 Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 2:00 AM
|
Bacteriocins are ribosomally synthesized antimicrobial peptides produced by various classes of bacteria, exhibiting broad-spectrum activity that makes them promising candidates for applications in food preservation and medicine. Their inherent stability under extreme pH, temperature, and salinity conditions further supports their functional versatility. However, the widespread industrial application of bacteriocins remains constrained by the low titres typically achieved during fermentation. Despite extensive efforts to optimize production using batch and fed-batch fermentation strategies, the resulting titres remain inadequate for economically viable large-scale manufacturing. This review aims to provide a comprehensive overview of novel process strategies developed over the past two decades to enhance bacteriocin yields during fermentation. One of the primary challenges is the inhibition of microbial growth due to the accumulation of lactic acid or the bacteriocin itself during production. To address this, in-situ product removal techniques—such as co-cultivation with lactic acid-consuming microorganisms, in-situ adsorption, filtration, and foam fractionation—have been explored, yielding notable improvements in bacteriocin titres. Additionally, stress-induced production strategies involving biological (e.g. co-culture with competing microbes), chemical (e.g. salinity and pH stress), and physical (e.g. agitation, temperature, and aeration stress) stimuli have also demonstrated success in enhancing bacteriocin synthesis. This review underscores the importance of these innovative fermentation approaches and highlights the need for further research focused on scaling up such processes. Advancing these strategies is critical to realizing the full potential of bacteriocins in food safety, antimicrobial therapy, and broader biomedical applications.

|
Scooped by
mhryu@live.com
Today, 1:52 AM
|
Chemical defenses are fundamental in organismal biology and widely used in medicine and agriculture. Plant defense chemistry evolves in response to various selective pressures, particularly herbivory, and theory has emphasized predicting toxin abundance and diversity. Here we test hypotheses about the evolution of structural innovation in chemical defense by combining molecular complexity metrics, metabolomics, molecular docking, and phylogenetic analyses, using milkweed cardenolides, steroidal glycosides that inhibit animal Na+/K+-ATPases. We identify the addition of a nitrogen–sulfur (N,S) heterocycle in highly substituted cardenolides as a major structural innovation that restores toxicity against coevolved natural enemies, such as the monarch butterfly. This toxicity is likely achieved by rigidifying the cardenolide scaffold and creating additional nonelectrostatic interactions within the Na+/K+-ATPase binding pocket, thereby enhancing binding affinity despite target-site resistance. Two biosynthetically distinct N,S-cardenolides, uscharin and labriformin, rank among the most complex structures in this chemical class and show divergent macroevolutionary histories: uscharin represents an ancestral character state with repeated losses, whereas labriformin has independently evolved multiple times in later-diverging lineages. This pattern across Asclepiadoideae indicates that the structural innovation evolved repeatedly, apparently limited by lineage-specific biosynthetic constraints among precursor pathways. N,S-cardenolides occur in over 75% of the 59 Asclepias species examined here, and species producing N,S-cardenolides exhibit greater cardenolide abundance, richness, metabolomic space, and toxicity against adapted organisms. More generally, structural innovation defines a distinct evolutionary axis in plant chemistry, enabling defense diversification and adaptive recovery of toxicity. Such innovations are predicted to build on existing molecular scaffolds in response to ecological challenges, here driven by coevolving specialist herbivores.

|
Scooped by
mhryu@live.com
Today, 1:44 AM
|
Resolving the biological and geological events that led to the origin of eukaryotes is an ongoing challenge in biology. A major step in the evolution of complex cellular life was the merger between an ancestral host cell and a bacterium (that became the mitochondrion) some two billion years ago. Recently, metagenomics has enabled the reconstruction of a broad diversity of genomes, referred to as the Asgard Archaea. The Asgards are monophyletic with eukaryotes on the tree of life. Asgards have an array of genes, previously thought exclusive to eukaryotes, involved in cellular trafficking, the ubiquitin system, endosomal sorting, and cytoskeleton formation, with growing evidence demonstrating the functions of these proteins mirror those in eukaryotes. This gene repertoire suggests that these Archaea are descendants of the archaeal host from which eukaryotes evolved. Increased sampling has revealed that Asgard lineages are metabolically versatile and play key roles in various ecosystems and uncovered evolutionary transitions between Archaea and eukaryotes, such as innovations in eukaryotic defense systems. The positioning of eukaryotes in the Asgards is debated, but eukaryotes appear to branch within the Heimdallarchaeia. Lineages within this group, particularly Hodarchaeales and Kariarchaeaceae, contain a broad repertoire of eukaryote-like traits, including high-energy yielding metabolisms. Observing and studying Asgard interactions with bacterial descendants of mitochondria in a modern setting will transform our understanding of the origin of complex cellular life.

|
Scooped by
mhryu@live.com
Today, 1:06 AM
|
Efficient genome writing in mammalian cells requires robust methods for integrating large DNA payloads. The previously described method mammalian Switching Antibiotic resistance markers Progressively for Integration (mSwAP-In) enables iterative, biallelic genome rewriting in mammalian stem cells with DNA payloads exceeding 100 kb. However, the lack of standardized vectors and certain technical constraints have limited its broader adoption. Here we present an improved plasmid toolkit designed to streamline the implementation of mSwAP-In. The toolkit includes two core vectors. pLP-TK (pCTC174) is a landing-pad plasmid compatible with Golden Gate assembly of genomic homology arms and supports both mSwAP-In and the recombinase-mediated cassette exchange method Big-IN. mSwAP-In MC2v2 (pKBA135) is a versatile Big DNA assembly and delivery vector that supports Gibson-based assembly and incorporates positive, negative, and fluorescent selection markers, as well as a backbone counterselection cassette to minimize unwanted plasmid integration. The vector architecture also enables propagation in yeast and bacterial hosts, inducible plasmid copy-number amplification in standard E. coli strains, and CRISPR/Cas9-mediated payload release through preinstalled guide RNA target sites. We further characterize the FCU1/5-FC counterselection system in mouse embryonic stem cells and define conditions that minimize its bystander toxicity. Finally, we provide a set of Cas9-gRNA expression plasmids optimized for common mSwAP-In applications. Together, these reagents constitute a standardized and experimentally validated toolkit that simplifies large-scale genome writing using mSwAP-In.

|
Scooped by
mhryu@live.com
March 16, 11:50 PM
|
Here, we introduce a new name into the bacterial energy conservation lexicon: facilitated fermentation. This name is necessary because the more familiar terms “respiration” and “fermentation” do not adequately describe how electron balancing is coupled to energy conservation for organisms that engage in this metabolism. Facilitated fermentation is when ATP is predominantly made via a substrate-level pathway that is redox-coupled to a terminal electron acceptor reduced outside of the cell. The coupling is often facilitated by an extracellular electron shuttle or outer membrane protein that shuttles electrons from the electron transport chain to the extracellular acceptor. Naming facilitated fermentation is timely because it has recently been demonstrated to support both growth and non-growth states in bacteria that are important in nature and disease. We hope that the introduction of this term will inspire future research to evaluate the extent of facilitated fermentation’s prevalence and impact in the microbial world and beyond.

|
Scooped by
mhryu@live.com
March 16, 11:38 PM
|
When environmental bacteria transition to laboratory conditions, a process termed domestication, the shift from the native habitat to a culture medium often reduces cell cultivability. Consequently, most bacteria remain uncultured using standard techniques, leaving the majority of their diversity unexplored. Here we introduce an enhanced domestication (EDEN) method for bacterial cultivation, which acclimatises environmental bacteria to culture media through a controlled and gradual exposure. To facilitate EDEN, we develop a 3D-printable microwell plate incorporating growth chambers integrated with a continuous-flow media reservoir. Using amplicon sequencing, we show that EDEN-acclimatised bacterial polycultures grow as distinct populations with significantly greater diversity and likely-uncultured taxa compared with standard cultivation methods. Similarly, EDEN-acclimatised bacterial monocultures show threefold greater diversity and tenfold more likely-uncultured taxa. EDEN also doubled the cultivability of agarose-encapsulated microcolonies. Finally, we demonstrate the utility of EDEN by isolating a previously uncultured bacterium exhibiting broad-spectrum antimicrobial activity against drug-resistant pathogens.

|
Scooped by
mhryu@live.com
March 16, 10:55 PM
|
The commercialization of cultivated meat remains limited by the high cost of cell culture medium. Here, we discuss how multi-omics characterization and computational modelling can enable the design of cell line-specific, serum-free medium using valorized sources, offering improved cost-efficiency and sustainability.

|
Scooped by
mhryu@live.com
March 16, 10:23 PM
|
Bacteriophages play a critical role in controlling bacterial populations, both in nature and as potential therapeutic agents. Their ability to replicate, compete against each other, and eradicate target cell populations is usually understood through a number of ‘life history parameters’, traditionally measured by population-level assays, which implicitly average the parameter’s value across a large number of infection events. Recent experiments suggest that bacteriophage life history parameters are subject to considerable stochasticity, raising the question of whether experimental and modelling efforts that do not account for this variability may overlook important factors in phage’s behavior, competitive fitness or therapeutic viability. Here, using agent-based simulations, we investigate the importance of stochasticity in lysis time and burst size of lytic bacteriophages in two common laboratory competition experiments: serial passage of well-mixed populations and plaque expansion across a bacterial lawn. We find that a phage’s analytic growth rate in isolation can be a poor predictor of its fitness advantage in simulated competition experiments. Specifically, when lysis times are tightly distributed, we identify a novel effect we name “population resonance”, through which a bacteriophage can display a significant fitness advantage over a competitor with a much greater growth rate in isolation. Our simulations also show that both serial passage and plaque expansion reward variability in lysis time more than expected, by increasing the phage resilience when resources are scarce.

|
Scooped by
mhryu@live.com
March 16, 6:29 PM
|
Biomolecular interactions lie at the core of cellular life, spanning diverse molecular modalities from small molecules to nucleic acids and proteins. Nevertheless, design strategies remain separated despite shared physicochemical principles of molecular recognition. Here we present AnewOmni, a unified generative framework trained on more than 5 million biomolecular complexes, that enables transferable molecular design across molecular scales by assembling chemically meaningful building blocks at atomic resolution. We further introduce programmable graph prompts to support user-defined chemical, topological, and geometric steering during generation, exploring hybrid and unconventional chemistries beyond canonical structures. We demonstrate that transferable learning of interaction patterns and physical constraints across molecular modalities is possible, via an atom-to-block latent space capturing both atomic details and structural priors. The framework successfully designed small molecules, peptides, and nanobodies targeting the challenging KRAS G12D switch II pocket, as well as orthosteric peptides and allosteric small-molecule inhibitors for PCSK9 in the absence of known binding site, achieving 23%-75% success with only low-throughput validation, bypassing modality-specific high-throughput screening. AnewOmni is the first to succeed in functional molecular design across all scales, from small chemical entities to large biologics, and represents a stepstone towards general molecular reasoning engines, advocating a generative foundation model for biomolecular interactions to enter regimes where data and human intuition remain limited.

|
Scooped by
mhryu@live.com
March 16, 6:07 PM
|
Genetically encoded biosensors are one of essential tools in biological research. They enable visualization of molecules of interest from the subcellular level to entire organism level in vivo and can be used to monitor presence of small molecules, gene expression, protein activity, and protein degradation. However, multiplexing fluorescent biosensors in plants is notoriously difficult due to signal bleed-through and strong autofluorescence from chlorophyll. In this study, we investigated the potential of multiplexing biosensors based on the selection of reporter fluorescent proteins. We characterized the emission spectra, fluorescence lifetimes, and relative brightness of diverse fluorescent proteins in plant leaves. We show that selected proteins exhibit comparable brightness, supporting their use in co-expression experiments and reliable quantification of individual signals. To separate overlapping signals, we applied two different linear unmixing approaches and compared them to results obtained without unmixing. We identified channel separation unmixing approach as the most suitable for biosensors. Additionally, we show how unmixing with the selected approach can be applied to separate autofluorescence and we validated this approach in virus-infected cells by following organelle dynamics in vivo. Overall, our work demonstrates that biosensors can be multiplexed, even when their emission spectra overlap.

|
Scooped by
mhryu@live.com
March 15, 4:21 PM
|
We present a living, synthetic bacterial cell made by transplanting a complete genome into a dead cell. After killing Mycoplasma capricolum cells by chemically crosslinking their genome with Mitomycin C (MMC), we installed synthetic Mycoplasma mycoides genomes into the resulting dead cells using Whole Genome Transplantation (WGT). During WGT, a synthetic donor genome is placed into a recipient cell, thereby reprogramming that cell to adopt a new genetic identity. WGT has only been demonstrated using species within one phylogenetic clade of Mollicutes bacteria. A major barrier to expanding WGT to diverse bacterial species has been the inability to inactivate the recipient genome, leading to false positive transplants due to homologous recombination of antibiotic resistance markers from the donor genome into the recipient cell genome. Here, we address this key limitation by removing reliance on an antibiotic resistance marker to select for transplants; recipient cells are dead unless revived by the installation of a new genome. Our work demonstrates a general approach to fully inactivate the recipient cell genome, reports the first living synthetic bacterial cell constructed from non-living parts, and advances WGT for building engineered or synthetic cells for diverse applications.

|
Scooped by
mhryu@live.com
March 15, 4:08 PM
|
Phenotypic heterogeneity within isogenic bacterial populations represents a fundamental adaptation strategy that enables pathogen survival across the selective pressures of host infection. Rather than uniformly responding to environmental challenges, bacterial populations diversify through mechanisms including phase variation, stochastic gene expression, asymmetric cell division, and intercellular communication, generating functionally specialized subpopulations that operate through bet-hedging and division of labor frameworks. This review synthesizes recent advances showing how host-derived signals tune bacterial switching dynamics and deterministically partition populations into discrete phenotypic states. These functionally specialized subpopulations have clinical implications, with antibiotic-tolerant persisters representing one example of how phenotypic heterogeneity drives treatment failure and enables infection recurrence, underscoring the need to understand and target this fundamental aspect of bacterial pathogenesis.

|
Scooped by
mhryu@live.com
March 14, 11:32 PM
|
The acid activity of enzymes, characterized by the minimum pH at which enzymes remain active (pHmin), is crucial for industrial applications in acidic environments. However, the rational design of acid-active enzymes remains challenging due to limited understanding of sequence-structure-activity relationships under acidic conditions. Here, we propose ACENet, a graph neural network that predicts enzyme pHmin by integrating surface features of protein structures with evolutionary representations derived from the large-scale protein language model ESM-2. ACENet achieved a Pearson correlation coefficient of 0.85 on the test dataset, significantly outperforming other deep learning baseline models and maintains stable pHmin predictions under various conditions. Even on a subset of the dataset with less than 20% homology, the PCC remains above 0.5, with an RMSE (Root mean square error) less than 1.4. ACENet also present excellent performance in the annotation of pHmin for homologous proteins and the predictive screening of minimal active pH in protein mutants. Remarkably, ACENet could identify the catalytic region as key determinants of acid activity through residue-level interpretability analysis. Overall, ACENet accelerates the development of highly efficient biocatalysts for diverse applications where acidic conditions predominate.
|

|
Scooped by
mhryu@live.com
Today, 1:54 AM
|
Ecosystems and human health are at serious risk due to the extensive application of pesticides in the agricultural system for controlling pests and diseases. The use of microbial consortia (MicroCons) has emerged as a promising solution for the remediation of pesticide-contaminated soil, offering a sustainable and eco-friendly alternative to physical and chemical methods; however, a systematic review on this aspect is still lacking. This comprehensive review provides an in-depth analysis of the current knowledge on microbial consortia-based remediation of pesticides in agricultural soil. Efficacy of single-strain vs multiple strains in MicroCons have been discussed to unravel the workload distribution between microbial strains in pesticide degradation. We also discuss the design and optimization of microbial consortia for remediation, highlighting the role of advanced tools and the mechanisms of MicroCons action. Furthermore, emerging trends and future directions in the field, including the potential of synthetic biology, machine learning (ML), and artificial intelligence (AI) are also covered. This review aims to critically expand the mechanistic understanding of how microbe-mediated remediation strategies might reduce pesticide phytotoxicity, enhance crop production in pesticide-stressed soils, and inspire future research and practices in MicroCons-based remediation to achieve the Sustainable Development Goals (SDGs).

|
Scooped by
mhryu@live.com
Today, 1:48 AM
|
Targeted protein degradation is a promising strategy for drug discovery, but designing effective PROTACs remains challenging, especially for proteins without well-defined binding sites. Current methods rely on modifying linkers between fixed ligands, which limits the diversity and innovation of the overall molecular architecture of PROTAC. Here, we introduce DeepDegradome, an AI-powered method that automates the structure-aware design of both small-molecule ligands and PROTACs. It employs a large fragment library constructed from public databases and applies an in-house docking method (iFitDock) to obtain initial binding fragments. DeepDegradome builds ligands by assembling these fragments based on the shape and physicochemical features of the target protein pocket. It can further construct PROTACs from these generated ligands, eliminating the dependency on predefined warheads or E3 ligands. Compared to other AI models, DeepDegradome produces more valid, drug-like molecules with higher predicted binding affinity. We demonstrate DeepDegradome’s effectiveness by designing and validating multiple potency inhibitors and PROTACs for two protein targets: WDR5 and CDK9. One synthesized compound showed excellent agreement between predicted and actual binding conformation confirmed by X-ray crystallography. By combining ligand and PROTAC design in one system, DeepDegradome offers a scalable and reliable tool for discovering new drugs against protein targets.

|
Scooped by
mhryu@live.com
Today, 1:13 AM
|
Pyomelanin has been extensively applied in various fields. However, the yield of pyomelanin isolated from natural producers is low. Engineering microbial biosynthesis is considered a sustainable and economically feasible alternative. We utilize engineered Komagataella phaffii to increase the yield of homogentisic acid by 66-fold by balancing three different biosynthetic modules and further synthesizing pyomelanin by oxidative polymerization. During this process, we establish a pyomelanin color screening system and apply this system to perform directed evolution of 3-deoxy-D-arabinoheptulosonate 7-phosphate synthase and semirational design modification of hydroxyphenylpyruvate dioxygenase. We report the difference of pyomelanin between oxidative polymerization under alkaline conditions and that under laccase treatment. After high-density fermentation in a 5-L fermenter for 204 h, strain Pyo29 achieve the highest titer (70.5 ± 0.7 g/L). Finally, we demonstrate the potential application value of pyomelanin. This study provides a feasible solution for high-yield pyomelanin biosynthesis with substantial industrial production. Pyomelanin is resistant to UV light, and has antibacterial and anti-inflammatory effects, however, the yield of pyomelanin isolated from natural producers is low. Here the authors engineer Komagataella phaffii to produce 70.5 g/L and isolate the product.

|
Scooped by
mhryu@live.com
Today, 12:48 AM
|
Prime editing (PE) and base editing (BE) are potent gene-editing techniques that avoid introducing exogenous DNA donors and double-strand breaks, but their large molecular size hinders efficient delivery and widespread application. Here, we engineer and evolve compact obligate mobile element-guided activity (OMEGA)-based PE and BE systems, termed OMEGA-PE and OMEGA-adenine base editor, which achieve superior editing efficiency and versatility compared with existing PE and BE tools in E. coli. Notably, OMEGA-PE enables rapid, programmable targeted mutagenesis with outstanding specificity and efficiency compared with current tools. Moreover, it exhibits excellent compatibility with several high-throughput screening platforms and can be used to enhance the efficiency of cofactor regeneration and protein translation. These groundbreaking advancements offer significant potential for gene editing, directed evolution, metabolic engineering, and protein expression optimization. Combining compact size with high performance, our OMEGA-based systems address key bottlenecks in current tools, offering new possibilities for basic research and industrial biotechnology.

|
Scooped by
mhryu@live.com
March 16, 11:44 PM
|
Gene expression in mycobacteria relies on both integrative and episomal E. coli-mycobacterium shuttle vectors. However, existing vectors have limited flexibility for adding epitope tags, selectable markers, and fluorescent reporters. To overcome this, we created a modular toolkit consisting of 32 integrative and eight episomal E. coli-mycobacterium shuttle vectors. Each vector features N- or C-terminal tags: SBP-HA-S, 3×-FLAG, GFP, and mCherry. The integrative vectors contain four different phage systems (L5, Giles, MS6, and Tweety), each with a distinct antibiotic resistance marker, enabling stable integration at various chromosomal sites. We validated the system in Mycobacterium smegmatis by expressing RbpA, an RNA polymerase-binding protein. Fluorescent tags confirmed Wag31’s polar localization, whereas co-expression with FtsZ illustrated their distinct localizations. The system’s flexibility was further shown by co-expressing four Mycobacterium tuberculosis sigma factors (SigA, SigC, SigG, and SigH) simultaneously. Additionally, an SBP-HA-S tandem affinity tag facilitated efficient two-step purification of RbpA with RNA polymerase and related regulators, reducing background compared to FLAG pull-down. This toolkit provides a versatile, reliable platform for gene complementation, protein localization, multi-protein co-expression, and native complex purification, supporting advanced genetic and cell biological studies in mycobacteria.

|
Scooped by
mhryu@live.com
March 16, 11:03 PM
|
Lignin forms the polyphenolic network in wood that enables trees to stand upright, transport water and ions, and resist microbial attack. Although its structural complexity is often seen as a limitation, its inherent multifunctionality offers opportunities. By strategically harnessing these features, new directions for advanced lignin-based materials can emerge.

|
Scooped by
mhryu@live.com
March 16, 10:37 PM
|
Using engineering biology to perform complex chemical synthesis offers a sustainable alternative to traditional processes that rely on finite fossil resources. A growing opportunity within this field lies in reclaiming carbon embedded in industrial and post-consumer waste—carbon otherwise lost to landfill, incineration or pollution. Here we report the bio-upcycling of poly(ethylene terephthalate) (PET) plastic waste into levodopa (l-DOPA), a frontline medication for Parkinson’s disease, using engineered E. coli. Two key bottlenecks—substrate import and feedback inhibition by the intermediate protocatechuate—were addressed through heterologous transporter expression and functional pathway separation across two microbial strains. To further improve sustainability, and as a proof-of-concept, Chlamydomonas reinhardtii was used to capture CO2 released during catechol generation. The resulting bioprocess operates under mild, aqueous conditions and achieves high l-DOPA titres (5.0 g l−1), with isolated product obtained at preparative scale from both industrial PET waste and a single post-consumer plastic bottle. This work demonstrates how engineering biology can transform plastic-derived aromatic monomers into high-value pharmaceuticals for the treatment of neurological disease in humans. The production of high-value chemicals can involve energy-intensive processes, necessitating sustainable production strategies. Here the authors present a circular bioeconomy approach, upcycling plastic waste through microbial conversion into levodopa, a medicine for Parkinson’s disease.

|
Scooped by
mhryu@live.com
March 16, 6:37 PM
|
Delivery of biomolecules into plant vascular tissues remains a barrier to managing diseases caused by insect vector-borne pathogens and to modifying phenotypes of established perennial crops. Inspired by the vascularized growth of crown galls induced by Agrobacterium tumefaciens, we repurposed the bacterium’s plant growth regulator (PGR) genes to engineer autonomously dividing, transgene-expressing plant cell structures termed symbionts. A plant transformation vector (pSYM) incorporating the IaaM, IaaH, Ipt and gene5 cassette from A. tumefaciens strain C58 together with a gene of interest on the same transfer DNA was delivered to stems of herbaceous and woody dicots using disarmed A. tumefaciens strain EHA105. Symbiont morphology, vascular differentiation, transgene expression, molecular mobility and protein secretion were evaluated using microscopy, fluorescent reporters, dye tracing, RNA silencing assays and mass spectrometry-based proteomics. pSym inoculation reproducibly generated symbionts across diverse host plant species that were vascularly integrated into their host plants and transgene expression ranging from heterogeneous niches to more uniform patterns. Small molecules moved between symbionts and host vascular tissues, whereas larger proteins exhibited more restricted mobility. Post-transcriptional gene silencing signals moved freely throughout the symbiont and slightly into adjacent stem tissue. Under tested field and greenhouse conditions in potato and tomato, respectively, gall or symbiont formation had no negative impacts on plant growth or tuber and fruit yield. In vitro, symbiont cultures abundantly secreted recombinant protein into surrounding media. Together, these results establish symbionts as a modular, plant bioengineering platform capable of producing and potentially delivering biomolecules without modifying the host plant genome, providing a foundation for vascular-targeted therapeutics and phenotype modulation in crops.

|
Scooped by
mhryu@live.com
March 16, 6:13 PM
|
Protein complexes are central to cellular function, but experimental determination of their structures remains challenging. Structure prediction methods require prior knowledge of stoichiometry - the number of copies of each protein entity within a complex. Current approaches rely on computationally expensive brute-force methods that run structure prediction on multiple stoichiometry combinations, often with limited accuracy. We introduce Stoic, a method that uses protein language model embeddings to predict protein complex stoichiometry. Our approach learns to identify interface residues that participate in protein-protein interactions, rather than relying on global sequence features. By integrating these interface-aware embeddings into a graph neural network, Stoic achieves fast and accurate stoichiometry prediction for both homomeric and heteromeric targets. Availability: Source code for inference and training along with web versions are available in the repository at https://github.com/PickyBinders/stoic.

|
Scooped by
mhryu@live.com
March 15, 4:36 PM
|
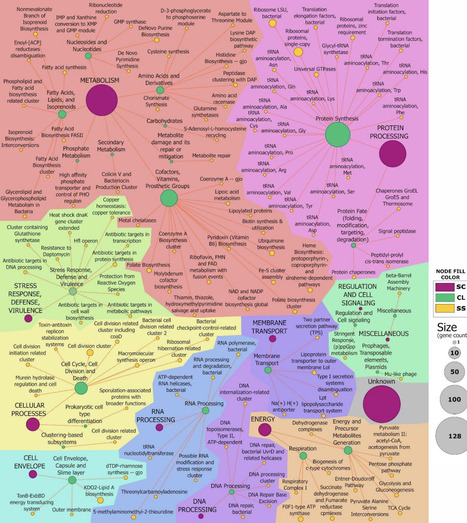
Pseudomonas chlororaphis ATCC 9446 is a non-pathogenic rhizobacterium with biotechnological relevance as a biocontrol agent and a promising chassis for synthetic biology. Understanding which genes are strictly required for survival is fundamental to both bacterial physiology and chassis engineering. Here, we generate a genome-wide map of genetic essentiality for P. chlororaphis using high-density Random Barcoded Transposon Mutagenesis (RB-TnSeq). To convert gene-level annotation into biological insight, we layered functional assignments from complementary annotation pipelines, integrating orthology/domain classifiers, ontology mapping, protein export, and delineation of secondary metabolism. These overlays reveal that essentiality concentrates in canonical information processing, envelope biogenesis, and central energy/cofactor and nucleotide metabolism, while large genomic regions are non-essential and therefore represent safe candidates for streamlining and pathway installation. Mapping essentiality onto biosynthetic gene clusters (BGCs) shows that most pathways are dispensable, but some essential genes co-localize, clarifying boundaries for safe editing around BGC loci. Comparison of experimentally determined essential genes with in silico predictions across additional P. chlororaphis genomes show strong overall agreement. Conversely, 32 essential gene orthogroups were found to be conserved across most genomes, yet were classified as non-essential by a prediction algorithm. Together, the resolved essential genome and its integrative functional interpretation provide a durable reference for P. chlororaphis biology and a functional blueprint that can be leveraged for rational streamlining in agricultural, biocontrol and industrial biotechnology.

|
Scooped by
mhryu@live.com
March 15, 4:17 PM
|
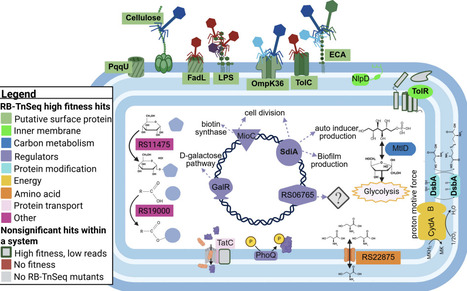
Bacteriophages (phages) are being cataloged at unprecedented rates and are recognized as key players in nutrient and energy cycling across ecosystems. Yet, the biological determinants of phage-host specificity and infection success remain poorly understood. Here, we used a randomly barcoded, genome-wide, loss-of-function transposon mutant library (RB-TnSeq) of Klebsiella sp. M5al, a plant-associated, nitrogen-fixing rhizobacterium, to identify bacterial genetic determinants necessary for the infection of 25 double-stranded DNA phages spanning five viral families. This approach identified 42 bacterial genes associated with phage infection, encompassing genes involved in receptor biosynthesis, regulatory pathways, electron transport, and unknown functions. Disruption of genes involved in receptor biosynthesis, such as glycosyltransferases involved in lipopolysaccharide (LPS) production, conferred cross-resistance to half of the phages, while intracellular gene disruptions had phage-specific effects. Taxonomically, we found significant differences in RB-TnSeq profiles across phage families and genera. While bacterial gene requirements are generally clustered by phage genus, we also observed notable divergences within genera. These differences were associated with variation in receptor usage, likely driven by divergence in tail fiber or other host recognition proteins. In some cases, differential reliance on bacterial intracellular factors suggested that even closely related phages may depend on distinct infection strategies, potentially involving phage-specific genes of unknown function. Our findings reveal key genetic dependencies shaping phage-host interactions and offer a framework for predicting infection outcomes in natural and applied settings.

|
Scooped by
mhryu@live.com
March 15, 4:05 PM
|
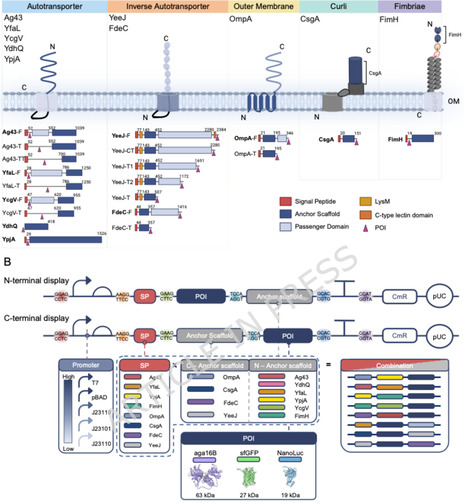
Bacterial cell surface display is a versatile platform technology for synthetic biology applications. A successful display platform often requires ad hoc testing of anchor scaffold types and orientations, as well as target protein types and orientations. To facilitate screening for the optimal display platform, this study designed and constructed a modular, combinatorial E. coli surface display toolkit. The modular toolkit not only screens for the optimal combination of diverse anchor proteins to display various bioparts but also benchmarks the performance of multiple surface display platforms in E. coli simultaneously. The design of the modular toolkit was based on five representative types of anchor scaffolds encompassing autotransporters, outer membrane proteins, curli-associated proteins, and fimbriae-associated proteins. Both amino- and carboxy-terminal display was examined across 20 designs of full or truncated scaffolds, each combined with multiple signal peptides, promoters, and terminators. The combinatorial toolkit was validated and optimized using three display reporters with different sizes and features—sfGFP (fluorescence), agarase (enzymatic activity), and NanoLuc (luminescence). The optimal display combinations were primarily dependent on the orientation of anchor scaffolds, which varied with reporter properties. Strong promoters, such as T7, differentially affect surface display activity on substrate-dependent (agarase, NanoLuc) and substrate-independent (sfGFP) proteins, highlighting expression tuning as a critical parameter for surface display optimization. Using this modular kit, two previously uncharacterized anchor scaffolds (YdhQ and YcgV) were identified as functional for surface display. Finally, the applicability of the toolkit across multiple E. coli genetic backgrounds was confirmed, supporting its utility for diverse biotechnological applications.
|
 Your new post is loading...
Your new post is loading...



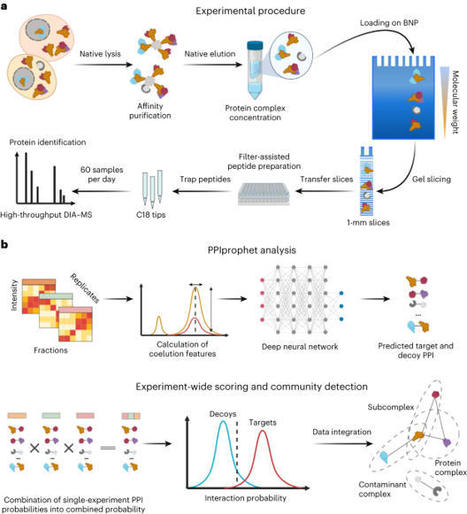





























ppi methods, As an alternative to AP–MS protein correlation methods exemplified by size-exclusion chromatography (SEC–MS)7,8 and blue-native-PAGE (BNP)9 coupled to MS10 have been introduced. They fractionate native complexes by their hydrodynamic radius and size, respectively, and each fraction is subsequently profiled by MS. Resulting cofractionation profiles are then used to define protein complexes