Your new post is loading...
 Your new post is loading...

|
Scooped by
Jean-Michel Ané
June 18, 5:21 PM
|
•Biofertilizers promoted the growth and nutrition of pea plants.
•Vermicompost and Azospirillum improved the soil organic matter.
•Mycorrhizal and Azospirillum improved the carbon of the microbial biomass.
•Height and weight improved with mycorrhizal and Azospirillum with vermicompost.
•Azospirrillum and mycorrhiza improved the profitability of the pea crop.

|
Scooped by
Jean-Michel Ané
June 16, 11:59 AM
|
Crops are increasingly exposed to drought and nutrient deficiencies, necessitating enhanced resistance to adverse conditions to meet the growing demands of the global population. While crop productivity has been greatly improved by integrating traits for high yield and stress tolerance through breeding, yield plateaus are now being observed. The rhizosheath, with physical and biological properties distinct from bulk soil, presents a promising target for enhanced tolerance to abiotic stresses such as drought and nutrient deficiencies. This multifunctional region contributes substantially to stress resistance and nutrient cycling, playing a pivotal role in the context of climate change and diminishing supplies of non-renewable fertilisers. We highlight the potential of the rhizosheath as a valuable breeding target to enhance crop productivity under diverse challenging environmental conditions.

|
Scooped by
Jean-Michel Ané
June 15, 10:34 AM
|
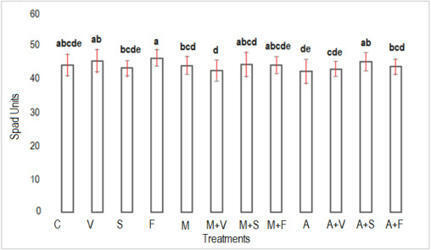
We reported that Glycyrrhiza uralensis inoculated with rhizobium tended to increase biomass production and glycyrrhizic acid (GL) production, in this study we have also achieved drastically increase in biomass and GL production in Glycyrrhiza glabra. At thirty days after inoculation (DAI), a significant increase in SPAD values was observed, and the expression of GL synthesis marker genes was also significantly increased. At 150 DAI, a significant increase in biomass was observed. Characteristically, it was also found that thick roots were enlarged by rhizobial inoculation. In addition, the expression of GL synthesis marker genes was also significantly increased. Moreover, GL content per unit root dry weight reached 4%, and GL production per plant increased six times compared to uninoculated plants. Moreover, we tried to reveal the mechanism of induction of GL production by rhizobial inoculation. Since it has been reported that the expression of jasmonic acid (JA) synthesis marker genes is increased by rhizobium in soybean, we investigated the expression of those genes in G. glabra, and found that GgMYC2 and GgJAR1 were up-regulated at Thirty DAI. Furthermore, methyl jasmonate treatment increased the expression of GL synthesis marker genes, suggesting that JA signaling is involved in the increased GL production due to rhizobial inoculation. These results aid in understanding the mechanism of increased GL production through the introduction of rhizobial symbiosis, and show the potential for providing a technology to significantly shorten the cultivation period for the production of Glycyrrhiza that meets the criteria for herbal medicines.

|
Scooped by
Jean-Michel Ané
June 13, 1:16 PM
|
Fungi are one of the most diverse and ecologically important groups of organisms on Earth. They exhibit remarkable diversity in their ecological roles, ranging from decomposers to mutualistic symbionts to parasites. They have a wide array of lifestyles, which reflect their diverse ecological roles and evolutionary adaptations to marine, aquatic, and terrestrial ecosystems. Fungi are osmotrophs that grow as filaments of cells (hyphae) into their food, secrete digestive enzymes across their cells’ chitinous walls, and absorb dissolved nutrients. The classification of fungal lifestyles is primarily based on how they obtain nutrients, with the major modes of nutrition being saprotrophy, parasitism, mutualism and commensalism. Here, we briefly explore these various lifestyles, illustrating their significance in ecosystems and their relationships with other organisms, and then discuss how comparative genomics provides novel insights into their evolutionary trajectories.

|
Scooped by
Jean-Michel Ané
June 13, 11:49 AM
|
Arbuscular mycorrhizal (AM) fungi are ancient plant mutualists that are ubiquitous across terrestrial ecosystems. These fungi are unique among most eukaryotes because they form multinucleate, open-pipe mycelial networks, where nutrients, organelles, and chemical signals move bidirectionally across a continuous cytoplasm. AM fungi play a crucial role in ecosystem functioning by supporting plant growth, mediating ecosystem diversity, and contributing to carbon cycling. It is estimated that plant communities allocate ∼3.93 Gt CO2e to AM fungi every year, much of which is stored as lipids inside the fungal network. Despite their ecological significance, the cellular biology of AM fungi remains underexplored. Here, we synthesise the current knowledge on AM fungal cellular structure and organisation. We examine AM fungal development at different biological levels — the hypha and its content, hyphal networks and AM fungal spores — and explore key cellular dynamics. This includes cell wall composition, cytoplasmic contents, nuclear and lipid organisation and dynamics, network architecture, and connectivity. We highlight how their unique cellular arrangement enables complex cytoplasmic flow and nutrient exchange processes across their open-pipe mycelial networks. We discuss how both established and novel techniques, including microscopy, culturing, and high-throughput image analysis, are helping to resolve previously unknown aspects of AM fungal biology. By comparing these insights with established knowledge in other, well-studied filamentous fungi, we identify critical knowledge gaps and propose questions for future research to further our understanding of fundamental AM fungal cell biology and its contributions to ecosystem health.

|
Scooped by
Jean-Michel Ané
June 13, 11:33 AM
|
Symbiotic nitrogen fixation represents a crucial yet energy-demanding strategy for legumes to survive in nutrient-poor soils. We highlight the multifaceted roles of NSP1 and NSP2 in this symbiosis and propose their function as ‘nutrient-responsive regulators’, integrating environmental signals, physiological status, and nutrient availability, to ensure nodulation occurs only under favorable conditions.

|
Scooped by
Jean-Michel Ané
June 12, 11:19 AM
|
Root nodule symbiosis allows for plant acquisition of reactive nitrogen through fixation of atmospheric molecular dinitrogen by nitrogen-fixing bacteria. Nodulation is a complex trait, with diverse modes of bacterial infection and nodule morphologies across species, reflecting evolutionary adaptation. Understanding ancient forms of this trait may carry advantages for its current utilization, since basal states likely reflect the least complexity. In this review we focus on the evolution of nodule development, particularly on events that have led to increased complexity of this symbiosis in later adaptations. We hypothesize that the ancestral form of nodulation comprises of an evolutionary coupling of nutrient-dependent lateral root development with apoplastic intercellular bacterial growth, alongside the acquisition or evolution of an ancestral chitinaceous signaling molecule by the microbial symbiont. Uncovering the evolutionary adaptations underpinning the extant diversity of this trait allows for a better understanding of the simplest ancestral state.

|
Scooped by
Jean-Michel Ané
June 12, 9:45 AM
|
Arbuscular mycorrhiza (AM) with soilborne Glomeromycota fungi was pivotal in the conquest of land by plants almost half a billion years ago. In flowering plants, it is hypothesized that AM is initiated by the perception of AM fungi-derived chito- and lipochito-oligosaccharides (COs/LCOs) in the host via Lysin Motif Receptor-Like Kinases (LysM-RLKs). However, it remains uncertain whether plant perception of these molecules is a prerequisite for AM establishment and for its origin. Here, we made use of the reduced LysM-RLK complement present in the liverwort Marchantia paleacea to assess the conservation of the role played by this class of receptors during AM and in CO/LCO perception. Our reverse genetic approach demonstrates the critical function of a single LysM-RLK, MpaLYKa, in AM formation, thereby supporting an ancestral function for this receptor in symbiosis. Binding studies, cytosolic calcium variation recordings and genome-wide transcriptomics indicate that another LysM-RLK of M. paleacea, MpaLYR, is also required for triggering a response to COs and tested LCOs, despite being dispensable for AM formation. Collectively, our results demonstrate that the perception of symbionts by LysM-RLK is an ancestral feature in land plants, and suggest the existence of yet-uncharacterized AM fungi signals.

|
Scooped by
Jean-Michel Ané
June 11, 5:22 PM
|
Background and aims
Biological nitrogen fixation (BNF) is a key factor in ensuring the sustainability of agricultural production and food security. This study aimed to estimate the savings generated by BNF in common bean (Phaseolus vulgaris), cowpea (Vigna unguiculata) and mung bean (Vigna radiata).
Material and methods
Data on the area, production and productivity of the crops in this study were obtained from the Brazilian agricultural censuses of 2006 and 2017. We estimated the amount of N (urea) necessary to produce 1000 kg of grain of each bean species in the absence of inoculation. The contribution of BNF to the N supply of each legume species was obtained from studies using the 15N natural abundance method.
Results
We highlight that in Brazil, the savings generated by the BNF were US$6 million with mung beans, US$59 million with cowpeas, and US$119 million with common beans, in 2017. The savings generated by BNF per hectare planted with common beans in 2017 increased by 40% compared to 2006, mainly due to the significant increase of 64% in the grain yield of this crop.
Conclusion
Our study highlights the importance of BNF in the production of these pulse species that allows a reduction in the use of N-fertilizers and an increase in savings in these production systems, which contributes to sustainability of agricultural production.

|
Scooped by
Jean-Michel Ané
June 9, 4:38 PM
|
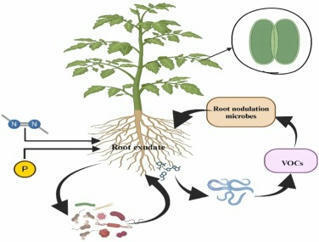
•The study showed the role of root exudate in drought stress mitigation.
•In drought conditions, increased naringenin and daidzein in exudates to attract diazotrophic bacteria.
•Proteoid roots enhance surface area, carboxylic acid release, and phosphorus uptake in drought conditions.
•The HvAACT1 gene aids citrate exudation at root apex, boosting elongation of root in drought conditions.
•This study is useful in formulating strategies for sustainable environment management.

|
Scooped by
Jean-Michel Ané
June 9, 2:06 PM
|
The environmental release of both engineered and non-engineered organisms for Biotechnologies Beyond Conventional Containment (BBCC) offers unique solutions to pressing global challenges, including the prevention of soil degradation, the attenuation of nitrogen pollution, the replacement of harmful pesticides and herbicides, the remediation of anthropogenic contaminants and ‘forever chemicals’ mitigation. An evaluation of impacts, both positive and negative, rather than arbitrary prohibitions, is crucial for advancing the responsible use of organisms intentionally released into the environment. The history of biological interventions demonstrates that organisms have successfully contributed to agriculture, pollution remediation, ecosystem restoration, waste upcycling, and pest control, yet their full potential remains constrained by regulatory hurdles that do not fully account for modern scientific advancements. At the same time, some releases serve as cautionary tales, having caused harm due to a lack of regulation and monitoring. Unlike chemicals released to the environment, organisms — particularly those designed or selected for specific functions — can be managed with built-in safeguards, ranging from physical and genetic containment strategies to controlled ecological interactions to mitigate risks while maximizing benefits. Advancements in precision engineering, computational modeling, and real-time monitoring technologies now allow for unprecedented accuracy in tracking, assessing, and controlling the environmental impact of released organisms — capabilities inaccessible when recombinant DNA technology first emerged 50 years ago. Many regulatory structures were developed decades before today’s explosion of biological knowledge and insight was even imaginable. This resulted in our current policies that have become restrictive, limiting the deployment of innovative and promising biological solutions. A new approach to risk analysis is now needed that accounts for changes in science, and in society, which assesses the environmental release of natural, evolved, and engineered organisms based on their functions rather than their origin or how they were developed. By modernizing these frameworks to emphasize continuous assessment, real-world data collection, and adaptive risk assessment and management, stakeholders can create a regulatory pathway for the sustainable, responsible, and evidence-based integration of environmental biological technologies.

|
Scooped by
Jean-Michel Ané
June 6, 6:34 PM
|
Background
Symbiotic nitrogen fixation (SNF) is a complex process regulated by numerous genes extensively studied in legumes that undergo intracellular infection, such as Lotus japonicus, Medicago truncatula, and Glycine max. However, the molecular and genetic mechanisms of SNF in legumes that rely on the intercellular infection pathway, such as peanut (Arachis hypogaea L.), remain poorly understood. In a previous study, we identified two chromosome segment substitution lines (CSSLs), 12CS_051 and 12CS_044, each contains a wild segment on homeologous regions of chromosomes A02 and B02 respectively, that are severely impaired in nitrogen fixation. In this study, we have compared the transcriptomes of those lines with that of their recurrent parent, Fleur11, in roots inoculated with the effective Bradyrhizobium vignae strain ISRA400 to identify candidate genes associated with the reduced nitrogen fixation observed in these CSSLs.
Results
A comparative analysis of the transcriptome profiles of the CSSLs and Fleur11 revealed significant changes in the expression of genes involved in plant immune signaling and key symbiotic genes, such as NIN, EFD, FEN1 or SNF-related transporters. These results align with the phenotypic differences observed during the symbiotic process in the CSSLs. When focusing on each QTL region, we found that only the orthologs of the symbiotic gene FEN1, which is responsible for the failure in the enlargement of infected cells in L. japonicus, exhibited a lack of expression in the two CSSLs compared to Fleur11. FEN1 encodes a homocitrate synthase that is essential for the nitrogenase activity. We hypothesize that changes in the expression of FEN1 could affect the nitrogenase activity, potentially leading to the unfair SNF observed in these lines.
Conclusions
In this study, we analyzed the expression profiles of two ineffective nitrogen-fixing chromosome segment substitution lines and identified FEN1 as a suitable candidate gene involved in peanut symbiosis. This research provides valuable insights into understanding and improving SNF in peanut.

|
Scooped by
Jean-Michel Ané
June 2, 3:27 PM
|
While intracellular symbiosis with rhizobia relies on Nod factor signaling through the conserved common symbiosis signaling pathway (CSSP), it remains unclear how legumes simultaneously manage interactions with commensal soil microbes. Using single cell RNA-sequencing, we show that commensal soil bacteria induce a Nod factor-independent transcriptional response in specific root hairs. This response is similar to the rhizobium response in the CSSP-deficient cyclops mutant, which is unable to accommodate rhizobia in root hair infection threads. Both responses include the nodulation gene NSP2 and a transcription factor, which we name ROOT HAIR DEFECTIVE 6 LIKE A (RHD6LA). We show that RHD6LA is required for facilitating infection thread formation in response to rhizobia and for preventing exaggerated root hair responses to commensal soil bacteria. The overlap between commensal and symbiotic signaling highlights the complexity of legume-microbe interactions at the root hair interface and suggests new mechanisms for microbial discrimination in rhizobium-responsive root hairs.
|

|
Scooped by
Jean-Michel Ané
June 17, 2:16 PM
|
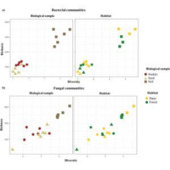
Unraveling the mechanisms underlying the maintenance of species diversity is a central pursuit in ecology. It has been hypothesized that ectomycorrhizal (EcM) in contrast to arbuscular mycorrhizal fungi can reduce tree species diversity in local communities, which remains to be tested at the global scale. To address this gap, we analyzed global forest inventory data and revealed that the relationship between tree species richness and EcM tree proportion varied along environmental gradients. Specifically, the relationship is more negative at low latitudes and in moist conditions but is unimodal at high latitudes and in arid conditions. The negative association of EcM tree proportion on species diversity at low latitudes and in humid conditions is likely due to more negative plant-soil microbial interactions in these regions. These findings extend our knowledge on the mechanisms shaping global patterns in plant species diversity from a belowground view.

|
Scooped by
Jean-Michel Ané
June 16, 11:54 AM
|
Globally, invasive plants, animals, and microbes are a dominant threat to biodiversity and ecosystems, inflicting over $1 trillion USD in damage annually. Hundreds of microbial invasive taxa are documented.
Currently, we are inoculating microbes into many environments in enormous numbers to fertilize agricultural soils, remediate contamination, control eutrophication, precipitate minerals, and restore ecosystem functions.
Given the astronomical numbers and myriad environments that are being inoculated with microbes, these manipulations risk creating microbial invasions in much the same way that catastrophic plant and animal invasions have been precipitated by intentional introductions by humans.
A mechanistic understanding and predictive framework for the potential for microbial inoculants to cause invasions is needed to balance their benefits with their risks of causing harmful effects.

|
Scooped by
Jean-Michel Ané
June 15, 10:32 AM
|
Plants and microbiomes have co-evolved for millennia. Through this co-evolution, microbiomes have become essential for plant nutrient acquisition, which involves plant signaling, microbial sensing, and acquiring and sharing nutrients. In this review, we synthesize recent advancements in the complex associations of molecular, physiological, and eco-evolutionary mechanisms that underpin microbe-facilitated plant nutrient uptake. Focusing on emerging insights in plant-microbial communication and metabolic pathways, we evaluate potential opportunities to harness plant microbiomes to sustainably supply nutrients in agricultural and natural ecosystems. However, further progress is constrained by key knowledge gaps. We propose an amended conceptual framework for advancement that includes a holistic understanding of eco-evolutionary relationships with explicit consideration of signaling and sensing mechanisms. Finally, we argue that advancing fundamental science by utilizing emerging analytical approaches in an integrated way is critical to develop effective microbiome-informed tools that can enhance plant nutrient acquisition and promote long-term food security and environmental sustainability.

|
Scooped by
Jean-Michel Ané
June 13, 12:59 PM
|
Breakthroughs in DNA sequencing have upended our understanding of fungal diversity. Only ∼155,000 of the 2–3 million fungal species on the planet have been formally described and named, and ‘dark taxa’ — species known only from sequences — represent the vast majority of species within the fungal kingdom. The International Code of Nomenclature requires physical type specimens to officially recognize new fungal species, making it difficult to name dark taxa. This is a significant problem for conservation because, without names, species cannot be recognized for environmental and legal protection. Symbiotic ectomycorrhizal (EcM) fungi play a particularly important role in forest carbon drawdown, but at present we have little understanding of how many EcM fungal species exist, or where to prioritize research activities to survey and describe EcM fungal lineages. In this review, we use global soil metabarcoding databases (GlobalFungi and the Global Soil Mycobiome consortium) to evaluate current estimates of the total number of EcM fungal species on Earth, outline the current state of undescribed EcM dark taxa, and identify priority regions for future dark taxa exploration. The metabarcoding databases include up to 219,730 EcM fungal operational taxonomic units (OTUs) detected from almost 39,500 samples. Using Chao richness estimates corrected for extrapolating species numbers from metabarcoding datasets, we predict that the global diversity of EcM fungi could be ∼25,500–55,500 species. Dark taxa — those that do not match species-level identities — account for 79–83% of OTUs. Oceania contains the highest percentage of dark taxa (87%), and Europe the lowest (78%). Priority ‘darkspots’ for future research occur predominantly in tropical regions, but also in selected temperate forests at both southern and northern latitudes. We propose concrete steps to reduce the prevalence of EcM darkspots, including performing targeted field surveys, barcoding fungaria voucher specimens, and developing new ways to describe and conserve fungal taxa from DNA alone.

|
Scooped by
Jean-Michel Ané
June 13, 11:45 AM
|
Plants accommodate diverse microbial communities, termed the microbiome, which can change dynamically during plant adaptation to varying environmental conditions. However, the direction of these changes and the underlying mechanisms driving them, particularly in crops adapting to the field conditions, are not well understood. Here, we investigate the root endosphere microbiome of rice (Oryza sativa ssp. japonica) across four consecutive cultivation seasons in a high-yield, non-fertilized, and pesticide-free paddy field, compared to a neighboring fertilized and pesticide-treated field. Using 16S rRNA amplicon and metagenome sequencing, we analyzed three Japonica cultivars—Nipponbare, Hinohikari, and Kinmaze. Our findings reveal that the root endosphere microbiomes diverge based on fertilization regime and plant developmental stages, while the effects of cultivar variation are less significant. Machine learning model and metagenomic analysis of nitrogenase (nif) genes suggest enhanced nitrogen fixation activity in the non-fertilized field-grown roots, highlighting a potential role of diazotrophic, iron-reducing bacteria Telmatospirillum. These results provide valuable insights into the assembly of the rice root microbiome in nutrient-poor soil, which can aid in managing microbial homeostasis for sustainable agriculture.

|
Scooped by
Jean-Michel Ané
June 12, 11:24 AM
|
NifA, the activator of nitrogenase, is sensitive to ammonium concentration, particularly within its N-terminal domain. In this work, genetically engineered mutants with N-terminal deletions of the nifA1 and nifA2 genes were constructed using overlap extension PCR to reduce the inhibitory effect of ammonium on nitrogenase expression in Rhodobacter capsulatus SB1003. Under 3 mM NH4+, the hydrogen production rate of ZX03 (nifA1-, nifA2-) reached 0.65 mmol L−1 h−1, with a 31.2 % increase in hydrogen production compared to the wild-type. When 8 mM NH4+ was used as the sole nitrogen source, H2 production in all strains decreased substantially compared to 5 mM NH4+. However, ZX03 demonstrated a 5.2-fold enhancement in hydrogen production relative to the wild-type under 8 mM NH4+, underscoring its improved ammonium tolerance. During hydrogen production, the gene expression levels of nifA and nifH in all mutant strains were significantly up-regulated under ammonium conditions compared to the wild-type. These findings reveal distinct roles of nifA1 and nifA2 in ammonium tolerance and hydrogen production.

|
Scooped by
Jean-Michel Ané
June 12, 9:49 AM
|
Many plants and animals, including humans, host diverse communities of microbes that provide many benefits. A key challenge in understanding microbiomes is that the species composition often differs among individuals, which can thwart generalization. Here, we argue that the key to identifying general principles for microbiome science lies in microbial metabolism. In the human microbiome and in other systems, every microbial species must find ways to harvest nutrients to thrive. The available nutrients in a microbiome interact with microbial metabolism to define which species have the potential to persist in a host. The resulting nutrient competition shapes other mechanisms, including bacterial warfare and cross-feeding, to define microbiome composition and properties. We discuss impacts on ecological stability, colonization resistance, nutrient provision for the host, and evolution. A focus on the metabolic ecology of microbiomes offers a powerful way to understand and engineer microbiomes in health, agriculture, and the environment.

|
Scooped by
Jean-Michel Ané
June 11, 5:25 PM
|
Acacia longifolia, a species native to Australia, is an aggressive invasive in Mediterranean-type ecosystems worldwide. Its success in diverse habitats, expanding from coastal dunes to forests, is often attributed to its ability to establish interactions with a variety of microbes, including bacteria and fungi. This study investigates the seed and root-nodule microbiomes of A. longifolia to understand the roles these microbial communities play in its adaptation and invasive behaviour. Using high-throughput sequencing, we characterized the bacteriome and mycobiome associated with the plant, considering nodules and seeds, and the surrounding soil in three different locations in Portugal with different climate conditions (North, Center and South), and a comparison between two different habitats (Dune versus Forest). Our results reveal a dynamic interaction between A. longifolia and its microbial partners, supporting the importance of these plant-microbe interactions in nutrient acquisition and stress tolerance for A. longifolia, ultimately leading to their impact in an invaded ecosystem. The seed microbiome of A. longifolia was less diverse than for nodules but with more functions assigned, while nodules showed a broader diversity, assigned to more specific functions. Here we provide evidence for the role of seed microbiota in germination and seed-to-seedling transition along with the beneficial role of nodulation in development and seedling-to-sapling switch. We also propose a local signature for microbial communities as we found a dissimilarity in microbial partners when considering habitat, with dune communities showing a functional plasticity, aiding A. longifolia to cope in such nutrient-limiting environment. For forests, functions more related with plant and microbe associations are evidenced, possibly facilitating interspecific competition. These findings contribute to an understanding of the plant-microbe interactions and dynamics that underpin A. longifolia ecological success as an invasive plant.

|
Scooped by
Jean-Michel Ané
June 11, 5:19 PM
|
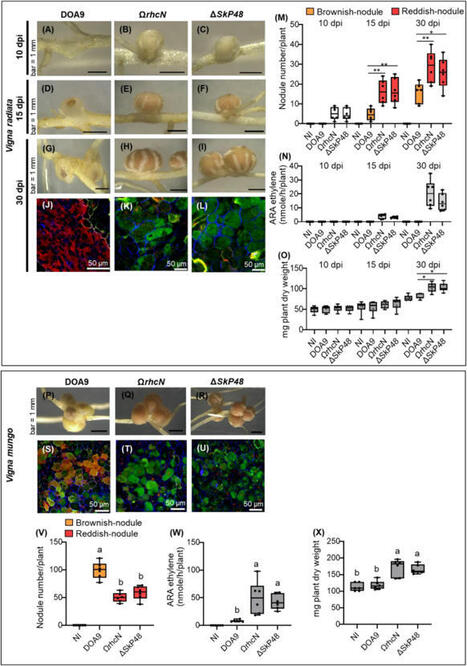
Bradyrhizobium sp. DOA9 can nodulate a wide spectrum of legumes; however, unlike other bradyrhizobia, DOA9 carries a symbiotic plasmid harboring type III secretion system (T3SS) and several effector (T3E) genes, one of which encodes a putative type III effector SkP48. Here, we demonstrated the pivotal roles of SkP48 from Bradyrhizobium sp. DOA9 in inhibiting nodulation of various Vigna species and Crotalaria juncea and suppressing nodulation efficiency of Arachis hypogaea. By contrast, the nodulation efficiency of a SkP48 mutant did not differ significantly with the DOA9 wild-type strain on Macroptilium atropurpureum and Stylosanthes hamata. The SUMO domain of SkP48 is primarily responsible for the blocking nodulation phenotype V. radiata. An evolutionary analysis revealed that the SkP48 which contains a shikimate kinase and a SUMO protease (C48 cysteine peptidase) domain, SkP48 is distinct from other effectors previously reported in other bradyrhizobia and pathogenic bacteria. Our findings suggest that the putative T3E SkP48 is a key factor suppressing nodulation and nodule organogenesis in several legumes by activation of effector-triggered immunity through salicylic acid biosynthesis induction, which is deleterious to rhizobial infection. In addition, nodulation may be modulated by the function of defensins involved in jasmonic acid signalling in V. radiata SUT1.

|
Scooped by
Jean-Michel Ané
June 9, 2:11 PM
|
Understanding the molecular basis of regulated nitrogen (N2) fixation is essential for engineering N2-fixing bacteria that fulfill the demand of crop plants for fixed nitrogen, reducing our reliance on synthetic nitrogen fertilizers. In Azotobacter vinelandii and many other members of Proteobacteria, the two-component system NifL-NifA controls the expression of nif genes that encode the nitrogen fixation machinery. The NifL-NifA system evolved the ability to integrate several environmental cues, such as oxygen, nitrogen, and carbon availability. The nitrogen fixation machinery is thereby only activated under strictly favorable conditions, enabling diazotrophs to thrive in competitive environments. Whilst genetic and biochemical studies have enlightened our understanding of how NifL represses NifA, the molecular basis of NifA sequestration by NifL depends on structural information on their interaction. Here, we present mechanistic insights into how nitrogen fixation is regulated by combining biochemical and genetic approaches with a low-resolution cryo-EM map of the oxidized NifL-NifA complex. Our findings define the interaction surface between NifL and NifA and reveal how this interaction can be manipulated to generate bacterial strains with increased nitrogen fixation rates able to secrete surplus nitrogen outside the cell, a crucial step in engineering improved nitrogen delivery to crop plants.

|
Scooped by
Jean-Michel Ané
June 6, 6:40 PM
|
The objective of this study was to characterize the arbuscular mycorrhizal fungi (AMF) community associated with local varieties of Pinot noir, Sauvignon blanc, and Chardonnay grown in the Malleco and Cautín valleys, in La Araucanía region, Chile. The research attempted to answer the following question: How do soil and climatic conditions, as well as cultivar characteristics could influence root colonization, spore abundance, and AMF composition in vineyards in this emerging region? Rhizosphere samples were collected from two locations and six grapevine cultivars. Root colonization rates and AMF spore abundance were measured, and the spores present were morphologically identified. In addition, differences in the AMF community were evaluated regarding cultivar and plant age. The results showed root colonization rates higher than 50%, with no significant differences between sites. However, variations in spore abundance and AMF community composition were observed among cultivars. Twelve AMF genera were identified, including Glomus, Sclerocystis, Dominikia, Rhizoglomus, Oehlia, and Paraglomus. Overall, Glomus rubiforme, Sclerocystis sp. CL1, Rhizoglomus irregulare, and Diversispora versiformis were the most abundant morphotypes. Additionally, R. irregulare, G. rubiforme, and Paraglomus occultum were consistently detected across nearly all analyzed rhizospheres. The presence of AMF genera varies according to cultivar, but not according to clones or plant age. It is hypothesized that differences in root architecture and root exudates of different grapevine cultivars are responsible for the observed variations in the composition of native AMF. These factors should be further investigated in future studies.

|
Scooped by
Jean-Michel Ané
June 3, 11:02 AM
|
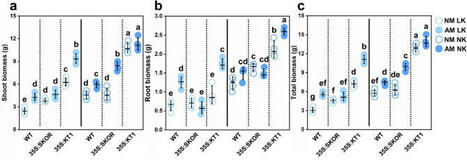
Potassium participates in a variety of plant physiological processes and has great impact on plant growth and stress adaptation. The absorption of potassium by Plant is mediated by potassium channels and transporters, and the Shaker potassium channel gene family plays an important role in potassium uptake. Arbuscular mycorrhizal (AM) fungi form ubiquitous symbioses with plants and increase plants’ potassium uptake. However, few studies have focused on the interaction of plant potassium channels from the Shaker gene family with AM fungi. In this study, the potassium uptake function of LbKT1 and LbSKOR (homologs of AKT1 and SKOR in Arabidopsis) from the Shaker gene family in Lycium barbarum was verified by the complementary assay using a yeast potassium uptake mutant. LbKT1 and LbSKOR were also overexpressed in tobacco to assess their influence on AM fungi under low and normal potassium conditions in a pot experiment. LbKT1 could rescue the phenotype of the yeast mutant, while LbSKOR could not. Overexpression of LbKT1 increased tobacco plant growth and potassium uptake and promoted the colonization of AM fungi. Meanwhile, overexpression of LbSKOR promoted potassium translocation from root to shoot and showed no obvious influence on the colonization of AM fungi. Our results suggested that the AM fungi could promote tobacco growth and potassium uptake, while the plant potassium status and the AM fungal colonization may form positive feedback in promoting tobacco potassium uptake and growth.
|
 Your new post is loading...
Your new post is loading...
 Your new post is loading...
Your new post is loading...