 Your new post is loading...

|
Scooped by
mhryu@live.com
March 15, 4:36 PM
|
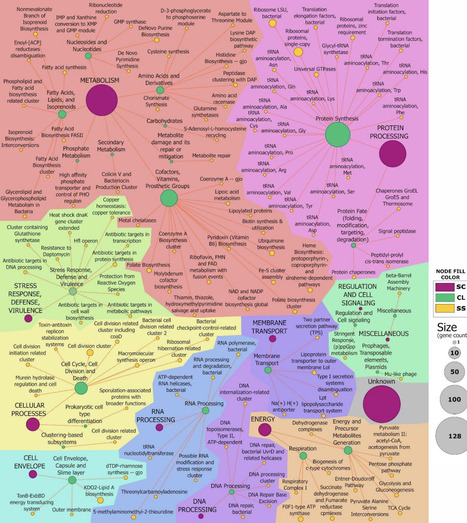
Pseudomonas chlororaphis ATCC 9446 is a non-pathogenic rhizobacterium with biotechnological relevance as a biocontrol agent and a promising chassis for synthetic biology. Understanding which genes are strictly required for survival is fundamental to both bacterial physiology and chassis engineering. Here, we generate a genome-wide map of genetic essentiality for P. chlororaphis using high-density Random Barcoded Transposon Mutagenesis (RB-TnSeq). To convert gene-level annotation into biological insight, we layered functional assignments from complementary annotation pipelines, integrating orthology/domain classifiers, ontology mapping, protein export, and delineation of secondary metabolism. These overlays reveal that essentiality concentrates in canonical information processing, envelope biogenesis, and central energy/cofactor and nucleotide metabolism, while large genomic regions are non-essential and therefore represent safe candidates for streamlining and pathway installation. Mapping essentiality onto biosynthetic gene clusters (BGCs) shows that most pathways are dispensable, but some essential genes co-localize, clarifying boundaries for safe editing around BGC loci. Comparison of experimentally determined essential genes with in silico predictions across additional P. chlororaphis genomes show strong overall agreement. Conversely, 32 essential gene orthogroups were found to be conserved across most genomes, yet were classified as non-essential by a prediction algorithm. Together, the resolved essential genome and its integrative functional interpretation provide a durable reference for P. chlororaphis biology and a functional blueprint that can be leveraged for rational streamlining in agricultural, biocontrol and industrial biotechnology.

|
Scooped by
mhryu@live.com
March 15, 4:17 PM
|
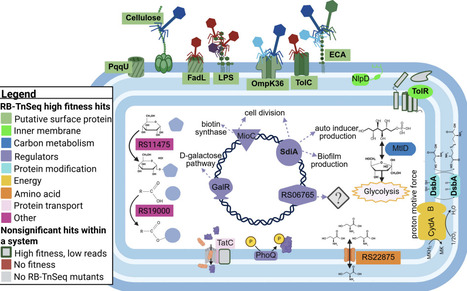
Bacteriophages (phages) are being cataloged at unprecedented rates and are recognized as key players in nutrient and energy cycling across ecosystems. Yet, the biological determinants of phage-host specificity and infection success remain poorly understood. Here, we used a randomly barcoded, genome-wide, loss-of-function transposon mutant library (RB-TnSeq) of Klebsiella sp. M5al, a plant-associated, nitrogen-fixing rhizobacterium, to identify bacterial genetic determinants necessary for the infection of 25 double-stranded DNA phages spanning five viral families. This approach identified 42 bacterial genes associated with phage infection, encompassing genes involved in receptor biosynthesis, regulatory pathways, electron transport, and unknown functions. Disruption of genes involved in receptor biosynthesis, such as glycosyltransferases involved in lipopolysaccharide (LPS) production, conferred cross-resistance to half of the phages, while intracellular gene disruptions had phage-specific effects. Taxonomically, we found significant differences in RB-TnSeq profiles across phage families and genera. While bacterial gene requirements are generally clustered by phage genus, we also observed notable divergences within genera. These differences were associated with variation in receptor usage, likely driven by divergence in tail fiber or other host recognition proteins. In some cases, differential reliance on bacterial intracellular factors suggested that even closely related phages may depend on distinct infection strategies, potentially involving phage-specific genes of unknown function. Our findings reveal key genetic dependencies shaping phage-host interactions and offer a framework for predicting infection outcomes in natural and applied settings.

|
Scooped by
mhryu@live.com
March 15, 4:05 PM
|
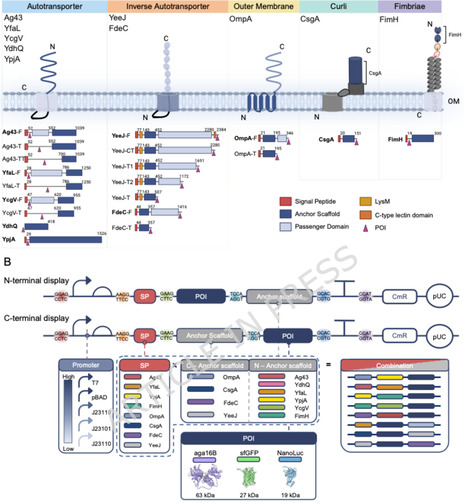
Bacterial cell surface display is a versatile platform technology for synthetic biology applications. A successful display platform often requires ad hoc testing of anchor scaffold types and orientations, as well as target protein types and orientations. To facilitate screening for the optimal display platform, this study designed and constructed a modular, combinatorial E. coli surface display toolkit. The modular toolkit not only screens for the optimal combination of diverse anchor proteins to display various bioparts but also benchmarks the performance of multiple surface display platforms in E. coli simultaneously. The design of the modular toolkit was based on five representative types of anchor scaffolds encompassing autotransporters, outer membrane proteins, curli-associated proteins, and fimbriae-associated proteins. Both amino- and carboxy-terminal display was examined across 20 designs of full or truncated scaffolds, each combined with multiple signal peptides, promoters, and terminators. The combinatorial toolkit was validated and optimized using three display reporters with different sizes and features—sfGFP (fluorescence), agarase (enzymatic activity), and NanoLuc (luminescence). The optimal display combinations were primarily dependent on the orientation of anchor scaffolds, which varied with reporter properties. Strong promoters, such as T7, differentially affect surface display activity on substrate-dependent (agarase, NanoLuc) and substrate-independent (sfGFP) proteins, highlighting expression tuning as a critical parameter for surface display optimization. Using this modular kit, two previously uncharacterized anchor scaffolds (YdhQ and YcgV) were identified as functional for surface display. Finally, the applicability of the toolkit across multiple E. coli genetic backgrounds was confirmed, supporting its utility for diverse biotechnological applications.

|
Scooped by
mhryu@live.com
March 14, 10:39 PM
|
With an increasing global cancer burden, the regulatory function of the human microbiome and its metabolites in tumor epigenetics has garnered significant interest. Microbial metabolites are not merely passive byproducts but serve as signaling molecules and epigenetic modulators, contributing to tumor progression through multiple overlapping pathways. Short-chain fatty acids (SCFAs) such as butyrate directly inhibit histone deacetylases to reactivate tumor suppressor genes, while secondary bile acids (BAs) induce gene silencing via DNA methylation remodeling by altering the FXR/TGR5 signaling pathway. Folate and vitamin B12 serve as substrates for DNA and histone methylation through one-carbon metabolism. A complex bidirectional feedback loop exists between microbial metabolism and tumor epigenetics: reprogramming driven by hypoxia or oncogenes alters metabolite flux, generating molecules such as lactate and succinate that not only remodel chromatin and the tumor microenvironment (TME) but also selectively promote the growth of metabolically adapted microbial species, thereby reinforcing epigenetic dysregulation. Despite growing mechanistic insights, establishing causality and correlating spatiotemporal dynamics and dose responses within the highly heterogeneous TME remain major challenges. Data integration across multi-omics remains limited by methodological and computational constraints. Resolving these issues will be critical for understanding the microbe–metabolite–epigenetic axis and advancing personalized precision oncology.

|
Scooped by
mhryu@live.com
March 14, 10:28 PM
|
Top-down trophic interactions are major drivers of microbiome dynamics, yet their outcomes are difficult to predict and their consequences for pathogen control remain unclear. We combine synthetic bacterial communities of varying complexity with field studies and microcosm assays to test whether microbivorous nematodes reorganize microbiomes to suppress soilborne disease. Field studies show stronger nematode-microbe associations around healthy plants, and microcosm assays confirm that nematode presence produces stable suppression, whereas microbe-only communities collapse under pathogen invasion. Nematode predation depletes non-preferred bacterial taxa and enriches metabolically versatile taxa within Proteobacteria, increasing community-level antagonistic potential and promoting complementary resource-use interactions linked to pathogen inhibition, yielding suppression beyond individual or pairwise effects. A minimal four-component feedback loop linking a nematode predator, plant pathogens, and two plant-associated bacteria with complementary functions accounts for the emergent outcome. Together, these results reveal an animal-mediated pathway of microbiome assembly that enhances resistance to pathogen invasion and provide a trophically informed framework for designing stable, disease-suppressive microbiomes in agriculture. Top-down trophic interactions are critical drivers of microbiome dynamics but are difficult to predict. Here, the authors demonstrate that nematode-driven shifts in microbiomes enhance pathogen suppression and promote plant health through a feedback loop involving nematodes and bacteria.

|
Scooped by
mhryu@live.com
March 14, 5:19 PM
|
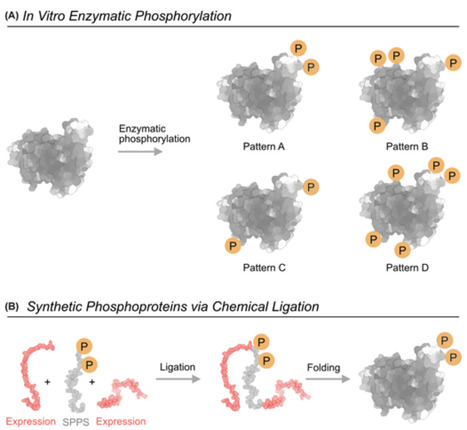
Protein phosphorylation is one of the most common and versatile regulatory mechanisms in cells. Most human proteins are phosphorylated at multiple sites, giving rise to large numbers of possible phosphorylation patterns. Each phosphorylation pattern can lead to a different functional or pathological outcome. Yet, linking defined phosphorylation patterns to specific biological functions remains a major experimental challenge. In this review we describe the main strategies to study phosphorylation patterns at the protein and domain levels and highlight how they complement each other. We first discuss cellular approaches, including phosphomimetics, kinase-based assays, and genetic code expansion, which allow working in a native environment but have their significant drawbacks. We then describe in vitro methods, such as enzymatic phosphorylation and semi-synthetic phosphoproteins generated by ligation, which afford mechanistic insights but result in low yields and are difficult to scale for producing libraries. We focus on synthetic phosphopeptide libraries as tools that offer precise control over the number and position of phosphosites and are uniquely suited for systematic mapping of phosphorylation patterns. This comes at a price of not working at the protein level, but rather at the domain level. Peptide libraries are often used for preliminary identification of key phosphorylations, later studied in detail at the protein level. We conclude that ideally more than one method should be used and that these methods should not be viewed as competing but rather as complementary. A combined use of several of these approaches provides a practical toolbox for dissecting how phosphorylation patterns regulate protein behavior.

|
Scooped by
mhryu@live.com
March 14, 5:12 PM
|
Optimization of the design-build-test-learn cycle remains a bottleneck for developing and efficiently manufacturing the next generation of bioproducts. To address this challenge, research has traditionally focused on either data-driven or mechanistic modelling, but the emerging consensus in the field highlights that their strategic integration, often termed hybrid modelling, offers significant advantages, particularly for complex systems where data is sparse. Here, we introduce a hybrid framework where these two methodological lineages are combined to frame development parameters within the metabolism of a production system, therefore achieving mechanism-informed predictions for follow-up experiments. We present three scenarios that exemplify how this framework can be leveraged to guide and accelerate the development of novel bioprocesses, even with small datasets available. We validate our framework by applying it to two heterologous peptide production scenarios in E. coli, explored through commonly relevant experimental factors such as inducer concentration and production strain. Using our framework, we identify key metabolic pathways and reactions that contribute to productivity and whose activity is modified by individual experimental factors like temperature and plasmid used. Furthermore, we show that the biological patterns extracted from this hybrid approach can complement the experimental design, informing predictive models of process performance. Our approach is general and can be tailored to a large array of processes, and thus holds potential for boosting both proof-of-concept and industrial projects, contributing to more efficient and sustainable biomanufacturing.

|
Scooped by
mhryu@live.com
March 14, 5:07 PM
|
Quantifying bacteria’s growth rates is essential for understanding their ecological roles and for building predictive models in environmental and clinical settings. Peak-to-trough ratios (PTRs) derived from shotgun metagenomes offer a culture-independent proxy for in situ growth rates of bacterial species, yet their reliable computation remains challenging. We introduce Pilea (https://github.com/xinehc/pilea), an alignment-free, sketching-based method that incorporates statistical models for robust PTR estimation. Pilea achieves speed improvements over existing methods while also enhancing accuracy, as demonstrated on both simulated and real datasets. By scaling efficiently to comprehensive reference collections such as the Genome Taxonomy Database (GTDB), Pilea enables large-scale analyses of bacterial growth dynamics across biomes, unlocking new insights for ecological research.

|
Scooped by
mhryu@live.com
March 14, 4:59 PM
|
Horizontal gene transfer (HGT) generates genetic variation in populations across all domains of life; however, most studies focus on individual transfers and functional information derived therefrom. This is useful but does not consider DNA transfer more broadly, that is, nongene transfers, donor–recipient dynamics, or trends and background levels that may help infer ecological information. Here, we review the mechanistic underpinnings of DNA transfer, literature from diverse fields that addresses HGT on a community basis and the associated methodological challenges, and propose a framework for conceptualizing the process of DNA transfer, highlighting DNA mobility as a feature of community ecology and DNA itself as a public good. These ideas coalesce to support DNA transfer as a fundamental ecological phenomenon that remains largely unmeasured.

|
Scooped by
mhryu@live.com
March 14, 4:50 PM
|
The stability of synthetic gene circuits is limited by cellular growth and division. Zhang et al. demonstrated how engineered condensates of transcription factors can stabilize gene expression and enhance bio-based production. This work establishes the spatial organization of transcription factors through phase separation as a powerful strategy for robust gene expression.

|
Scooped by
mhryu@live.com
March 14, 4:45 PM
|
Robots play an ever-expanding role in society by performing a broad range of tasks. However, there are growing concerns about their environmental sustainability, as many conventional robotic systems rely on materials that are neither renewable nor degradable. Consequently, significant efforts are being made to develop eco-friendly robots built from sustainable and biodegradable materials. In this context, plants represent a promising direction, as the biomaterials composing plants are biodegradable, and their inherent multifunctionality as living organisms, including sensing, actuation, energy harvesting, and self-healing, makes them strong candidates for realizing biodegradable robotic systems. Moreover, they are abundant and renewable resources. Recent studies have demonstrated plant-based robotic systems that harness some of these features, helping to establish plant robotics as an emerging research field. Among the many functions plants offer, actuation is pivotal, as it enables physical robotic motion, such as locomotion and grasping, which substantially broadens the potential applications of plant robots. Focusing on plant movement, this article reviews key plant species and their behaviors through the perspective of actuation characteristics. It also examines the current landscape of plant-based robotic systems and outlines future research directions in this rapidly growing field.

|
Scooped by
mhryu@live.com
March 14, 4:21 PM
|
Physiologically relevant biosensors are in increasingly high demand, yet existing ones are severely limited in the number and type of biomarkers that are detected. The lack of biorecognition elements for most medically relevant biomarkers restricts the development of next generation single and continuous use monitors. Over billions of years, microbes have evolved a vast array of proteins to sense and metabolize small molecules, including those pertinent to human health. Of particular interest to us is the identification and subsequent integration of new microbial redox enzymes into electronic biosensors building off the established electrochemical technology of the continuous glucose monitor. Here we deploy genomic screening to identify analyte specific redox enzymes for biosensor development. As a proof of concept, we report the first electrochemical enzyme-based nicotine biosensor from a novel microbial enzyme, and use a variant with improved catalytic performance to enhance sensor performance. The biosensor detects nicotine over 0.4-100 μM, a range relevant to nicotine concentrations present in active smoker sweat, saliva, gastric juice, and urine. This microbial mining approach for discovering redox enzymes expands the sensing parts toolbox available over conventional antibodies and aptamers.

|
Scooped by
mhryu@live.com
March 14, 4:04 PM
|
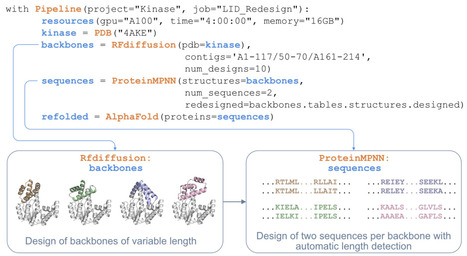
Deep learning methods for protein structure generation, sequence design, and structure and property prediction have created unprecedented opportunities for protein engineering and drug discovery. However, using these tools often requires navigating incompatible software environments, diverse input/output formats, and high-performance computing infrastructure, any of which may hinder adoption by primarily experimental chemical biology laboratories. Here we present BioPipelines, an open-source Python framework that allows researchers to define multi-step computational design workflows in a few lines of code. Additionally, its robust yet modular architecture provides a straightforward way to expand the toolkit with different functionalities, particularly by leveraging coding agents, with little effort. The framework currently integrates over 30 tools encompassing structure generation, sequence design, structure prediction, compound screening, and analysis. The same workflow code can be prototyped interactively in a Jupyter notebook and then submitted for production-scale runs without modification. We demonstrate applications in inverse folding, gene synthesis, de novo protein design, compound library screening, iterative binding site optimization, and fusion-protein linker optimization. We hope this framework will empower researchers, allowing them to focus on the scientific question rather than computational logistics. https://github.com/locbp-uzh/biopipelines
|

|
Scooped by
mhryu@live.com
March 15, 4:21 PM
|
We present a living, synthetic bacterial cell made by transplanting a complete genome into a dead cell. After killing Mycoplasma capricolum cells by chemically crosslinking their genome with Mitomycin C (MMC), we installed synthetic Mycoplasma mycoides genomes into the resulting dead cells using Whole Genome Transplantation (WGT). During WGT, a synthetic donor genome is placed into a recipient cell, thereby reprogramming that cell to adopt a new genetic identity. WGT has only been demonstrated using species within one phylogenetic clade of Mollicutes bacteria. A major barrier to expanding WGT to diverse bacterial species has been the inability to inactivate the recipient genome, leading to false positive transplants due to homologous recombination of antibiotic resistance markers from the donor genome into the recipient cell genome. Here, we address this key limitation by removing reliance on an antibiotic resistance marker to select for transplants; recipient cells are dead unless revived by the installation of a new genome. Our work demonstrates a general approach to fully inactivate the recipient cell genome, reports the first living synthetic bacterial cell constructed from non-living parts, and advances WGT for building engineered or synthetic cells for diverse applications.

|
Scooped by
mhryu@live.com
March 15, 4:08 PM
|
Phenotypic heterogeneity within isogenic bacterial populations represents a fundamental adaptation strategy that enables pathogen survival across the selective pressures of host infection. Rather than uniformly responding to environmental challenges, bacterial populations diversify through mechanisms including phase variation, stochastic gene expression, asymmetric cell division, and intercellular communication, generating functionally specialized subpopulations that operate through bet-hedging and division of labor frameworks. This review synthesizes recent advances showing how host-derived signals tune bacterial switching dynamics and deterministically partition populations into discrete phenotypic states. These functionally specialized subpopulations have clinical implications, with antibiotic-tolerant persisters representing one example of how phenotypic heterogeneity drives treatment failure and enables infection recurrence, underscoring the need to understand and target this fundamental aspect of bacterial pathogenesis.

|
Scooped by
mhryu@live.com
March 14, 11:32 PM
|
The acid activity of enzymes, characterized by the minimum pH at which enzymes remain active (pHmin), is crucial for industrial applications in acidic environments. However, the rational design of acid-active enzymes remains challenging due to limited understanding of sequence-structure-activity relationships under acidic conditions. Here, we propose ACENet, a graph neural network that predicts enzyme pHmin by integrating surface features of protein structures with evolutionary representations derived from the large-scale protein language model ESM-2. ACENet achieved a Pearson correlation coefficient of 0.85 on the test dataset, significantly outperforming other deep learning baseline models and maintains stable pHmin predictions under various conditions. Even on a subset of the dataset with less than 20% homology, the PCC remains above 0.5, with an RMSE (Root mean square error) less than 1.4. ACENet also present excellent performance in the annotation of pHmin for homologous proteins and the predictive screening of minimal active pH in protein mutants. Remarkably, ACENet could identify the catalytic region as key determinants of acid activity through residue-level interpretability analysis. Overall, ACENet accelerates the development of highly efficient biocatalysts for diverse applications where acidic conditions predominate.

|
Scooped by
mhryu@live.com
March 14, 10:37 PM
|
Infertility affects ~1 in 6 people of reproductive age and remains difficult to treat because causes are heterogeneous and diagnostics are incomplete. Recent evidence reframes the female reproductive tract as a low-biomass but biologically active microbial ecosystem. Dysbiosis, typically loss of protective Lactobacillus species (notably L. crispatus) with overgrowth of anaerobic pathobionts, is associated with implantation failure and recurrent pregnancy loss. Framing conditions such as chronic endometritis and reproducible low-Lactobacillus endometrial profiles as dysbiosis-related disorders clarifies opportunities for prevention, companion diagnostics and microbiome-directed therapies. This narrative review contrasts receptive (Lactobacillus-dominant) versus dysbiotic states and summarizes mechanisms linking microbiota to fertility: microbial metabolites (lactic acid, short-chain fatty acids) support epithelial barrier function and immune tolerance, whereas dysbiosis provokes inflammation that impairs implantation. Although observational data consistently associate Lactobacillus dominance with better outcomes, evidence quality is low-to-moderate due to retrospective designs, methodological heterogeneity, and a lack of adequately powered, diagnostic-stratified randomised trials. The review highlights precision microbial therapeutics under development, single-strain next-generation probiotics, synthetic consortia, engineered live biotherapeutics, postbiotics, targeted phage/endolysins and vaginal microbiota transplantation, and proposes a diagnostic-driven roadmap that matches microbiome endotypes and clinical contexts (e.g., preconception vs. immediate embryo transfer) to specific interventions. Regulatory and safety issues for reproductive biologics are also considered. The reproductive microbiome is a promising translational frontier but currently offers a consistent signal rather than definitive proof of benefit. To translate promise into practice requires standardised low-biomass sampling/reporting, mechanistic validation in human-relevant models and diagnostic-stratified randomised trials with staged endpoints, alongside strategies to address engraftment, formulation and regulatory pathways.

|
Scooped by
mhryu@live.com
March 14, 10:24 PM
|
The CRISPR–Cas9 system provides adaptive immunity against invading genetic elements through a dual-RNA-guided DNA cleavage mechanism. This system relies on the precise assembly of a ribonucleoprotein (RNP) complex composed of the Cas9 endonuclease, a CRISPR-derived RNA (crRNA), and a trans-activating CRISPR RNA (tracrRNA). Around 100 anti-CRISPR proteins that inhibit CRISPR–Cas systems have been identified, and the mechanisms by which they act are increasingly being elucidated. However, the inhibitory mechanisms of many Acrs, including AcrIIA7, remain poorly understood. Here, we present the structure of AcrIIA7 and uncover a previously unrecognized mechanism by which it inhibits Cas9 function. Structural and biochemical analyses reveal that AcrIIA7 specifically binds to tracrRNA, preventing its association with crRNA and thereby blocking formation of the active Cas9 RNP complex. This tracrRNA hijacking mechanism represents a unique strategy for CRISPR inhibition, in which an anti-CRISPR protein targets an RNA scaffold essential for Cas9 activation rather than interacting directly with the Cas9 protein. Our findings provide the first structural insight into tracrRNA-targeted anti-CRISPR activity and highlight RNA–RNA interaction interfaces as vulnerable nodes in CRISPR–Cas immunity. CRISPR–Cas9 immunity relies on forming a Cas9–crRNA–tracrRNA ribonucleoprotein complex. Here, the authors show that AcrIIA7 blocks this process by binding tracrRNA, preventing its pairing with crRNA.

|
Scooped by
mhryu@live.com
March 14, 5:15 PM
|
Jasmonic and salicylic acids are key hormones involved in plant responses to pests and pathogens. Existing fluorescence-based approaches to imaging plant defence hormones are constrained by the need for external illumination and by autofluorescence of plant tissues, while luminescence-based ones require exogenous substrates. Here, we use jasmonate- and salicylate-responsive promoters to engineer autoluminescent plants that report hormone signalling activity with up to a 53-fold contrast. Using consumer-grade cameras, we image reporter Arabidopsis thaliana and Nicotiana benthamiana plants throughout normal development and in response to pest and pathogen attacks, visualising local and systemic responses. Because the luminescence is self-sustained, these reporters enable non-invasive, substrate-free imaging of defence signalling over extended time courses without specialized equipment. Salicylic acid and jasmonic acid are two major hormones regulating plant response against pests and pathogens. Here, the authors configured the autoluminescence pathway to be conditional on activities of these hormones, achieving direct visualisation of plant defense responses.

|
Scooped by
mhryu@live.com
March 14, 5:09 PM
|
Across phyla and hosts, morphological plasticity serves as a common strategy for overcoming host defenses and optimizing growth. Whether toggling between yeast and hyphae to invade tissues, as seen in C. albicans and O. novo-ulmi, or shedding the cell wall for concealment like E. muscae, structural transitions are crucial in shaping the pathogenesis of these fungi. While efforts to block morphogenesis in C. albicans are underway, the broader importance of fungal “shapeshifting” remains underappreciated. Morphological plasticity is a recurring strategy that underlies the virulence of diverse fungal pathogens. Indeed, while the three examples we highlighted exploit yeast-hyphal transitions, there are other morphological transitions that fungi leverage for pathogenesis. Cryptococcus neoformans, for example, forms Titan cells—massive, multinucleate fungal cells that bud to produce normal sized daughters—while Coccidioides species form large, endospore-filled spherules. Understanding how pathogenic fungi regulate diverse morphological transitions could have far-reaching implications for developing effective antifungal treatments across phyla.

|
Scooped by
mhryu@live.com
March 14, 5:03 PM
|
Archaeal antiviral defense systems remain poorly characterized despite recent advances in understanding prokaryotic immunity. Here, we analyze 7747 archaeal genomes, the largest and most diverse dataset to date, revealing a striking disparity in defense system prevalence and diversity compared to Bacteria. Nearly one-third of archaeal genomes have no detected systems beyond CRISPR-Cas and restriction-modification (in contrast to only 2.2% bacterial genomes), and only 50–55% contain CRISPR-Cas systems, far below previous estimates. Many known defense systems appear restricted to Bacteria, while several single-gene putative candidate systems (PDCs) recently identified through a guilt-by-embedding approach are enriched in Archaea. Phylogenetic analyses suggest that PDC-S70 and PDC-M05 likely originated in Archaea, representing rare archaeal contributions to the prokaryotic immune repertoire. Consistent with earlier studies, our findings support the existence of deep evolutionary links between archaeal and eukaryotic systems for argonautes and viperins. These analyses highlight both the underexplored nature and the evolutionary significance of archaeal immunity, calling for expanded efforts to uncover archaeal-specific systems and improve our understanding of immune evolution across domains of life.

|
Scooped by
mhryu@live.com
March 14, 4:54 PM
|
The large size of widely used CRISPR-Cas enzymes limits their delivery for therapeutic applications. Cas12j nucleases offer a hypercompact alternative but show modest editing efficiency. To overcome this limitation, we identified eight novel Cas12j orthologs from viral metagenomes, which in their native form exhibit low editing activity in mammalian cells. We therefore engineered T5 exonuclease-Cas12j fusions, resulting in substantially enhanced genome-editing activity across multiple mammalian cell types, reaching levels comparable to established compact CRISPR-Cas editors. Intriguingly, robust cellular editing occurred in the presence of a previously unrecognized trinucleotide sequence context within the target DNA. Furthermore, we developed Cas12j-based adenine base editors by coupling catalytically inactive Cas12j orthologs with adenine deaminase, enabling efficient A-to-G base conversion in mammalian cells. This study expands the CRISPR toolbox by establishing engineering principles that convert compact Cas12j nucleases into efficient and modular genome-editing platforms well suited for delivery-constrained therapeutic applications.

|
Scooped by
mhryu@live.com
March 14, 4:47 PM
|
Foods derived from cultured cocoa, coffee, or other plant cells are nearing commercialization. Given the novelty of the production process and product characteristics, it is advisable to address food safety early during product development. International harmonization of a case-by-case approach for pre-market safety assessments should be pursued.

|
Scooped by
mhryu@live.com
March 14, 4:42 PM
|
Biodegradable adhesives, unlike their traditional counterparts, are engineered to bond to biological tissues while naturally degraded over time, thereby eliminating the need for removal procedures and reducing the risk of chronic inflammation. These unique features are particularly suitable for temporary biomedical applications such as wound closure, internal sealing, or integration with electronics for active/passive functions. The adhesive performance arises from the strategic combination of biodegradable polymers and adhesion mechanisms that dynamically interact with tissue surfaces. This review introduces recent advancements in biodegradable adhesives through a mechanism-based framework, focusing on five key adhesion strategies: physical interlocking, hydrogen bonding, catechol chemistry, amine-carboxyl coupling, and covalent bonding via diazirine or isocyanate linkages. For each strategy, representative material systems, functional properties, and biomedical implementations that enable strong, conformal adhesion under wet and physiological environments are highlighted, and with a discussion of current challenges and future directions toward intelligent, multifunctional bioadhesives for clinical uses are concluded.

|
Scooped by
mhryu@live.com
March 14, 4:13 PM
|
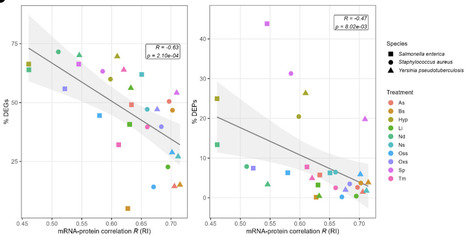
Diverse bacterial pathogens have evolved complex regulatory mechanisms to adapt to various environmental stresses during infection. The uncertainty in mRNA-protein levels in response to environmental stressors complicates our understanding of bacterial physiology and their adaptation to stressful environments. To examine this issue, we have integrated transcriptomics and proteomics data on three human bacterial pathogens Salmonella enterica Typhimurium[CE4.1][KA4.2], Yersinia pseudotuberculosis, and Staphylococcus aureus under ten infection-relevant stress conditions. We observed positive correlations between mRNA and protein levels, which were decreased under different stress conditions. Essential genes exhibited higher expression levels with lower variation across the conditions and stronger mRNA-protein correlations compared to non-essential genes, highlighting their critical role in bacterial adaptability and survival. Moreover, we identified a substantial number of genes with stress-induced non-correlating mRNA-protein levels, particularly under conditions triggering strong stress responses. Particularly this level was dramatically lowered for osmotic stress specific genes affected by impaired translational activity under osmotic stress. Our findings highlight the prevalence of non-correlating mRNA-protein levels and the potential role of post-translational modifications in modulating protein levels in response to environmental stressors during infection. This study provides a comprehensive framework for integrating transcriptomics and proteomics data and identifies potential gene products that might significantly impact the ability of diverse bacterial pathogens to adapt to hostile infection environments.
|
 Your new post is loading...
Your new post is loading...





























3st, agarase activity was determined by agar hydrolysis patterns such as pitting around colonies on the agar plate or staining the plate by iodine solution. pFAST was evaluated by fluorescence emission under UV light (354 nm) after spotting the fluorophore substrate HBR (4-hydroxybenzylidene rhodanine) over the colony.. substrate HBR was added to cells prior to fluorescence measurement.