Your new post is loading...
 Your new post is loading...

|
Scooped by
Jean-Michel Ané
February 5, 2013 5:40 PM
|
Legumes control the nitrogen-fixing root nodule symbiosis in response to external and internal stimuli, such as nitrate, and via systemic autoregulation of nodulation (AON). Overexpression of the CLV3/ESR-related (CLE) pre-propeptide-encoding genes GmNIC1 (nitrate-induced and acting locally) and GmRIC1 (Bradyrhizobium-induced and acting systemically) suppresses soybean nodulation dependent on the activity of the nodulation autoregulation receptor kinase (GmNARK). This nodule inhibition response was used to assess the relative importance of key structural components within and around the CLE domain sequences of these genes. Using a site-directed mutagenesis approach, mutants were produced at each amino acid within the CLE domain (RLAPEGPDPHHN) ofGmRIC1. This approach identified the Arg1, Ala3, Pro4, Gly6, Pro7, Asp8, His11, and Asn12 residues as critical to GmRIC1 nodulation suppression activity (NSA). In contrast, none of the mutations in conserved residues outside of the CLE domain showed compromised NSA. Chimeric genes derived from combinations of GmRIC1 and GmNIC1 domains were used to determine the role of each pre-propeptide domain in NSA differences that exist between the two peptides. It was found that the transit peptide and CLE peptide regions of GmRIC1 significantly enhanced activity of GmNIC1. In contrast, the comparable GmNIC1 domains reduced the NSA of GmRIC1. Identification of these critical residues and domains provides a better understanding of how these hormone-like peptides function in plant development and regulation.

|
Rescooped by
Jean-Michel Ané
from Rhizobium Research
January 31, 2013 8:56 AM
|
The ability of the photosynthetic Bradyrhizobium strains ORS285 and ORS278 to nodulate soybeans was investigated. While the nod gene-deficient ORS278 strain induced only bumps on soybean roots, the nod gene-containing strain ORS285 formed nitrogen-fixing nodules. However, symbiotic efficiency differed drastically depending on both the soybean genotypes used and the culture conditions tested. Giraud E, Xu L, Chaintreuil C, Gargani D, Gully D, Sadowsky MJ. (2013). Appl Environ Microbiol. Jan 25. [Epub ahead of print]
Via IvanOresnik

|
Rescooped by
Jean-Michel Ané
from Plants and Microbes
January 28, 2013 8:57 AM
|
The goal of this symposium is to bring together researchers working in a wide range of disciplines in plant biology, ecology and evolution in the field of plant interactions with other organisms. The central objective is to discuss and integrate information from different approaches and perspectives in order to create a synthetic framework for understanding these interactions. Our hope is to have a diverse and dynamic group of scientists, who are willing to step out of their disciplinary comfort zone and engage in an effort to participate in discussions spanning the range from molecular approaches to ecosystem implications.
Via Kamoun Lab @ TSL

|
Rescooped by
Jean-Michel Ané
from Rhizobium Research
January 24, 2013 8:59 AM
|
Resources from the Sinorhizobium meliloti Rm1021 open reading frame (ORF) plasmid libraries were used in a medium-throughput method to construct a set of 50 overlapping deletion mutants covering all of the Rm1021 pSymA megaplasmid except the replicon region. Each resulting pSymA derivative carried a defined deletion of approximately 25 ORFs. Various phenotypes, including cytochrome crespiration activity, the ability of the mutants to grow on various carbon and nitrogen sources and the symbiotic effectiveness of the mutants with alfalfa were analyzed. This approach allowed us to systematically evaluate the potential impact of regions of Rm1021 pSymA for their free-living and symbiotic phenotypes.
Yurgel SN, Mortimer MW, Rice J, Humann J, Kahn ML. (2013). Appl Environ Microbiol. Jan 18. [Epub ahead of print]
Via IvanOresnik

|
Rescooped by
Jean-Michel Ané
from Rhizobium Research
January 12, 2013 12:12 PM
|
Abstract Microevolution and origins of Bradyrhizobium populations associated with soybeans at two field sites (A and B, 280 km apart in Canada) with contrasting histories of inoculation was investigated using probabilistic analyses of six core (housekeeping) gene sequences. These analyses supported division of 220 isolates in five lineages corresponding either to B. japonicum groups 1 and 1a or to one of three novel lineages within the genus Bradyrhizobium. None of the isolates from site A and about 20% from site B (the only site with a recent inoculation history) were attributed to inoculation sources. The data suggest that most isolates were of indigenous origin based on sequence analysis of 148 isolates of soybean-nodulating bacteria from native legumes (Amphicarpaea bracteata and Desmodium canadense). Isolates from D. canadense clustered with B. japonicum group 1, whereas those from A. bracteata were placed in two novel lineages encountered at soybean field sites. One of these novel lineages predominated at soybean sites and exhibited a significant clonal expansion likely reflecting selection by the plant host. Homologous recombination events detected in the 35 sequence types from soybean sites had an effect on genetic diversification that was approximately equal to mutation. Interlineage transfer of core genes was infrequent and mostly attributable to gyrB that had a history of frequent recombination. Symbiotic gene sequences (nodC and nifH) of isolates from soybean sites and native legumes clustered in two lineages corresponding to B. japonicum and B. elkani with the inheritance of these genes appearing predominantly by vertical transmission. The data suggest that soybean-nodulating bacteria associated with native legumes represent a novel source of ecologically adapted bacteria for soybean inoculation. Tang J, Bromfield ES, Rodrigue N, Cloutier S, Tambong JT. (2012). Ecol Evol. Dec;2(12):2943-61. doi: 10.1002/ece3.404. Epub 2012 Oct 22.
Via IvanOresnik

|
Rescooped by
Jean-Michel Ané
from Rhizobium Research
January 3, 2013 4:50 PM
|
SummaryMedicago truncatula and Sinorhizobium meliloti form a symbiotic association resulting in the formation of nitrogen-fixing nodules. Nodule cells contain large numbers of bacteroids which are differentiated, nitrogen-fixing forms of the symbiotic bacteria. In the nodules, symbiotic plant cells home and maintain hundreds of viable bacteria. In order to better understand the molecular mechanism sustaining the phenomenon, we searched for new plant genes required for effective symbiosis.We used a combination of forward and reverse genetics approaches to identify a gene required for nitrogen fixation, and we used cell and molecular biology to characterize the mutant phenotype and to gain an insight into gene function.The symbiotic gene DNF2 encodes a putative phosphatidylinositol phospholipase C-like protein. Nodules formed by the mutant contain a zone of infected cells reduced to a few cell layers. In this zone, bacteria do not differentiate properly into bacteroids. Furthermore, mutant nodules senesce rapidly and exhibit defense-like reactions.This atypical phenotype amongst Fix(-) mutants unravels dnf2 as a new actor of bacteroid persistence inside symbiotic plant cells. Bourcy M, Brocard L, Pislariu CI, Cosson V, Mergaert P, Tadege M, Mysore KS, Udvardi MK, Gourion B, Ratet P. (2012). New Pytol. article first published online: 27 DEC DOI: 10.1111/nph.12091
Via IvanOresnik
A new year’s resolution for your consideration…
Here is one more resolution for your list: “In 2013, I will communicate about plants to a non-specialist audience.” I think you know WHY (plant science is underfunded, plants are perceived as boring, or less important than animals, etc.), but here are some suggestions for HOW. Participate in Fascination of Plants Day 2013 (http://www.plantday12.eu/home.htm#). Contact your local organizer to find out what’s planned, and either join in or create an event. Here is a summary of the 2012 event to inspire you (http://www.plantday12.eu/downloads2013/Success_EPSOglobal_FoPD2012.pdf). Give a public lecture at a botanic garden, library, senior center, community group or other venue. You can talk about your research or about the roles of plants in our lives. Some of the Teaching Tools materials are easily adapted to a non-specialist audience, including “Why Study Plants?”, “Genetic Improvements in Agriculture”, “Plants, Food and Human Health”, and “Medicinal Plants” (out in Jan 2013). (http://www.plantcell.org/site/teachingtools/teaching.xhtml). Bring plants and a plant-based activity into a school classroom for an hour. Contact your local schools with a concrete proposal, and they’ll help to find a teacher who can work your idea into their curriculum. Many teachers have a very strict curriculum they have to follow, but if you can show them how your presentation fits into their needs they’re usually willing to bring you in as an invited guest. Here are some resources to guide and inspire you: Activities you can do with school children: (http://bit.ly/12SGZ41) and lots more here (http://www.aspb.org/education/NEWK12.CFM) and here (http://www.apsnet.org/edcenter/K-12/Pages/default.aspx) and here (http://www.saps.org.uk/). Here’s a first-person account of a day in the classroom: (http://blogs.scientificamerican.com/culturing-science/2011/08/31/it-only-takes-one-day-bringing-scientists-into-the-classroom/). Here’s a UK resource to help you connect with teachers (https://www.sciencelearningcentres.org.uk/). Or, volunteer to be a mentor for a school group through Planting Science (http://www.plantingscience.org/) or at a school science fair: see more on p. 16 of this ASPB newsletter (http://newsletter.aspb.org/2004/marapr04.pdf). Share a popular science book about plants. You can donate a copy to your public library, school library, or to your favourite high school biology teacher. Here are a few I’ve read recently that I’d like to share: “What a Plant Knows: A Field Guide to the Senses” by Daniel Chamovitz “Seed to Seed: The Secret Life of Plants” by Nicholas Harberd “Hybrid: The History and Science of Plant Breeding” by Noel Kingsbury "The Emerald Planet: How Plants Changed Earth's History” by David Beerling “Eating the Sun” by Oliver Morton “The Secret Life of Trees” by Colin Tudge “Reinventing Life: A Guide to our Evolutionary Future” by Jeffrey Coker “Reaching for the Sun: How Plants Work” by John King “Tomorrows Table” by Pamela Ronald and Raoul Adamchak “The New Oxford Book of Food Plants” by J.G. Vaughan and C.A. Geissler “The Natural History of Medicinal Plants” by Judith Sumner "A Private Life of Plants" by David Attenborough (book and DVD!) (if I missed any of your favorites let me know and I’ll add them!) Plant scientists do have to shoulder a heavier burden of responsibility for communicating about our discipline than do animal biologists, but we also have a strong, supportive community and plenty of well-researched resources to make it easier. Have a very happy, fruitful year!
Via Mary Williams

|
Scooped by
Jean-Michel Ané
December 31, 2012 1:11 PM
|
The interaction between legumes and arbuscular mycorrhizal (AM) fungi is vital to the development of sustainable plant production systems. Here, we focus on a putative MYB-like (LjMAMI) transcription factor (TF) previously reported to be highly upregulated in Lotus japonicus mycorrhizal roots. Phylogenetic analyses revealed that the protein is related to a group of TFs involved in phosphate (Pi) starvation responses, the expression of which is independent of the Pi level, such as PHR1. GUS transformed plants and quantitative reverse transcription PCR revealed strong gene induction in arbusculated cells, as well as the presence of LjMAMI transcripts in lateral root primordia and root meristems, even in the absence of the fungus, and independently of Pi concentration. In agreement with its putative identification as a TF, an eGFP-LjMAMI chimera was localized to the nuclei of plant protoplasts, whereas in transgenic Lotus roots expressing the eGFP-LjMAMI fusion protein under the control of the native promoter, the protein was located in the nuclei of the arbusculated cells. Further expression analyses revealed a correlation between LjMAMI and LjPT4, a marker gene for mycorrhizal function. To elucidate the role of the LjMAMI gene in the mycorrhizal process, RNAi and overexpressing root lines were generated. All the lines retained their symbiotic capacity; however, RNAi root lines and composite plants showed an important reduction in root elongation and branching in the absence of the symbiont. The results support the involvement of the AM-responsive LjMAMI in non-symbiotic functions: i.e. root growth.

|
Scooped by
Jean-Michel Ané
December 30, 2012 1:01 PM
|
Medicago truncatula and Sinorhizobium meliloti form a symbiotic association resulting in the formation of nitrogen-fixing nodules. Nodule cells contain large numbers of bacteroids which are differentiated, nitrogen-fixing forms of the symbiotic bacteria. In the nodules, symbiotic plant cells home and maintain hundreds of viable bacteria. In order to better understand the molecular mechanism sustaining the phenomenon, we searched for new plant genes required for effective symbiosis.We used a combination of forward and reverse genetics approaches to identify a gene required for nitrogen fixation, and we used cell and molecular biology to characterize the mutant phenotype and to gain an insight into gene function.The symbiotic gene DNF2 encodes a putative phosphatidylinositol phospholipase C-like protein. Nodules formed by the mutant contain a zone of infected cells reduced to a few cell layers. In this zone, bacteria do not differentiate properly into bacteroids. Furthermore, mutant nodules senesce rapidly and exhibit defense-like reactions.This atypical phenotype amongst Fix− mutants unravels dnf2 as a new actor of bacteroid persistence inside symbiotic plant cells.

|
Scooped by
Jean-Michel Ané
December 20, 2012 10:50 AM
|
Diversified grasslands that contain native plant species can produce biofuels, support sustainable grazing systems, and produce other ecosystem services. However, ecosystem service production can be disrupted by invasion of exotic perennial plants, and these plants can have soil-microbial “legacies” that may interfere with establishment and maintenance of diversified grasslands even after effective management of the invasive species. The nature of such legacies is not well understood, but may involve suppression of mutualisms between native species and soil microbes. In this study, we tested the hypotheses that legacy effects of invasive species change colonization rates, diversity, and composition of arbuscular-mycorrhizal fungi (AMF) associated with seedlings of co-occurring invasive and native grassland species. In a glasshouse, experimental soils were conditioned by cultivating three invasive grassland perennials, three native grassland perennials, and a native perennial mixture. Each was grown separately through three cycles of growth, after which we used T-RFLP analysis to characterize AMF associations of seedlings of six native perennial and six invasive perennial species grown in these soils. Legacy effects of soil conditioning by invasive species did not affect AMF richness in seedling roots, but did affect AMF colonization rates and the taxonomic composition of mycorrhizal associations in seedling roots. Moreover, native species were more heavily colonized by AMF and roots of native species had greater AMF richness (number of AMF operational taxonomic units per seedling) than did invasive species. The invasive species used to condition soil in this experiment have been shown to have legacy effects on biomass of native seedlings, reducing their growth in this and a previous similar experiment. Therefore, our results suggest that successful plant invaders can have legacies that affect soil-microbial associations of native plants and that these effects can inhibit growth of native plant species in invaded communities.

|
Scooped by
Jean-Michel Ané
December 19, 2012 12:03 PM
|
During the course of evolution, mainly leguminous plants have acquired the ability to formde novo structures called root nodules. Recent studies on the autoregulation and hormonal controls of nodulation have identified key mechanisms and also indicated a possible link to other developmental processes, such as the formation of the shoot apical meristem (SAM). However, our understanding of nodulation is still limited by the low number of nodulation-related genes that have been identified. Here, we show that the induced mutation tricot (tco) can suppress the activity of spontaneous nodule formation 2, a gain-of-function mutation of the cytokinin receptor in Lotus japonicus. Our analyses of tco mutant plants demonstrate that TCO positively regulates rhizobial infection and nodule organogenesis. Defects in auxin regulation are also observed during nodule development in tco mutants. In addition to its role in nodulation, TCO is involved in the maintenance of the SAM. The TCO gene was isolated by a map-based cloning approach and found to encode a putative glutamate carboxypeptidase with greatest similarity to Arabidopsis ALTERED MERISTEM PROGRAM 1, which is involved in cell proliferation in the SAM. Taken together, our analyses have not only identified a novel gene for regulation of nodule organogenesis but also provide significant additional evidence for a common genetic regulatory mechanism in nodulation and SAM formation. These new data will contribute further to our understanding of the evolution and genetic basis of nodulation.

|
Scooped by
Jean-Michel Ané
December 15, 2012 12:01 PM
|
On a recent trip to Ukraine, it became obvious that high soil temperatures and dry surface conditions there have greatly reduced rhizobia populations.

|
Rescooped by
Jean-Michel Ané
from Rhizobium Research
December 14, 2012 9:01 AM
|
Nitrogen fixing wheat 'possible in the next 20 years'FarmersWeeklyThe transfer of genes from legumes and oats into wheat will allow the production of cereal crops capable of fixing nitrogen and with resistance to take-all in the next 20 years,...
Via IvanOresnik
|

|
Rescooped by
Jean-Michel Ané
from Rhizobium Research
February 1, 2013 10:45 AM
|
The intensive application of fertilizers during agricultural practices has led to an unprecedented perturbation of the nitrogen cycle, illustrated by the growing accumulation of nitrates in soils and waters, and of nitrogen oxides in the atmosphere. Besides increasing use efficiency of current N fertilizers, priority should be given to put on value the process of biological nitrogen fixation, through more sustainable technologies that reduce the undesired effects of chemical N fertilization of agricultural crops. Wider legume adoption, supported by coordinated legume breeding and inoculation programs are approaches at hand. Also available are biofertilizers based on microbes that help to reduce the needs of N fertilization in important crops like cereals. Engineering in cereals the capacity to fix nitrogen, either by themselves or in symbiosis with nitrogen-fixing microbes, are attractive future options that nevertheless require more intensive and internationally coordinated research efforts. Although nitrogen-fixing plants may be less productive, at some point agriculture must significantly reduce the use of warming (chemically synthesized) N and give priority to the biological nitrogen fixation, if it is to sustain both food production and environmental health for a continuously growing human population. Olivares J, Bedmar EJ, Sanjuan J.(2013). Mol Plant Microbe Interact. Jan 29. [Epub ahead of print]
Via IvanOresnik

|
Rescooped by
Jean-Michel Ané
from Mycorrhizal fungal genomes
January 28, 2013 8:58 AM
|
Arbuscular mycorrhizal fungi (AMF) represent an ecologically relevant and evolutionarily intriguing group of land plant symbionts, which produce multinucleated spores and hyphae that are currently thought to have propagated clonally for over 500 million years. This long-term absence of sex in AMF is a puzzling evolutionary feature that has sparked scientific interest for some time, but a provoking explanation for their successful evolutionary history in the absence of an obvious sexual cycle is that these organisms may have cryptic sex, or a parasexual life cycle, allowing them to recombine alleles and compensate for deleterious mutations. Interestingly, the recent acquisition of large sequence data from many AMF species can finally allow this hypothesis to be tested more extensively. In this perspective, we highlight emerging evidence based on sequence data for the potential of AMF to have sexual reproduction, and propose a number of routes that could be taken to further explore the presence (or absence thereof) of sex in this poorly studied, yet highly relevant, fungal group. Rohan Riley, Nicolas Corradi Fungal Ecology Volume 6, Issue 1, February 2013, Pages 44–49 http://dx.doi.org/10.1016/j.funeco.2012.01.010
Via Stefano Ghignone

|
Rescooped by
Jean-Michel Ané
from Rhizobium Research
January 28, 2013 8:56 AM
|
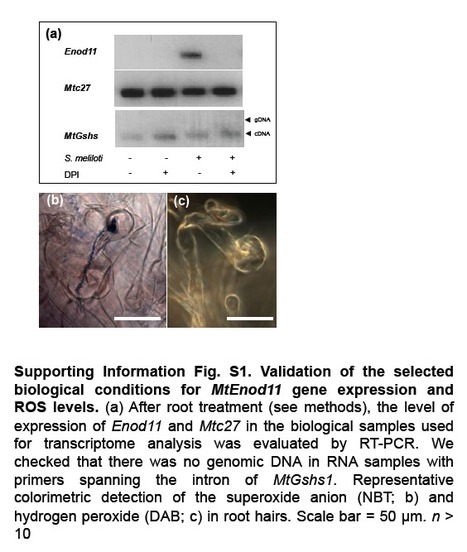
Reactive oxygen species (ROS), particularly hydrogen peroxide (H(2) O(2) ), play an important role in signalling in various cellular processes. The involvement of H(2) O(2) in the Medicago truncatula-Sinorhizobium meliloti symbiotic interaction raises questions about its effect on gene expression. A transcriptome analysis was performed on inoculated roots of M. truncatula in which ROS production was inhibited with diphenylene iodonium (DPI). In total, 301 genes potentially regulated by ROS content were identified 2 d after inoculation. These genes included MtSpk1, which encodes a putative protein kinase and is induced by exogenous H(2) O(2) treatment. MtSpk1 gene expression was also induced by nodulation factor treatment. MtSpk1 transcription was observed in infected root hair cells, nodule primordia and the infection zone of mature nodules. Analysis with a fluorescent protein probe specific for H(2) O(2) showed that MtSpk1 expression and H(2) O(2) were similarly distributed in the nodule infection zone. Finally, the establishment of symbiosis was impaired by MtSpk1 downregulation with an artificial micro-RNA. Several genes regulated by H(2) O(2) during the establishment of rhizobial symbiosis were identified. The involvement of MtSpk1 in the establishment of the symbiosis is proposed. Andrio E, Marino D, Marmeys A, de Segonzac MD, Damiani I, Genre A, Huguet S, Frendo P, Puppo A, Pauly N. (2013). New Phytol. Jan 24. doi: 10.1111/nph.12120. [Epub ahead of print]
Via IvanOresnik

|
Scooped by
Jean-Michel Ané
January 18, 2013 2:39 PM
|
During the early 1980s, Bauer and associates reported that nodulation potential in primary roots of soybean seedlings following inoculation with rhizobia was significantly reduced in developmentally younger regions. They suggested that this phenomenon might be due to a fast-acting regulatory mechanism in the host that prevented excessive nodulation. However, the molecular mechanism of this fast-acting regulatory response remains uncertain. Here, we sought to elucidate components of this regulatory mechanism by investigating the expression of the NSP1and NSP2 genes that encode a GRAS transcription factor required for nodule initiation. First, we confirmed that younger regions of Lotus japonicus roots also show a reduction in nodule numbers in response toMesorhizobium loti. Then, we compared the expression levels of NSP1and NSP2 in developmentally younger regions of primary roots. After inoculation with M. loti, expression of NSP1 was transiently induced whereas that of NSP2 was significantly downregulated 1 day after inoculation. This result implicates that downregulation of NSP2 might cause a fast-acting regulatory mechanism to prevent further nodulation. Next we overexpressed NSP2 in wild type plants. Overexpression resulted in the clustering of nodules in the upper region of the root but strong suppression of nodulation in the lower region. By contrast, overexpression of NSP2 in har1 hypernodulating mutants resulted in an increased number of nodule primordia even in the root tip region. These results indicate that HAR1 negatively regulates NSP2-induced excessive nodule formation in the developmentally younger regions of roots.

|
Rescooped by
Jean-Michel Ané
from Rhizobium Research
January 12, 2013 11:01 AM
|
Calcium (Ca2+) is a key secondary messenger in many plant signaling pathways. One such pathway is the SYM pathway, required in the establishment of both arbuscular mycorrhizal and rhizobial root symbioses with legume host plants.1 When the host plant has perceived the diffusible signals from the microbial symbionts, one of the earliest physiological responses are Ca2+ oscillations in and around the nucleus.2 These oscillations are essential for activating downstream gene expression, but the precise mechanisms of encoding and decoding the Ca2+ signals are unclear and still under intense investigation. Here we put forward a hypothesis for the mechanism of the cation channel DMI1. Myriam Charpentier, Teresa Vaz Martins, Emma Granqvist, Giles Oldroyd and Richard Morris (2013). Volume 8, Issue 2 February eLocation ID: e22894
Via IvanOresnik

|
Scooped by
Jean-Michel Ané
January 1, 2013 4:19 PM
|
Scientist Fighting Microbes with Microbes Scientist Researchers traced communication back to the fungus Glomus mosseae, which forms a symbiotic relationship with plant root hair known as a mycorrhizal network by inserting itself into the root...

|
Scooped by
Jean-Michel Ané
December 31, 2012 1:18 PM
|
In order to accommodate the physiologically incompatible processes of photosynthesis and nitrogen fixation within the same cell, unicellular nitrogen fixing cyanobacteria have to maintain a dynamic metabolic profile in the light as well as the dark phase of a diel cycle. The transition from the photosynthetic to the nitrogen-fixing phase is marked by the onset of various biochemical and regulatory responses, which prime the intracellular environment for nitrogenase activity. Cellular respiration plays an important role during this transition, quenching the oxygen generated by photosynthesis and by providing energy necessary for the process. Although the underlying principles of nitrogen fixation predict unicellular nitrogen fixing cyanobacteria to function in a certain way, significant variations are observed in the diazotrophic behavior of these microbes. In an effort to elucidate the underlying differences and similarities that govern the nitrogen fixing ability of unicellular diazotrophic cyanobacteria, we analyzed six members of the genus Cyanothece. Cyanothece 51142, a member of this genus, has been shown to perform efficient aerobic nitrogen fixation and hydrogen production. Our study revealed significant differences in the patterns of respiration and nitrogen fixation among the Cyanothece strains that were grown under identical culture conditions, suggesting that these processes are not solely controlled by cues from the diurnal cycle but that strain specific intracellular metabolic signals play a major role. Despite these inherent differences, the ability to perform high rates of aerobic nitrogen fixation and hydrogen production appears to be a characteristic of this genus.

|
Scooped by
Jean-Michel Ané
December 30, 2012 1:05 PM
|
Nodulation in legumes requires the recognition of rhizobially made Nod factors. Genetic studies have revealed that the perception of Nod factors involves LysM domain receptor-like kinases, while biochemical approaches have identified LECTIN NUCLEOTIDE PHOSPHOHYDROLASE (LNP) as a Nod factor-binding protein. Here, we show that antisense inhibition of LNP blocks nodulation in Lotus japonicus. This absence of nodulation was due to a defect in Nod factor signaling based on the observations that the early nodulation gene NODULE INCEPTION was not induced and that both Nod factor-induced perinuclear calcium spiking and calcium influx at the root hair tip were blocked. However, Nod factor did induce root hair deformation in the LNP antisense lines. LNP is also required for infection by the mycorrhizal fungus Glomus intraradices, suggesting that LNP plays a role in the common signaling pathway shared by the rhizobial and mycorrhizal symbioses. Taken together, these observations indicate that LNP acts at a novel position in the early stages of symbiosis signaling. We propose that LNP functions at the earliest stage of the common nodulation and mycorrhization symbiosis signaling pathway downstream of the Nod factor receptors; it may act either by influencing signaling via changes in external nucleotides or in conjunction with the LysM receptor-like kinases for recognition of Nod factor.

|
Scooped by
Jean-Michel Ané
December 29, 2012 12:13 PM
|
Researchers find new way to bring poor-quality land into cultivation to grow food even in harsh environments.

|
Scooped by
Jean-Michel Ané
December 19, 2012 3:56 PM
|
To date, little research has been published that quantifies the impact of soil desiccation (e.g. drought) on endemic B. japonicum populations. Pena-Cabriales and Alexander (1979) reported a biphasic decline inRhizobium japonicum numbers in soils undergoing drying. The initial rapid phase of R. japonicum loss coincided with major water loss (simulated drought) whereas the secondary and subsequently slower decline in numbers was governed by the soil water content of specific soils. Pena-Cabriales and Alexander (1979) also noted that organic matter content provided little protection againstR. japonicum desiccation and loss. In a parallel study Oso-Afiana and Alexander (1982) reported similar results when comparing strains of R. japonicum and cowpea rhizobia under desiccation. As one would expect, variation exists among Rhizobium spp. as well as individual isolates within species in their response to soil desiccation, though significant losses were evident regardless of species or isolate (Foulds, 1971; Trotman and Weaver, 1995).
In the Internet era, communicating with friends and colleagues via social networks constitutes a significant proportion of our daily activities. Similarly animals and plants also interact with many organisms, some of which are pathogens and do no good for the plant, while others are beneficial symbionts. Almost all plants indulge in developing social networks with microbes, in particular with arbuscular mycorrhizal fungi, and emerging evidence indicates that most employ an ancient and widespread central ‘social media’ pathway made of signaling molecules within what is called the SYM pathway. Some plants, like legumes, are particularly active recruiters of friends, as they have established very sophisticated and beneficial interactions with nitrogen-fixing bacteria, also via the SYM pathway. Interestingly, many members of the Brassicaceae, including the model plant Arabidopsis thaliana, seem to have removed themselves from this ancestral social network and lost the ability to engage in mutually favorable interactions with arbuscular mycorrhizal fungi. Despite these generalizations, recent studies exploring the root microbiota of A. thaliana have found that in natural conditions, A. thaliana roots are colonized by many different bacterial species and therefore may be using different and probably more recent ‘social media’ for these interactions. In general, recent advances in the understanding of such molecular machinery required for plant–symbiont associations are being obtained using high throughput genomic profiling strategies including transcriptomics, proteomics and metabolomics. The crucial mechanistic understanding that such data reveal may provide the infrastructure for future efforts to genetically manipulate crop social networks for our own food and fiber needs. Muthusubramanian Venkateshwaran, Jeremy D Volkening, Michael R Sussman, Jean-Michel Ané
Via Nicolas Denancé

|
Rescooped by
Jean-Michel Ané
from Mycorrhizal fungal genomes
December 14, 2012 9:01 AM
|
Within the framework of the Mycorrhizal Genomics Initiative, the JGI is sequencing aphylogenetically and ecologically diverse suite of mycorrhizal fungi (Basidiomycota and Ascomycota), which include the major clades of symbiotic species associating with trees and woody shrubs. Analyses of these genomes will provide insight into the diversity of mechanisms for the mycorrhizal symbiosis, including endo- and ectomycorrhiza. This scoop points to the official project page hosted on Mycorweb.
Via Stefano Ghignone
|
 Your new post is loading...
Your new post is loading...
 Your new post is loading...
Your new post is loading...