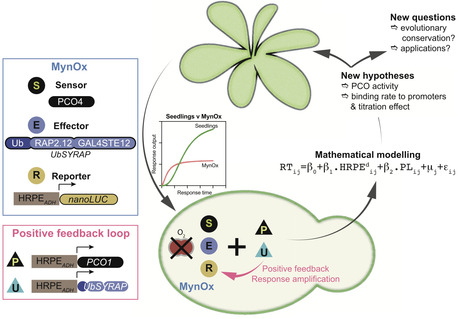
Plants face hypoxic conditions either chronically, as particular tissues are characterized by fluctuating or stable low oxygen levels, or acutely, when flooded. In vascular plants, transcriptional adaptive responses to hypoxia are rapidly mounted by Ethylene Response Factors VII (ERFVIIs), regulated by Plant Cysteine Oxidases (PCOs) through the cysteine branch of the N-degron pathway (Cys-NDP) for oxygen sensing. However, this relatively simple regulatory circuit, consisting of both constitutively expressed as well as hypoxia-inducible ERFVIIs and PCOs, interacts with diverse signaling cues and pathways invoked by hypoxia. To understand the share of the PCO-mediated oxygen sensing mechanism in the production of hypoxia responses, we insulated the PCO/ERFVII circuit from Arabidopsis thaliana and adapted it to Saccharomyces cerevisiae. Using a reporter gene to monitor the output of the circuit allowed us to compare the speed and amplitude of response to hypoxia in the engineered yeast and the source organism. Hypoxia triggered ERFVII stabilization both in Arabidopsis and yeast, leading to a similarly fast transcriptional response that was however larger in plants. A simple hypoxia-inducible feedback loop improved the amplitude of response in yeast, demonstrating the importance of this regulation in the endogenous PCO/ERFVII circuit. Finally, computational modeling of the yeast circuit enabled us to identify promoter competition and presence of hypoxia-inducible PCOs as key parameters that shape early hypoxia responses in plant cells.

|
Scooped by
mhryu@live.com
April 15, 1:39 AM
|
Abiotic stress is a major problem which threatens agricultural productivity and global food security. Drought, extreme temperatures, salinity, and heavy metal contaminations cause disturbances in plant cellular homeostasis, impair metabolic processes, and reduce crop yields. Plants possess innate defense mechanisms against abiotic stresses by expressing stress-responsive genes, accumulating osmo-protectants, and the activation of antioxidant enzymes. Failure of these defense mechanisms under prolonged stress conditions leads to homeostatic imbalances. Recently, nanotechnological approaches like nanoparticle-mediated delivery systems have been developed to enhance the plants' resilience against stress conditions. Engineered nanoparticles (ENPs) have unique physicochemical properties such as a high surface area, high reactivity, and a tunable surface chemistry. These properties enable the nanoparticles to interact with plant systems at molecular and cellular levels, modulate stress signaling pathways, trigger the upregulation of stress-responsive genes, and reduce oxidative stress by increasing antioxidant enzymes' activity. However, under certain environmental conditions, ENPs may also induce oxidative damages and exhibit phytotoxicity. This review encompasses a comprehensive overview on the role of nanoparticles in abiotic stress management, along with detailed insights into the biosafety and environmental toxicity of ENPs. Overall, the review highlights the novel insights of ENPs-plant interactions and identifies existing knowledge gaps through a systematic literature review to guide future research towards sustainable agriculture.

|
Scooped by
mhryu@live.com
April 15, 1:22 AM
|
As ubiquitous features of every natural environment, microbes have profoundly shaped eukaryotic biology throughout evolution. Circadian clocks evolved in all domains of life as central regulators that align physiology with environmental cycles, yet whether they respond directly to microbial signals remains unknown. Here, we demonstrate that evolutionarily diverse microbes potently reset mammalian cellular clocks and can drive phase shifts in plants and algae, indicating cross-kingdom effects of microbes on circadian rhythms. In mammals, exposure to soluble bacterial components distinct from canonical innate immune ligands induces acute PER2 upregulation independently of Bmal1 or nascent transcription. A targeted inhibitor screen and biochemical assays implicate p38 MAPK as a modulator of this response. Taken together, this positions bacterial exposure as a previously unrecognized circadian clock input, revealing a new axis of host-microbe interaction that modulates biological timing at the cellular level.

|
Scooped by
mhryu@live.com
April 15, 12:55 AM
|
Streptomyces species have long been recognized as prolific sources of novel natural products, generating a wide array of clinically-relevant antimicrobial and cytotoxic compounds. In this review, we examine 174 publications from 2021 to 2024 that describe 716 previously unreported secondary metabolites isolated from Streptomyces strains sourced from a range of marine and terrestrial environments. Natural product researchers used a variety of chemical and biological techniques, including genome mining, molecular networking, and heterologous expression, to uncover novel compounds with a diverse array of structures, including cyclic and linear peptides, macrocyclic, polyaromatic, and other polyketides, terpenoids, alkaloids, and azoxy compounds, among others. The variety of bioactivities exhibited by these secondary metabolites, including antibacterial, anticancer, antifungal, antiviral, and anti-inflammatory effects, illustrates the enduring potential of the Streptomyces genus to produce promising new bioactive natural products.
|

|
Scooped by
mhryu@live.com
Today, 12:44 AM
|
The fitness and virulence of Pseudomonas aeruginosa rely on its ability to maintain a functional pool of ribosomes, which are essential for protein synthesis. This review explores the intricate ways of ribosome protection, rescue and hibernation, by which P. aeruginosa preserves ribosome functionality under stress. These processes enhance the adaptability and resistance of this pathogen to ribosome-targeting antibiotics and present significant challenges to current therapeutic strategies. By highlighting recent discoveries and identifying promising directions for future research, this review aims to explore potential targets for innovative drug discovery.

|
Scooped by
mhryu@live.com
Today, 12:13 AM
|
Plant synthetic biology is a highly innovative field that aims to better understand, redesign, and reprogram plants. Progress in recent years has been driven by technical developments such as modular cloning, gene circuits, genome editing, and synthetic genomics, which have expanded the field beyond traditional single-gene strategies. However, most efforts remain focused on angiosperms, leaving much of plant diversity underexplored and limiting potential applications due to the inherent complexity of these systems. Simpler, underutilized plant systems, particularly bryophytes, provide an alternative experimental platform for rapid tool development and, more broadly, for the establishment of universal as well as synergistic bioengineering approaches across different plant lineages. This review highlights recent advances that underscore the importance of bryophytes as plant synthetic biology systems. Progress across mosses, hornworts, and liverworts is discussed, with particular emphasis on Marchantia polymorpha as a leading model.

|
Scooped by
mhryu@live.com
Today, 12:02 AM
|
Internal chemical modifications of mRNA constitute a key epitranscriptomic layer of gene regulation in eukaryotes. Although N6-methyladenosine (m6A) has been the most intensively studied, accumulating evidence reveals important roles of non-m6A mRNA modifications, such as 5-methylcytosine, N4-acetylcytidine and pseudouridine, in plants. These modifications modulate diverse aspects of mRNA metabolism, including alternative splicing, stability, translation and long-distance transport, and thereby shape plant development and environmental adaptation. Here we summarize current advances in understanding non-m6A mRNA modifications in plants, emphasizing their regulatory roles in mRNA metabolism and their effects on plant development and stress resilience. We also review the detection technologies for non-m6A modifications and discuss key challenges and future directions towards elucidating their regulatory functions. This Review highlights emerging roles of non-m6A modifications, such as 5-methylcytosine, N4-acetylcytidine and pseudouridine, in plant mRNAs, outlining their effects on mRNA processing, development and stress adaptation, as well as advances in detection methods and future directions.

|
Scooped by
mhryu@live.com
April 15, 11:16 PM
|
Engineering small-molecule binding proteins de novo remains a significant challenge as even advanced generative models struggle to model the atom-level details of protein-ligand interactions with sufficient accuracy. Higher experimental success rates have resulted from methods that explicitly scaffold predefined binding interactions into helical bundles. Here we introduce a scaffolding strategy that generalizes to alpha-beta architectures. By screening thousands of combinatorially assembled protein-ligand interactions against diverse de novo backbones with finely varied pocket geometries, the protocol allows for high-fidelity accommodation of target interaction geometries. Our protocol then integrates physics-based and deep learning methods for optimization of interfacial interactions and sequence-structure compatibility, considerably improving in silico design metrics. Applying this method to two chemically similar steroids achieved a notable experimental success rate (4/26 designs bind their targets), and NMR structures of two designs are in good agreement with design models. Our generalizable, atomically precise approach offers a robust framework for small-molecule binder design, effectively eliminating the need for high-throughput screening.

|
Scooped by
mhryu@live.com
April 15, 5:22 PM
|
Protein allostery underlies most information and energy processing in biology and the development of artificial allosteric proteins is a key objective of synthetic biology and biotechnology. We show that machine-learning-engineered minimal ligand-binding domains act as efficient receptors in single-component allosteric switches, despite lacking global conformational change. Such colorimetric, luminescent and electrochemical biosensors of small molecules, peptides and proteins can be compiled into intramolecular YES and AND logic gates. Furthermore, we report fully synthetic allosteric switches composed of artificial receptor and reporter domains. Hydrogen/deuterium exchange mass spectrometry and 19F nuclear magnetic resonance analyses suggest that ligand binding reduces the conformation entropy of the system, increasing the catalytic activity of the reporter domain. The potential practical utility of this approach is demonstrated by engineering Escherichia coli cells with steroid-dependent antibiotic resistance and by developing bioelectronic devices capable of quantifying steroid hormones. Allosteric biosensors are constructed by combining reporter proteins with machine-learning-designed receptor domains.

|
Scooped by
mhryu@live.com
April 15, 5:03 PM
|
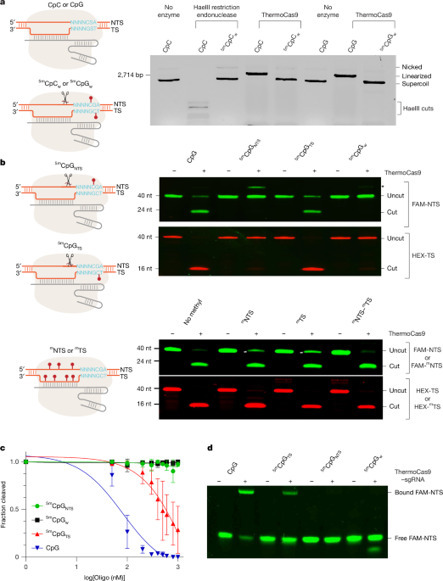
The bacterial CRISPR–Cas9 nuclease has become a powerful genome manipulation tool for a wide range of organisms. However, it has yet to fully leverage the pervasive presence of DNA methylation in genomes. Here, to fill this gap, we report biochemical, structural and human genome-editing characterizations of a methylation-sensitive Cas9 (ThermoCas9). ThermoCas9 efficiently binds to and cleaves DNA upstream of its protospacer adjacent motif (PAM) 5′-NNNNCGA-3′ or 5′-NNNNCCA-3′ in vitro. Methylation of the fifth cytosine in either PAM sequence (5mCpG or 5mCpC), however, significantly inhibits ThermoCas9 activity. Cryo-electron microscopy structures of ThermoCas9 in pre-cleavage and post-cleavage states at 2.8 Å and 2.2 Å resolution, respectively, reveal the molecular basis for the stringent requirement of the unmethylated cytosine in PAM binding and provide guidance for further enzyme engineering. We demonstrate methylation-sensitive editing by ThermoCas9 in human cell lines with distinct DNA methylation landscapes. Moreover, we demonstrate that a catalytically enhanced ThermoCas9 efficiently targets luminal expression signature genes that are consistently hypomethylated in patients with breast cancer. Owing to its sensitivity to DNA methylation, ThermoCas9 can specifically target cells with disease-related hypomethylation, which adds another layer of precision to genome-editing technologies. ThermoCas9, a genome-editing enzyme that is sensitive to the DNA methylation status of the target locus, is characterized and shows promise for targeting hypomethylated DNA regions in cancer cells.

|
Scooped by
mhryu@live.com
April 15, 4:53 PM
|
Understanding peptide properties is often assumed to require modeling long-range molecular interactions, motivating complex graph neural networks and pretrained transformers. Whether such long-range dependencies are essential remains unclear. We investigate if simple, domain-specific molecular fingerprints can capture peptide function without these assumptions. Atomic-level representations aim to provide richer information than purely sequence-based models and better efficiency than structural ones. Across 132 datasets, including LRGB and five additional peptide benchmarks, models using count-based ECFP, Topological Torsion, and RDKit fingerprints with LightGBM achieve state-of-the-art accuracy. Despite encoding only short-range molecular features, these models outperform GNNs and transformer-based approaches. Control experiments confirm that fingerprints, though inherently local, suffice for robust peptide property prediction. Our results challenge the presumed necessity of long-range interaction modeling and highlight molecular fingerprints as efficient, interpretable, and lightweight alternatives.

|
Scooped by
mhryu@live.com
April 15, 1:37 PM
|
Fermentative Clostridium species associated with rice roots can contribute substantially to biological nitrogen fixation in anoxic paddy soils, yet whether their biological nitrogen fixation is regulated by the redox chemistry of rhizosphere remains unclear. Here we show that iron plaques on rice roots function as terminal electron acceptors that reprogram Clostridium fermentation and thereby enhance biological nitrogen fixation. In nitrogen-fixation microcosms, Clostridium sensu stricto I was selectively enriched under plaque-associated Fe(III)-reducing conditions, coinciding with elevated nitrogen fixation. Metabolomic profiling coupled with metabolic flux analysis revealed that Fe(III) reduction redirects a portion of carbon and electron flow from low-energy-yield solventogenesis toward high-energy-yield acidogenesis. This shift increases cellular ATP generation and expands the reductant pool, thereby benefiting the energetic and reductant demands of nitrogenase. Integrated transcriptomic and metagenomic analyses further identified NosR, a flavin mononucleotide-binding protein that is upregulated during Fe(III) reduction and may facilitate electron delivery to plaque-associated Fe(III). Our findings establish a mechanism in which iron plaque reduction optimizes fermentation for biological nitrogen fixation, providing fundamental insights into coupled Fe–N cycling in rice rhizospheres and suggesting potential strategies for sustainable nitrogen management in flooded agroecosystems.

|
Scooped by
mhryu@live.com
April 15, 12:33 PM
|
Reactive oxygen species are essential for cellular signalling and redox homeostasis, but their accumulation causes cellular oxidative stress. In inflammatory bowel disease, oxidative stress is linked to chronic inflammation and alterations in the gut microbiota. We hypothesised that these alterations may result from the impact of reactive oxygen species on the interactions between bacteria and their viruses, bacteriophages. We followed the evolution of three E. coli strains and a virulent bacteriophage in a chemostat under continuous growth and studied the impact of oxidative stress on this community. We show that both the bacteriophage and its three hosts persisted in the system over 10 days, but the relative abundance of bacteriophages was decreased in the presence of reactive oxygen species. Oxidative stress also limited bacteriophage population diversity by favouring the selection of specialist bacteriophages with a narrower host range. Concomitantly, reactive oxygen species accelerated the evolution of bacterial resistance to bacteriophages and drove the fixation of genomic mutations in genes related to cell surface structures or located in mobile genetic elements. These results highlight that oxidative stress impacts the evolutionary dynamics between bacteria and bacteriophages with consequences for microbiota diversity and potential implications in the context of intestinal inflammation.

|
Scooped by
mhryu@live.com
April 15, 1:43 AM
|
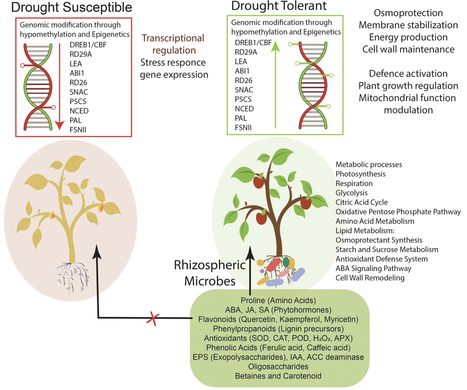
The frequent occurrence of drought, salinity and heat disasters due to global climate change has become a problem that cannot be ignored and seriously restricts food security and sustainable agricultural development. The role of rhizobacteria in the response of plants to abiotic stress has an important guiding significance in improving plant growth. This review summarizes the response of plant rhizosphere microbial communities to abiotic stress, analyzes the mechanism by which rhizosphere-related bacteria assist plants to resist abiotic stress, and expounds on the interaction between soil physical and chemical properties, the plant root metabolome, and the rhizosphere microbiome under abiotic stress. This review systematically summarizes the core roles and mechanisms of rhizobacteria in plants' defense against abiotic stress. Stress reshapes the rhizosphere microecology, with drought enriching Firmicutes and Actinobacteria, salt stress increasing Bacteroidetes abundance, and heat stress expanding the dominance of thermotolerant bacteria. Microbial diversity and network structure undergo adaptive reorganization. Streptomyces and Bacillus, as the twin stars aiding plants in enhancing stress resistance, provide medium- to long-term protection through rich secondary metabolites and mycelial networks, while Bacillus achieves acute responses via rapid spore germination, signal induction, and nutrient competition. Rhizobacteria improve soil nutrient availability by regulating carbon, nitrogen, and phosphorus cycles, secreting organic acids and enzymes, and induce plant osmotic adjustment, antioxidant, and anti-ethylene signaling networks through extracellular polysaccharides, volatile organic compounds, plant hormones, and 1-aminocyclopropane-1-carboxylic acid (ACC) deaminase pathways, thereby systematically enhancing the host's water use efficiency and membrane stability. Future research should integrate multi-omics and field validation to precisely construct rhizosphere bacterial communities, providing theoretical basis and technical routes for green agriculture.

|
Scooped by
mhryu@live.com
April 15, 1:34 AM
|
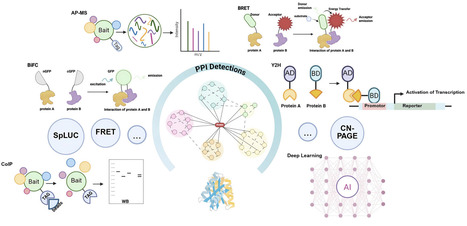
Protein–Protein interactions (PPI) wire plant cells, assembling metabolons, routing signals, and coordinating organelle crosstalk. We review experimental platforms and the computational signals long used to predict PPIs. While experimental platforms and traditional computational approaches have long been employed for PPIs prediction, recent advances in artificial intelligence offer unprecedented opportunities to map plant interactomes comprehensively. To provide a systematic overview, we categorize current methodologies into four thematic families: (i) sequence-centric predictors utilizing protein language models to extract evolutionary features; (ii) structure-based predictors integrating coevolutionary signals to reconstruct 3D complex arrangements; (iii) network-level learners employing graph architectures to capture global interactome topology; and (iv) geometric and generative methods leveraging symmetry-aware networks for specific site identification and de novo design. Despite rapid gains, plant applications are constrained by paralog expansion, compartmentalization, dynamic microenvironments, and the sparse availability of gold standards in the field. Next-generation plant AI PPI models should be organelle-aware, multimodal, rigorously benchmarked, structure-gated, and condition-validated.

|
Scooped by
mhryu@live.com
April 15, 1:06 AM
|
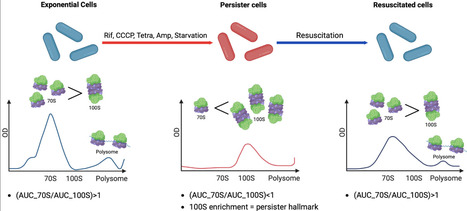
Persister cells survive antibiotic exposure and contribute to infection relapse, yet the molecular features that distinguish them from actively growing cells remain incompletely defined. Here, we used sucrose gradient-based ribosome sedimentation profiling to characterize ribosome complex distributions in E. coli persister cells and monitored their dynamics during resuscitation. Rifampicin-induced persister cells were characterised by pronounced enrichment of translationally inactive 90–100S ribosome complexes and a concomitant reduction in 70S ribosomes relative to exponentially growing cells. Upon nutrient replenishment, ribosome distributions progressively shifted toward higher 70S and polysome (complexes of multiple ribosomes simultaneously translating a single mRNA) levels, coinciding with growth recovery, indicating that resuscitation involves gradual remodelling of ribosome states rather than abrupt restoration of active translation. Functional analysis of ribosome-associated factors demonstrated that RMF, HPF and RaiA promote 100S ribosome accumulation and enhance persister formation, whereas deletion of rmf severely impaired both 100S formation and persistence. In contrast, loss of HflX did not measurably affect persister formation, consistent with a role downstream of persister establishment. In multiple stress-induced persister models including rifampicin, tetracycline, CCCP and starvation, as well as in a clinically relevant E. coli O157:H7 (EHEC) strain, ribosome distributions consistently exhibited a quantitative reversal of the AUC_70S/AUC_100S ratio (Ratio < 1.0) relative to exponentially growing cells (Ratio > 1.0). Collectively, these findings demonstrate that this shift in the 70S-to-100S balance is a consistent and shared feature of E. coli persister physiology and that ribosome state distributions link persister formation to resuscitation dynamics. These findings provide a quantitative ribosome-state framework that may inform the development of anti-persistence strategies targeting ribosome hibernation factors.
|
 Your new post is loading...
Your new post is loading...