Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 4:22 PM
|
Microbial communities are shaped by complex metabolic interactions, whereby the byproducts of one organism influence the physiology of others. This is exemplified in the microbial nitrogen cycle, where diffusion of free intermediates can drastically reshape the chemical landscape of the environment. One such intermediate, nitrous oxide (N2O), is often overlooked as biologically inert. However, emerging evidence suggests this gas may inhibit the activity of some cobalamin-dependent enzymes through a reaction with the cofactor. This raises the possibility that, through such an interaction, N2O-producing organisms may shape the microbial communities in which they reside, selecting against organisms that rely on these sensitive cobalamin enzymes. At the plant root, a hotspot of microbial activity, the impact of such interactions may be especially important. To investigate this, we focused on microbial N2O production and its effect on methionine biosynthesis, a ubiquitous bacterial process carried out by cobalamin-dependent (MetH) or independent (MetE) methyltransferases. In this study, we show that deleting metE and forcing reliance on MetH sensitizes the denitrifier Pseudomonas aeruginosa to exogenous and self-produced N2O. We extend these findings to plant-associated bacteria, where we find that a significant portion of an Arabidopsis thaliana rhizosphere culture collection relies exclusively on cobalamin-dependent methionine synthases and experimentally demonstrate their sensitivity to N2O. Finally, we show that the growth of one MetH-reliant rhizosphere isolate is suppressed in co-culture with N2O-producing P. aeruginosa. Together, these findings suggest that N2O producers can shape microbial ecology at the plant root.

|
Scooped by
mhryu@live.com
Today, 3:37 PM
|
Aflatoxin contamination in agricultural products poses a critical global food safety threat, demanding an urgent paradigm shift from conventional postcontamination quantification to proactive early warning strategies. This comprehensive review critically examines the revolutionary transition toward biomarker-driven biosensing platforms that enable the interception of aflatoxigenesis before toxin accumulation. By systematically analyzing stage-specific biomarkers─including fungal spores/volatile organic compounds, regulatory genes, metabolic proteins, and critical biosynthetic precursors─we elucidate how temporal biosensing creates crucial intervention windows. The review further evaluates advances in nanomaterial-enhanced transducers, multiplexed detection platforms, and AI-integrated sensor arrays while addressing persistent challenges in bioreceptor specificity and field deployment. Future progress hinges on the development of quantitative biomarker–toxin correlation models, advancing point-of-care devices, and implementing integrated monitoring networks. Distinct from existing reviews, this work presents a novel chronobiology-aligned framework for aflatoxin contamination by integrating biomarker discovery with biosensing innovation to enable proactive risk management and strengthen global food security.

|
Scooped by
mhryu@live.com
Today, 3:29 PM
|
Transfer RNAs (tRNAs) are essential components of the protein synthesis machinery. Their biogenesis is a highly regulated process that involves the incorporation of numerous post-transcriptional chemical modifications, essential for tRNA folding, cellular stability and function. The sequential process by which these modifications are introduced remains poorly characterized. Previous studies have suggested the existence of modification hierarchies, particularly in the anticodon-loop region, but also among tRNA core modifications. Here, aiming to understand the molecular mechanisms by which modifications are incorporated in a bacterial model organism, we employed a combination of NMR spectroscopy and biochemical methods to characterize the maturation process of several E. coli tRNAs. By monitoring tRNA maturation in a time-resolved fashion by NMR, we observed a conserved temporal pattern in the incorporation of the Ψ55, T54, and m7G46 modifications. We also show that Ψ55 stimulates the incorporation of T54 in E. coli tRNAPhe, tRNAVal and tRNAAsp, and stimulates that of m7G46 in tRNAPhe and tRNAAsp. Importantly, we also provide general insights into the impact of modifications on tRNA structural properties, and show that while post-transcriptional modifications generally have a structuring effect that reduces conformational heterogeneities, these effects are tRNA-dependent, with certain tRNAs being more affected than others. These findings provide fundamental insights into the molecular aspects of tRNA maturation in E. coli.

|
Scooped by
mhryu@live.com
Today, 2:59 PM
|
Ammonia-oxidizing archaea (AOA) are among the most abundant microorganisms in the ocean, playing a fundamental role in the marine nitrogen cycle. Although temperature and trace metal availability each individually influence the growth and activity of marine AOA, there is only a very limited understanding of the interactive effects of these two major factors on AOA in the rapidly changing ocean. Here, we show that the iron requirements of the model marine AOA species Nitrosopumilus maritimus SCM1 are highly sensitive to temperature changes. A 5 °C increase in growth temperature reduced SCM1 iron requirements by >80%, and was associated with a substantial increase in iron use efficiencies (IUE, mol C fixed/h/mol cellular Fe) under iron-limited and warming conditions. A thermally enhanced IUE enables SCM1 to more efficiently utilize scarce available iron supplies to support its growth. Whole-cell proteomic analysis revealed that iron limitation decreased expression of a ferredoxin and increased expression of a copper-dependent plastocyanin that became more pronounced with warming, suggesting coordinated electron transport response regulation under combined iron and temperature stress. The global impacts of these temperature-dependent changes to AOA iron demands were assessed using sensitivity experiments with a state-of-the-art biogeochemical model. Simulations showed that impacts on nitrification were concentrated at higher latitudes, but the alterations to ammonia concentrations were redistributed toward lower latitudes by mode and intermediate water transport. These findings reveal a previously unrecognized mechanism by which ocean warming may alleviate iron limitation of AOA, enhance their ecological competitiveness, and reshape ocean nitrogen cycling throughout marine ecosystems.

|
Scooped by
mhryu@live.com
Today, 11:43 AM
|
Lignin is the most abundant renewable source of aromatic carbon on Earth and a central yet historically underutilized component of lignocellulosic biomass. Its complex and heterogeneous molecular architecture has long constrained efficient and selective conversion into value-added products, despite its high aromatic carbon content and chemical functionality. Recent advances in lignin extraction, fractionation, modification, and application-driven design have substantially expanded the range of achievable material and chemical performance within circular bioeconomy frameworks. This review provides a comprehensive and critically integrated assessment of lignin valorization that explicitly links plant biosynthesis and structural diversity to industrial convertibility, functional materials development, and sustainability performance. Green extraction technologies—including deep eutectic solvent and hydrotropic systems—are evaluated with respect to lignin structural quality, energy demand, solvent recovery, and downstream compatibility. Targeted chemical and enzymatic modification strategies enabling more reproducible lignin streams are discussed alongside applications in carbon fibers, nanomaterials, adhesives, bioplastics, cementitious systems, and additive manufacturing. Quantitative benchmarking against fossil-based incumbents identifies application domains where lignin-derived materials already achieve comparable performance, as well as areas where intrinsic structural limitations remain. In parallel, catalytic depolymerization pathways toward renewable aromatic chemicals are assessed from both mechanistic and systems-level perspectives. Environmental and economic implications are critically examined using recent life-cycle and techno-economic evidence, highlighting the influence of allocation choices, energy integration, and comparison with lignin incineration for energy recovery. Overall, this review clarifies how application-targeted lignin design and system-level sustainability assessment are essential for translating lignin’s biological complexity into scalable, competitive solutions for sustainable materials and chemicals.

|
Scooped by
mhryu@live.com
Today, 10:50 AM
|
Identifying evolutionarily remote antimicrobial peptides (AMP) is crucial for discovering underexplored clinical candidates to combat antibiotic resistance. Existing experimental and computational methods are limited by their reliance on sequence identity to known AMPs, missing distant homologues. Here we introduce HMD-AMP, a protein language model-based approach for AMP discovery. HMD-AMP outperforms previous methods in identifying evolutionarily distant AMPs and enables the discovery of unknown and highly potent AMPs from metagenomic data. Applied to host and gut microorganism genomes of nine mammals, HMD-AMP revealed over 37 million predicted AMPs. Of 91 high-confidence sequences experimentally validated, 74 showed strong antibacterial activity and 48 were evolutionarily remote from known AMPs. Four of these AMPs exhibited broad-spectrum antibacterial activity at low effective concentrations and showed low toxicity, with the most potent peptide demonstrating therapeutic efficacy in a mouse model of peritoneal E. coli infection. This study introduces an effective strategy to uncover AMPs. A protein-language-based model enables improved discovery of evolutionarily distant antimicrobial peptides (AMPs). High-confidence AMPs showed strong antibacterial activity, with the most potent demonstrating efficacy in a mouse model of E. coli infection.

|
Scooped by
mhryu@live.com
Today, 12:25 AM
|
The healthy human vaginal microbiota is typically dominated by one species of Lactobacillus: L. iners, L. crispatus, L. jensenii, or L. gasseri. L. iners, the most prevalent vaginal microbe globally, is the most fastidious of the vaginal lactobacilli, has the smallest genome, and produces less lactic acid (only the L-isoform). L. iners is also less protective against bacterial vaginosis, and uniquely encodes a cholesterol-dependent cytolysin, inerolysin, suggesting it may be a pathobiont. Despite its central role in the health of over one billion females, L. iners biology remains poorly understood in part due to a lack of genetic editing tools. Here, we present findings that L. iners is naturally competent and can be transformed easily by exogenous DNA. Natural competence was leveraged to disrupt the iny gene encoding inerolysin, and comGA, encoding the ATPase component of the competence pilus. Both gene disruptions were accomplished using PCR assembled DNA fragments comprising a drug resistance gene cassette (tetM or ermB) flanked by ~2 kb regions of homology to the L. iners chromosome. We further demonstrate that comGA is essential for L. iners transformation. The ability to rapidly perform targeted deletions in L. iners with in vitro generated DNA templates provides a straightforward and much needed method to probe the genetics and physiology of these important vaginal bacteria.

|
Scooped by
mhryu@live.com
Today, 12:19 AM
|
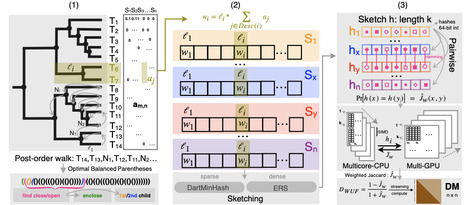
We introduce a new algorithm, DartUniFrac, and a near-optimal implementation with GPU acceleration, up to three orders of magnitude faster than the state of the art and scaling to millions of samples (pairwise) and billions of taxa. DartUniFrac connects UniFrac with weighted Jaccard similarity and exploits sketching algorithms for fast computation. We benchmark DartUniFrac against exact UniFrac implementations, demonstrating that DartUniFrac is statistically indistinguishable from them on real-world microbiome and metagenomic datasets.

|
Scooped by
mhryu@live.com
March 3, 11:47 PM
|
Genome architecture reorganizes over evolutionary time to support complex multicellularity without a proportional expansion of coding DNA. We conducted a cross-kingdom comparative analysis using high-quality RefSeq assemblies annotated by the NCBI Genome Annotation Pipeline, restricting the dataset to chromosome-level or complete genomes. Scaling relationships among genome size, gene content, and coding DNA content reveal compositional transitions that distinguish prokaryotic, unicellular eukaryotic, and multicellular lineages. Beyond ∼40 Mb of genic content, coding expansion slows and saturates, indicating compositional constraints that shaped the rise of multicellularity. These results establish scaling laws that quantify how noncoding sequence expansion dominates genome growth in complex eukaryotes.

|
Scooped by
mhryu@live.com
March 3, 11:01 PM
|
The intensifying climate crisis necessitates a global transition from fossil fuels to renewable energy sources. To meet this demand, metabolic engineering has become a pivotal strategy for developing microorganisms as efficient cell factories capable of producing fuels and fuel precursors. Among the biofuel platforms, fatty acid–based fuels are particularly promising, offering energy densities comparable to those of petroleum-based fuels. Recent advances in systems metabolic engineering, including metabolic pathway optimization, cofactor balancing, and dynamic regulation, have significantly improved the microbial production of key fuels and intermediates such as alka(e)nes, and fatty acid esters. In this review, we discuss recent progress in metabolic engineering strategies for microbial production of representative fatty acid-based fuels, highlighting current technological challenges and future directions.

|
Scooped by
mhryu@live.com
March 3, 10:43 PM
|
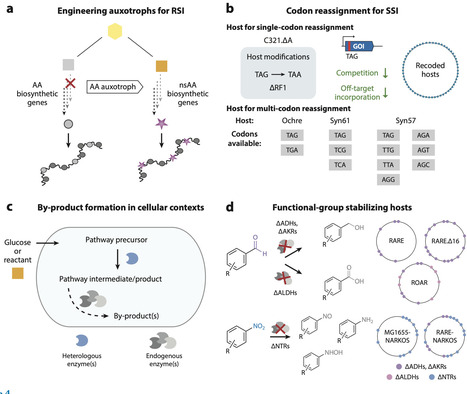
Genetic code expansion (GCE) is the ability to encode polypeptide building blocks beyond the standard 20 the ribosome uses for protein translation, known as nonstandard amino acids (nsAAs). The broadening of chemical functionalities in proteins produced by live cells has generated substantial value across fundamental and applied research settings. However, a common limitation of GCE approaches is their reliance on the supplementation of chemically synthesized nsAAs to cell culture media. To overcome this limitation of nsAA sourcing, efforts have engineered systems for nsAA biosynthesis, often in the same host that performs GCE. In recent years, these works have reported new chemical targets obtained through biosynthesis, as well as additional rationale for combining metabolic engineering and GCE, particularly for synthetic biology applications. Here, we review this rapidly advancing field and provide our perspectives on technical and conceptual innovations.

|
Scooped by
mhryu@live.com
March 3, 7:01 PM
|
Trimethoprim is a clinically important antibiotic used for the routine treatment of urinary tract infections as a cost-effective first-line choice for treatment. A unique feature of the drug is that it can have bacteriostatic or bactericidal effects depending upon the metabolites available in the environment. Bacteriostatic activity requires the absence of nucleoside from the growth media. Conversely, bactericidal activity requires the presence of a nucleoside and the amino acids glycine and methionine. Mechanistically, bacteriostatic action does not appear to be dependent upon protein or RNA synthesis, whereas protein synthesis does not appear to be essential for bactericidal activity. Instead, this is likely due to the inhibition of DNA synthesis and the triggering of programmed cell death, involving the suicide module mazEF. In summary, trimethoprim has a complex mechanism of action that should be considered when researching the antibiotic and informing growth media design to test susceptibility.

|
Scooped by
mhryu@live.com
March 3, 4:17 PM
|
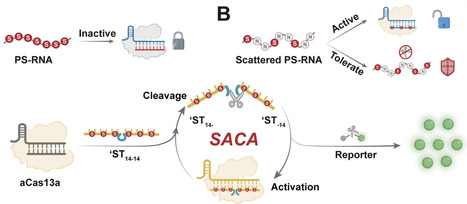
The CRISPR/Cas system is a powerful tool for molecular diagnostics, but its reliance on linear amplification constrains sensitivity, particularly for in situ imaging. Here, we discovered that phosphorothioate (PS)-modified activators can modulate Cas enzyme conformation via hydrophobic anchoring. By adjusting the PS modification sites, we achieved precise control over Cas activation and trans-cleavage resistance. Guided by this mechanism, we proposed a tailored design strategy featuring a “scattered” PS modification to engineer a linear “Coordinator” probe. This design effectively decouples Cas enzyme activation from substrate trans-cleavage resistance, enabling the construction of a Scattered PS Nucleic Acid-driven Cas Autocatalytic system (SACA). SACA achieves exponential amplification without external enzymes, enhancing Cas12a and Cas13a sensitivity by 50 000-fold and 10 000-fold, respectively. Furthermore, the superior biostability and structural simplicity of these linear probes endow SACA with excellent compatibility, facilitating precise in situ imaging of HPV16 and HPV18 mRNA in cervical cancer cells. This study not only advances the understanding of Cas enzyme regulation by chemically modified nucleic acids but also establishes a new paradigm for precise and efficient molecular diagnostics.
|

|
Scooped by
mhryu@live.com
Today, 3:57 PM
|
Metagenomics provides broad insights from microbial communities, but more biological relevant phenotypes are attributed to subtle changes at the strain-level rather than species. Despite development of several tools using different algorithms, resolving individual strains from short-read pair-end sequencing data remains challenging. We developed MetaStrainer, a tool capable of reconstructing strain genotypes from metagenomic data. Compared with existing approaches, MetaStrainer substantially increases genotype accuracy, correctly identifies the number of strains, and accurately estimates their relative abundances. Accuracy of reconstructed genotypes is robust to choice of mapping reference.

|
Scooped by
mhryu@live.com
Today, 3:32 PM
|
This study identified a putative cold-adapted acetylxylan esterase in Glutamicibacter soli Em07 via a metagenome-assembled genome. The gene encoding this enzyme was cloned and heterologously expressed in E. coli. SDS-PAGE gel electrophoresis analysis showed that the protein has a molecular weight of 33.24 kDa. Using 1-naphthyl acetate as a substrate, the enzyme activity was optimal at 20 °C and pH 9. Furthermore, the enzyme exhibited excellent cold adaptation, alkali resistance, and salt tolerance. It demonstrated OCP pesticide-degrading activity: 66.48% degradation of carbaryl, 92.14% of cypermethrin, and 97.78% of malathion, underscoring its strong potential in environmental remediation. Notably, this esterase emerged as the first to simultaneously possess cold adaptation, alkali resistance, and salt tolerance. These results positioned the enzyme as a promising candidate for bioremediation strategies in multiextreme environments. Further research will investigate its activity on other persistent organic pollutants.

|
Scooped by
mhryu@live.com
Today, 3:25 PM
|
In eukaryotic cells, essential functions are often confined within organelles enclosed by lipid membranes. Increasing evidence, however, highlights the role of membrane-less organelles (MLOs), formed through liquid-liquid phase separation (LLPS). MLO assemblies are typically initiated by driver proteins, which form a scaffold to recruit additional client molecules. By leveraging expanding MLO datasets and modern machine learning approaches, we developed LLPSight, an ML-based predictor of LLPS-driving proteins. The model was trained using rigorously curated datasets: a positive set of proteins experimentally confirmed to drive LLPS in vivo and a negative set of soluble, unstructured proteins not associated with LLPS. For the features, we employed a cutting-edge approach using embeddings from protein Language Models. LLPSight achieves the highest F1 score (0.885) among existing tools, enabling more efficient discovery of new LLPS drivers eagerly awaited by researchers for experimental validation. An additional key feature of LLPSight is its ability to perform proteome-wide analyses; application to the human proteome yielded promising targets. LLPSight can be obtained from authors upon request.

|
Scooped by
mhryu@live.com
Today, 12:49 PM
|
The rhizosphere microbiome plays a crucial role in the resistance to soilborne plant diseases. However, the principles needed to explain and predict which microbiota will be effective against soilborne pathogens are still lacking due to the complexity of the soil microbial community. We hypothesized that, independent of particular microbial strains, a high diversity is associated with, or increases the probability of, effective suppression. We tested this hypothesis by demonstrating that random combinations of rhizosphere microbial isolates, with the same bacterial diversity, had an equal impact on suppressing root diseases. The incidence of root rot was significantly reduced when soil bacterial diversity was high. We further investigated how high-diversity bacterial communities suppress root rot by constructing synthetic bacterial communities (SynComs). The results suggest that high bacterial diversity suppresses pathogens through mechanisms potentially including nutrient competition and the formation of physical barriers on the root surface. Our study highlights that high bacterial diversity is beneficial for suppressing soilborne plant diseases, offering a nonchemical and sustainable approach for crop disease management.

|
Scooped by
mhryu@live.com
Today, 11:40 AM
|
Translation is carried out by the most conserved assemblies in biology. Among these assemblies, the ribosome and RNase P are central players. These ancient ribonucleoprotein complexes achieved structural and functional maturity by the last universal common ancestor (LUCA) of life. In prior work, we reconstructed the evolutionary history of the ribosome using its three-dimensional structure, based on accretion and molecular fingerprints that date back to life’s earliest stages. Here, we extend our structural phylogenetic framework—based on the accretion model—to RNase P, a ribonucleoprotein responsible for processing pre-tRNAs. By sampling RNase P RNA (RPR) sequences and structures across phylogeny and partitioning them into RNA fragments based on insertion fingerprints, we characterize the state of RPR at LUCA and reconstruct the chronology of its emergence. The chronology reveals that RNase P, like the ribosome, accreted modular RNA elements over evolution, while preserving the structure of preexisting elements, thus maintaining a structural record. We used interactions with tRNA to link and unify the evolutionary trajectories of RPR and rRNA. These results support the view that RNase P and the ribosome coevolved as part of a functionally integrated system. The ancestral catalytic sites of rRNA and RPR formed by the same process, fusion of two stem–elbow–stem elements. Analysis of these two coevolving RNAs also suggests that some of their accreted elements share common ancestry. Application of the accretion model requires correct secondary structures and was successful for RPR only when the traditional secondary structure was corrected by reorganizing a pseudoknot.

|
Scooped by
mhryu@live.com
Today, 10:46 AM
|
Multicellular life arose in a world dominated by microorganisms, a reality that has imposed a constant and pervasive selective pressure on all subsequent complex organisms. The immune system has been historically defined by its role in pathogen clearance through resistance mechanisms. However, a complementary and equally critical strategy is to enable the peaceful and inevitable coexistence with microorganisms, allowing each host species to shelter a unique associated microbiome. The term tolerance holds multiple meanings in immunology, yet all underlie a balanced and cooperative host-microorganism relationship. Each represents a different aspect of how the immune system limits tissue damage while maintaining functionality in the presence of microbial or inflammatory stimuli. Using the intestinal mucosa as a paradigm, we explore how epithelial barrier integrity, toxin neutralization, tissue repair, and stress response underpin disease tolerance; how microbial exposure calibrates innate immunity via epigenetic and metabolic reprogramming (LPS tolerance); and how the gut microenvironment fosters the generation of tolerogenic antigen-presenting cells and microbe-specific regulatory T cells to enforce immunological tolerance. We further explore how the microbiota itself is a potent inducer of these tolerogenic pathways and highlight IL-10 as a major hub, connecting different tolerogenic circuits. Finally, we examine the hygiene hypothesis, arguing that lifestyle changes during the Anthropocene disrupt these finely tuned tolerance mechanisms, thereby contributing to the rising incidence of immune-mediated diseases. We posit that these tolerance programs are fundamental prerequisites for engendering host-microbiota symbiosis, a relationship forged over millennia of co-evolution and endangered in the contemporary world.

|
Scooped by
mhryu@live.com
Today, 12:21 AM
|
Marine plastic litter, including microplastics, has a profound impact on the ocean and its wildlife, and strategies to remove/eliminate it are needed. Microbial biodegradation, particularly by bacteria, offers a potential solution, where a link between hydrocarbon and plastic-degradation has been hypothesized. This study screened the plastic-degrading potential of 18 bacterial strains isolated from 1-month-old biofilms developed in three submerged plastic fishing nets (braided polyethylene (PE), braided nylon, thin nylon). In addition, three highly efficient hydrocarbon-degrading strains were also tested. Strains were cultivated on solid minimal media with fishing net small pieces (new/unused nets) added as a carbon source for 1 month, followed by tributyrin-agar assays to assess esterase/lipase activity. Eleven bacteria exhibited enhanced growth with net polymers, mainly from the genera Sulfitobacter, Rhodococcus, Bacillus, and Pseudomonas, and eight of which bacteria also demonstrated esterase/lipase activity. Then, genes encoding hydrocarbon or plastic-degrading enzymes (alkB and almA homologs, PETase-like enzymes) were screened by PCR in the 21 mentioned bacteria and in ca.100 other strains found in submerged nets biofilm. Amplification of the investigated genes was predominantly observed in Actinomycetes strains. Genome mining of six promising strains revealed hits with enzymes linked to degradation of synthetic polymers like polyethylene terephthalate, low-density PE and nylon. The workflow developed here enabled the selection of marine bacteria with plastic-degrading potential, sourced from biofilms of submerged plastic fishing nets and hydrocarbon-enriched environments.

|
Scooped by
mhryu@live.com
Today, 12:17 AM
|
From soil to the gut, communities composed of thousands of microbes perform functions such as carbon sequestration and immune system regulation. Here, we introduce a data-driven approach that explains how community function can be traced to just a few groups of microbes or genes. In gut communities, our neural-network based clustering algorithm correctly recovers known functional groups. In the ocean metagenome, it distills ~500 gene modules down to three sparse groups highlighting survival strategies at different depths. In soils, it distills ~4400 bacterial species into two groups that enter a mathematical model of nitrate metabolism. By combining interpretable ML with strain isolation and sequencing experiments, we connect the metabolic specialization of each group to community-wide responses to perturbations. This integrated approach yields simple structure-function maps of microbiomes, allowing the discovery of molecular mechanisms underlying human and environmental health. More broadly, we illustrate how to do function-informed dimensionality reduction in biology.

|
Scooped by
mhryu@live.com
March 3, 11:29 PM
|
Natural evolution is high-dimensional; organisms adapt to many pressures at once, across substrates, environments, and genetic backgrounds. Yet most directed evolution methods flatten this landscape to a single selection axis, hiding tradeoffs, and limiting what can be learned. Phage-assisted continuous evolution (PACE) is uniquely suited for multivariate selection because horizontal gene transfer couples genotype to propagation and allows the same phage lineage to traverse different selection environments. In practice, implementing this at scale has been prohibitive because each selection demands its own host culture, and every culture must be held for days to weeks within a narrow, infectable density window using continuously responsive bioreactors. In this work, TurboPRANCE is presented as an open-source, queueable robotic platform that integrates ~200 independently controlled turbidostats with 96 parallel PACE lagoons under closed-loop control. Each turbidostat operates as a fully separate unit that can be equilibrated and initiated on its own schedule, enabling asynchronous starts and sustained operation without intervention. Automated media formulation, programmable dosing, on-deck sterilization, and adaptive scheduling coordinate growth control with the changing needs of the robotic workflow, dynamically adjusting dilution and transfer timing around formulation, sampling, and handling steps to keep each culture at consistent infectable densities despite unpredictable method demands. Cultures can be multiplexed and titrated into lagoons at defined ratios, swapped in and out on a schedule, or kept fully separate across experiments, creating a combinatorial space of selection pressures and programs that is effectively unbounded. Additionally, to enable high-throughput evolutionary tracking that scales with TurboPRANCE, Nanopore long-read sequencing was combined with DeepVariant, a deep learning-based variant caller, enabling population-level tracking of evolving variants. The result is a system that generates high-resolution time-resolvable evolutionary trajectories and large parallel datasets spanning diverse selection regimes, yielding dense, multivariate training data to map and engineer complex fitness landscapes at scale.

|
Scooped by
mhryu@live.com
March 3, 10:53 PM
|
Traditional 16S rRNA gene and Internal Transcribed Spacer region amplicon sequencing provides only relative abundance, often leading to biased ecological interpretations. To overcome this limitation, we developed Accu16S/AccuITS, an absolute quantification method for bacterial and fungal amplicons based on synthetic internal spike-in DNA with known copy numbers. By adding internal standards prior to PCR and sequencing, absolute microbial abundances can be calculated using standard curve regression. Accu16S/AccuITS exhibits sensitivity and consistency comparable to quantitative PCR and is applicable to diverse sample types. A single sequencing run simultaneously yields relative abundance, total absolute abundance, and taxon-specific absolute abundance. Case studies across diverse ecosystems demonstrate that absolute quantification provides ecologically and functionally meaningful insights beyond those obtained from relative abundance analyses.

|
Scooped by
mhryu@live.com
March 3, 10:39 PM
|
Filamentous fungi, particularly Aspergillus species, play a crucial role in industrial biotechnology due to their exceptional protein secretion systems and robust metabolic adaptability. However, variability in heterologous protein production presents a major bottleneck. Here, we combine multi-omics analyses and synthetic biology to identify msnA as a key regulator of recombinant protein production in Aspergillus spp. Overexpression of msnA enhances secretion of diverse proteins (e.g., glycoside hydrolases, fluorescent proteins) by up to 2.8-fold in A. nidulans, A. niger, and A. oryzae, demonstrating functional conservation across Aspergillus species. Mechanistically, transcriptomic studies reveal that msnA reprograms cellular resource allocation by upregulating vesicle transport and fatty-acid degradation while downregulating secondary metabolism, thereby redirecting energy toward recombinant protein synthesis and secretion. Remarkably, the overexpression of msnA significantly improves the efficiency of recombinant protein secretion, even with lower transcriptional expression. In some species, this enhancement occurs despite a decrease in total secreted protein. Our research identifies msnA as a potential target gene for optimizing fungal cell factories, offering a framework to enhance recombinant protein production and increase the efficiency of Aspergillus-based systems in industrial applications.

|
Scooped by
mhryu@live.com
March 3, 6:48 PM
|
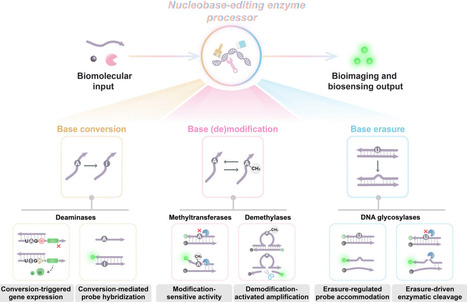
Nucleobases, the fundamental units of DNA and RNA, undergo diverse chemical modifications and lesions that regulate genomic stability, epigenetic states, and cellular functions. Nucleobase-editing enzymes, including deaminases, methyltransferases/demethylases, and DNA glycosylases, catalyze precise base conversions, modification/demodification, or excision reactions at high resolution without inducing double-stranded breaks. Originally studied in transcriptomic diversification, epigenetic regulation, and DNA repair, these enzymes are now increasingly repurposed as programmable actuators in biosensing and bioimaging applications. In this review, we first outline the catalytic principles of representative nucleobase-editing enzymes, emphasizing substrate recognition, reaction mechanisms, and physiological functions. We then highlight how these enzymes are integrated into biosensing and bioimaging modules across three major modes: nucleobase conversion, where site-directed deamination enables facile fluorescent protein reporter translation or specific DNA probe hybridization signals; nucleobase modification/demodification, where methylation and demethylation events regulate downstream selective enzymatic biocatalysis or programmble functional nucleic acid activation; and nucleobase erasure, where glycosylase-mediated base excisions facilitate specific probe accommodation or effient enzyme-catalyzed amplification. Such nucleobase-editing enzyme-driven systems offer high specificity, efficient amplification, and high compatibility with physiological conditions, enabling sensitive and spatiotemporally resolved monitoring of nucleic acids, proteins, and cellular processes. Finally, we discuss the advantages, challenges, and future directions of this emerging field. Advances in enzyme engineering, delivery strategies, and multiple circuitry integration are expected to yield next-generation bioanalytical platforms with improved precision, scalability, and clinical applicability. Collectively, these developments establish nucleobase-editing enzymes as a versatile molecular toolbox that bridges fundamental enzymology with applied biotechnology for diagnostics, therapeutic monitoring, and synthetic biology.
|
 Your new post is loading...
Your new post is loading...