Your new post is loading...
 Your new post is loading...

|
Scooped by
The Sainsbury Lab
Today, 9:57 AM
|
Bacteria-derived lipopeptides are immunogenic triggers of host defences in metazoans and plants. Root-associated rhizobacteria produce cyclic lipopeptides that activate induced systemic resistance against microbial infection in various plant species. Whether and how these molecules are perceived at the plant cell surface remains elusive. Here we reveal that immune activation in Arabidopsis thaliana by the lipopeptide elicitor surfactin is mediated via a specific interaction with membrane sphingolipids. It relies on host membrane remodelling and subsequent activation of mechanosensitive ion channels. This mechanism leads to host defence potentiation and resistance to the necrotrophic fungus Botrytis cinerea and appears distinct from pattern-triggered immunity induced by classical host pattern recognition receptors. These results reveal a previously uncharacterized mechanism through which lipopeptides derived from non-pathogenic bacteria activate plant immune responses. This work unveils a non-canonical lipid-driven mechanism for plant immune activation by a bacterial lipopeptide. Perception at the cell membrane leads to deformation, mechanosensing and early signalling and culminates in systemic resistance priming.

|
Scooped by
The Sainsbury Lab
April 27, 4:18 AM
|
Dioecious plants, which have distinct male and female individuals, constitute ~5% of angiosperm species and have emerged frequently and independently from hermaphroditic ancestors. Although recent molecular studies of sex determination have started to reveal the diversity of the genetic systems underlying dioecy, research on the evolution of dioecy is limited, especially in monocots. Here, we explore the molecular basis of sex determination in the monocot Dioscorea tokoro, a dioecious wild yam endemic to East Asia. Chromosome-scale and haplotype-resolved genome assemblies and linkage analysis suggested that this plant has a male heterogametic sex-determination (XY) system, with sex-determination regions located on chromosome 3. Sequence comparison between the X- and Y-chromosomes and read coverage analysis revealed X- and Y-specific regions in putative pericentromeric chromosome regions. Within the Y-specific region, we propose two candidate genes that are likely involved in sex determination: BLH9, encoding a homeobox protein, and HSP90, encoding a molecular chaperone. BLH9 functions in a similar way as AtBLH9 in Arabidopsis thaliana. BLH9 could be involved in suppression of female organ development, whereas HSP90 might be required for pollen development. These results shed light on the complex evolution of dioecy in plants.

|
Scooped by
The Sainsbury Lab
April 23, 4:22 AM
|
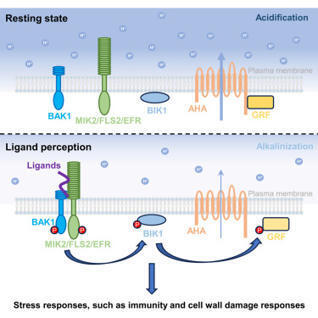
Extracellular alkalinization has long been recognized as a hallmark of plant cell-surface receptor activation, including during pattern-triggered immunity (PTI), yet the mechanisms driving elicitor-induced alkalinization and its role in plant signaling remain unclear. Here, we demonstrate that inhibition of autoinhibited H+-ATPases (AHAs) is required for elicitor-induced extracellular alkalinization. This alkalinization is essential for immune and cell-wall damage signaling mediated by diverse plasma membrane-localized receptor kinases (RKs), likely through modulation of ligand-receptor interactions. Mechanistically, RKs transduce elicitor-triggered signaling via the receptor-like cytoplasmic kinase BOTRYTIS-INDUCED KINASE 1 (BIK1), which inhibits AHA activity by disrupting AHA-GENERAL REGULATORY FACTOR (GRF) interactions through a conserved phosphorylation event. This phosphorylation-driven extracellular alkalinization module is required for disease resistance and cell-wall damage responses initiated by ligand-RK pairs. Our findings uncover a conserved phosphorelay circuit that broadly regulates extracellular alkalinization to coordinate RK signaling, illuminating a general mechanism for RK activation and stress resilience.

|
Scooped by
The Sainsbury Lab
April 8, 4:31 AM
|
Plants secrete a variety of proteases as a defense response during infection by microbial pathogens. However, the relationship between their catalytic activities and antimicrobial functions remains largely unknown. Particularly, few biologically relevant substrates of these proteases have been identified. Huanglongbing (HLB) has been a major threat to the citrus industry worldwide. The HLB-associated bacterium, "Candidatus Liberibacter asiaticus" (Las), was previously shown to deploy an inhibitor of papain-like cysteine proteases (PLCPs) to promote disease in citrus. In this study, we identified an outer membrane protein (OMP) of Las, LasOMP1, as a substrate of the citrus PLCP CsRD21a. LasOMP1 is one of the most highly expressed genes in Las. CsRD21a cleaves LasOMP1 and produces cleaved peptide products, which could be detected in vitro and in HLB-diseased citrus plants. We found that CsRD21a targets the N-terminal portion of LasOMP1, potentially at an extracellular loop region. Importantly, transgenic sweet orange overexpressing CsRD21a showed reduced Las populations and improved plant growth, highlighting that engineering this protease is a promising strategy to enhance HLB resistance in citrus. Together, our work reveals a pathogen-derived substrate of plant PLCPs and suggests bacterial OMPs may be direct targets of plant defense.

|
Scooped by
The Sainsbury Lab
March 31, 11:16 AM
|
The wheat blast fungus Magnaporthe oryzae pathotype Triticum (MoT) poses a severe threat to global wheat (Triticum aestivum L.) production, yet the molecular mechanisms underlying tissue invasion remain poorly understood. We performed dual RNA-seq analysis of MoT-inoculated wheat leaves at 0, 24, 36, and 48 hpi, mapping reads separately to the wheat and M. oryzae genomes to capture stage-specific host responses and pathogen gene expression across progressive infection stages. Wheat exhibited pronounced stage-specific transcriptional reprogramming, with peak differential gene expression at 36 hpi and visible symptoms at 48 hpi. The 24 hpi stage was characterized by rapid induction of immune- and defense-related pathways, including innate immunity and detoxification processes, along with downregulation of cell wall and membrane biosynthesis. By 36 hpi, wheat maintained sustained activation of immune and detoxification pathways, while chloroplast- and photosynthesis-associated genes were broadly repressed, consistent with transcriptional features of metabolic constraint. At 48 hpi, coinciding with lesion initiation, transcriptomes showed persistent, metabolically costly immune and defense responses together with extensive suppression of photosynthesis- and chloroplast-associated functions, which were associated with metabolic strain and a transition toward necrosis. Analysis of pathogen-derived reads revealed temporal induction of multiple effector candidates, including known M. oryzae orthologs and additional effector-like proteins, highlighting coordinated temporal patterns between host immune and metabolic response as well as stage-specific pathogen effector expression. Together, these findings provide a temporal framework for wheat blast susceptibility and highlight key host pathways and effector candidates that define critical windows for functional dissection of MoT virulence and wheat susceptibility.

|
Scooped by
The Sainsbury Lab
March 31, 11:10 AM
|
Photoperiodism, a seasonal response mechanism that relies on day-length measurement by the circadian clock, is a major regulator of flowering time (Imaizumi 2025). The external coincidence model proposes that the alignment between the internal circadian phase and external light signals triggers photoperiodic responses (Pittendrigh and Minis 1964). It has been challenging to establish robust models for the quantitative relationship between clock and flowering time, due to a lack of tools to modulate the clock quantitatively. Here, using a circadian period-lengthening small molecule, we demonstrated the quantitative modulation of the critical day-length for flowering in monocots.

|
Scooped by
The Sainsbury Lab
March 31, 11:04 AM
|
The accelerating growth of plant science knowledge presents a major challenge for researchers seeking to extract accurate, up-to-date knowledge from an increasingly fragmented and domain-specific corpus. General-purpose large language models (LLMs), while powerful, often misinterpret plant science terminology and lack mechanisms for source traceability. We created PlantScience.ai, a virtual plant biology scientist powered by our automated scientific knowledge graph construction pipeline (AutoSKG). PlantScience.ai exhibits expert-level reasoning in plant biology and maintains scholarly rigour in its citations. Through continuous learning, it integrates the latest research, ensuring that its knowledge base remains current and scientifically robust. Apart from providing the answers to the scientific questions, PlantScience.ai can interact with human scientists, follow instructions, and retrieve information with citation awareness, grounding each response in primary sources to ensure accuracy and verifiability. PlantScience.ai marks a pivotal advance toward a collaborative scientific paradigm in which virtual and human plant scientists work synergistically to accelerate discovery while preserving the unique value of human insight. PlantScience.ai is available at https://plantscience.ai.

|
Scooped by
The Sainsbury Lab
March 10, 6:59 AM
|
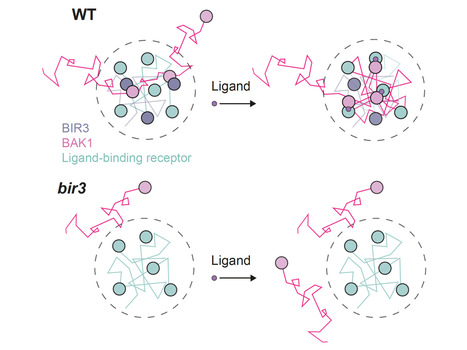
Plasma membrane-localized receptors operate as dynamic signaling complexes and integrative networks, yet the spatial and temporal regulation of these interactions remain largely unknown. Here, by analyzing the components of a minimal Arabidopsis leucine-rich repeat receptor kinase network, we describe the differential diffusion and organization of receptor complex components and unveil the nanoscale spatial and temporal logic underlying the formation of receptor kinase complexes. The ligand-binding receptors FLS2 and BRI1, and the accessory receptor BIR3, are organized in plasma membrane nanodomains, within which the co-receptor BAK1 diffuses and is spatially arrested upon ligand perception. BAK1 spatial arrest relies on extracellular domain (ECD)-ECD interactions but does not require receptor complex activation. Mathematical modelling, single molecule imaging and bio-assays infer that accessory receptors maintain a dynamic pool of co-receptors in the vicinity of ligand-binding receptors to promote ligand-induced complex formation and signaling. We propose that ligand-induced receptor kinase complex formation is a deterministic process defined by the relative nanoscale spatial positioning of individual signaling and regulatory components.

|
Scooped by
The Sainsbury Lab
March 4, 6:42 AM
|
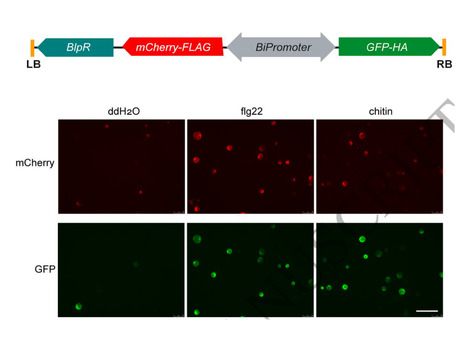
Immune activation in plants triggers extracellular alkalinization, presumably by inhibiting plasma membrane H+-ATPases. The precise role and underlying mechanisms of this process remain poorly understood. Here, we show that Pseudomonas syringae bacteria induce apoplastic alkalinization not only at the site of infection but also in neighboring distal tissues to prime defenses and disease resistance in Arabidopsis. We show that several calcium-dependent protein kinases phosphorylate Ser899 of two major autoinhibited H+-ATPases to dampen their activity, leading to alkalinization. The distal alkalinization is accompanied by the transcriptional activation of phytocytokines, including plant elicitor peptides, serine-rich endogenous peptides, and their receptors. We show that these phytocytokines promote distal alkalinization and disease resistance, whereas the apoplastic alkalinization sensitizes the phytocytokine perception that further induces phytocytokine genes. Our study suggests that apoplastic alkalinization and phytocytokine gene expression mutually potentiate and act as a combined signal that propagates in local-distal communication and disease resistance priming.

|
Scooped by
The Sainsbury Lab
February 20, 6:33 AM
|
Trans-species RNA interference (tsRNAi), in which plants produce small RNAs (sRNAs) to silence target genes in pathogens, has emerged as a promising strategy for disease control. However, whether tsRNAi constitutes an endogenous, regulated immune response remains unclear. Here, we show that ARGONAUTE10 (AGO10) plays a critical role in pathogen-induced tsRNAi. Loss of AGO10 in Arabidopsis abolished pathogen gene silencing during infection, leading to hypersusceptibility to oomycete and fungal pathogens. Importantly, AGO10 rapidly responds to pathogen infection through increased protein accumulation and re-location into discrete cytoplasmic condensates, thus promoting the production of trans-species sRNAs at the pathogen infection sites. This immune responsiveness relies on the N terminal intrinsically disordered region (IDR) of AGO10, which is responsible for sensing and responding to immune activation. Specific features in the IDR partitions AGO10 into two deeply diverged subgroups, AGO10a and AGO10b, with the immune responsiveness and defense function evolutionarily conserved in AGO10a but not AGO10b. Together, these findings establish tsRNAi as a bona fide, evolutionarily conserved immune response and position AGO10 as a signal-responsive hub linking pathogen perception to tsRNAi-based defense.

|
Scooped by
The Sainsbury Lab
February 17, 6:46 AM
|
Transposon silencing via RNA-directed DNA methylation (RdDM) is mediated primarily through 24-nucleotide (nt) small interfering RNAs (siRNAs) and two plant-specific RNA polymerases, Pol IV and Pol V. We generated and characterized RNA-interference (RNAi) lines targeting the largest subunit of Pol IV and Pol V and the second-largest subunit shared by Pol IV and Pol V in soybean. Our analyses showed that the canonical roles of Pol IV and Pol V in RdDM as described in Arabidopsis, whereby Pol IV produces 24-nt siRNAs and Pol V recruits siRNAs to homologous loci to trigger DNA methylation, are conserved in soybean. Our analyses also uncovered functions of Pol IV and Pol V in that they repress defense response genes en masse. These genes normally undergo RdDM and are silenced, but they are de-repressed when Pol IV and Pol V are knocked down. Furthermore, the de-repression of a set of defense-related genes channeled their RNAs into the RNAi pathway to produce 21-22-nt siRNAs. Knocking down Pol IV and Pol V either singly or together led to increased resistance against the oomycete pathogen Phytophthora sojae, suggesting that Pol IV- and Pol V-mediated gene silencing regulates plant immunity.

|
Scooped by
The Sainsbury Lab
February 11, 11:33 AM
|
Resistance gene enrichment sequencing (RenSeq) was developed in 2013. It has accelerated the cloning of plant NLR genes and has contributed to resistance breeding for multiple crop plants, such as potato, wheat, and rice. By combining with other strategies, many applications were developed, such as cDNA-RenSeq, dRenSeq, SMRT-RenSeq, RLP/KSeq, AgRenSeq, and MutRenSeq. These methods have been widely applied in different crops. In this protocol, we present a step-by-step guide for applying RenSeq in gene cloning and Pan-NLRome analysis. The protocol covers bait design, library preparation, target enrichment, and downstream bioinformatic analysis. This methodology can make RenSeq more accessible to researchers working with different crops and enhance our understanding of plant resistance genes in the age of pan-genome.

|
Scooped by
The Sainsbury Lab
February 11, 5:32 AM
|
Herbivorous insects can shape the epidemiology of disease in plants by vectoring numerous phytopathogens. While the consequences of infection are often well-characterized in the host plant, the extent to which phytopathogens alter the physiology and development of their insect vectors remains poorly understood. In this review, we highlight how insect-borne phytopathogens can promote vector fitness, consistent with theoretical predictions that selection should favor a mutualistic or commensal phenotype. In doing so, we define the metabolic features predisposing plant pathogens to engage in beneficial partnerships with herbivorous insects and how these mutualisms promote the microbe's propagation to uninfected plants. For the vector, the benefits of co-opting microbial pathways and metabolites can be immense: from balancing a nutritionally deficient diet and unlocking a novel ecological niche to upgrading its defensive biochemistry against natural enemies. Given the independent origins of these tripartite interactions and a number of convergent features, we also discuss the evolutionary and genomic signatures underlying microbial adaptation to its dual lifestyle as both a plant pathogen and an insect mutualist. Finally, as host association can constrain the metabolic potential of microbes over evolutionary time, we outline the stability of these interactions and how they impact the virulence and transmission of plant pathogens.
|

|
Scooped by
The Sainsbury Lab
April 28, 7:52 AM
|
Plant diseases remain a major threat to food security, reducing crop yield worldwide. A central challenge in crop protection is that effective immunity often comes at a cost. Plants must continuously balance growth with defense, and this balance complicates molecular breeding strategies aimed at improving disease resistance. When immune pathways are constitutively activated, plants are often better protected against pathogens, but the price for this protection is reduced growth or fitness. This defense-growth trade-off has therefore become a major obstacle to engineering disease resistance in crops.

|
Scooped by
The Sainsbury Lab
April 23, 8:54 AM
|
When organisms encounter pathogens, they rapidly activate complex defense programs to ensure survival. While these immune responses are vital, they often also incur trade-offs, such as reduced growth and development and must therefore be tightly controlled. In this study, we reveal that the steroid hormones brassinosteroids (BRs) contribute to this control in Arabidopsis thaliana by repressing immunity-related genes. We provide evidence that the BR-regulated basic helix-loop-helix (bHLH) transcription factor CESTA (CES), along with its homologs BR ENHANCED EXPRESSION (BEE)1-3, mediate DNA methylation changes at transposable element (TE)-rich loci containing nucleotide-binding leucine-rich-repeat (NLR)-type receptor genes, including SUPPRESSOR OF NPR1-1 CONSTITUTIVE 1 (SNC1). These CES-induced methylation changes correlate with altered splicing of SNC1 pre-mRNA, a process that requires the BR receptor BRASSINOSTEROID INSENSITIVE 1 (BRI1). In support, we show that CES associates with components of the chromatin remodeling and splicing machinery. Together, our findings reveal a previously unrecognized BR-induced mechanism that modulates the epigenetic and post transcriptional regulation of immune genes, enabling plants to prioritize growth over defense.

|
Scooped by
The Sainsbury Lab
April 22, 6:27 AM
|
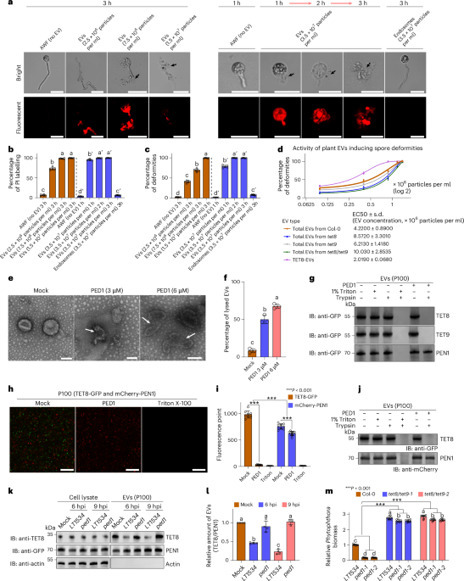
Extracellular vesicles (EVs) transport biologically active molecules and are known to mediate host defence against microbial pathogens, including plant fungal pathogens. However, the mechanism by which pathogens disrupt EV-dependent defence remains unclear. Here we show that Phytophthora capsici, a global crop pathogen, counteracts EV-mediated plant defence through targeted lipase activity. We show that Arabidopsis releases EVs containing tetraspanin (TET), specifically TET8- and TET9-EVs, which damage germinated spores of Phytophthora, reducing infection. As a counter-defence, Phytophthora secretes an infection-induced apoplastic lipase, Plant Extracellular Vesicle Destroyer 1 (PED1), which targets TET8- and TET9-EVs. This occurs via interaction with the EV membrane-localized protein Defective Glycosylation 1 (DGL1), which directly interacts and co-localizes with TET8 and TET9 on the EV membrane. PED1 damages TET8- and TET9-EVs through its lipase activity towards campesteryl esters, suppressing EV-mediated plant defence. Our study reveals a mechanism used by Phytophthora to counteract EV-mediated host defence. Phytophthora secretes an infection-induced apoplastic lipase, damaging Arabidopsis extracellular vesicles and suppressing plant defence against infection.

|
Scooped by
The Sainsbury Lab
April 7, 6:09 AM
|
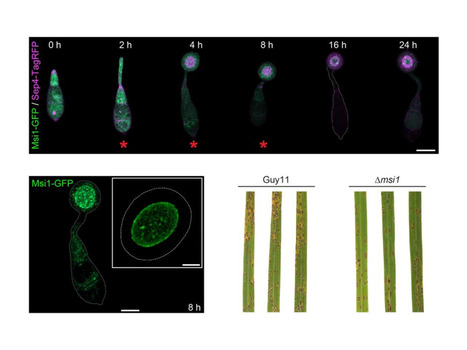
Septin GTPases are essential cytoskeletal regulators that organize membranes and scaffold protein complexes to control cytokinesis, polarity, and morphogenesis. How septins execute these functions remains poorly understood, and comprehensive, stage-resolved interaction maps are lacking. Here, we define a quantitative, time-resolved septin interactome in the rice blast fungus Magnaporthe oryzae using immunoprecipitation coupled to mass spectrometry. We map more than 350 interactors of septins Sep3, Sep4, Sep5 and Sep6, revealing a dynamic network required for appressorium-mediated plant infection. Beyond canonical roles in cytoskeletal organisation and polarity, septins associate with proteins linked to membrane remodelling, metabolism, and virulence, deployed during host invasion. Integration with ultra-high-throughput yeast two-hybrid analysis defines a high-confidence septin interactome and identifies previously uncharacterised factors, including Msi1, a BAR domain protein required for invasive growth. Together, these findings establish septins as dynamic organisers of infection-related processes and provide a framework for understanding how cytoskeletal scaffolds coordinate fungal pathogenesis.

|
Scooped by
The Sainsbury Lab
March 31, 11:12 AM
|
Nucleotide-binding, leucine-rich repeat (NLR) receptors are widespread intracellular immune sensors across kingdoms. Plant G10-type coiled-coil (CCG10)-NLRs constitute a distinct phylogenetic clade that remains poorly characterized. Here, we identified a gain-of-function mutant of wheat autoimmunity 3 (WAI3GOF), which encodes a constitutively active CCG10-NLR resulting from a residue substitution in the leucine-rich repeat (LRR) domain. Cryo-electron microscopy (cryo-EM) analysis reveals that activated WAI3 assembles into a distinctive octameric resistosome. Arabidopsis RPS2, another CCG10-NLR, also forms an octamer, indicating a conserved structural property across monocot and dicot plants. The WAI3 resistosome induces a prolonged and sustained increase in cytosolic calcium, likely facilitated by a unique channel architecture arising from its divergent coiled-coil (CC) domain configuration. Notably, this domain arrangement may be shared by plant NLRs that lack the conserved EDVID (Glu-Asp-Val-Ile-Asp) motif in their CC domains. Together, our findings uncover a conserved yet previously uncharacterized NLR resistosome structure and provide insights into the plant immune receptor plasticity.

|
Scooped by
The Sainsbury Lab
March 31, 11:07 AM
|
The wheat resistance gene Pm4 encodes a kinase fusion protein and has gained particular attention as it confers race-specific resistance against two major wheat pathogens: powdery mildew and blast. Here we describe the identification of AvrPm4, the mildew avirulence effector recognized by Pm4, using UV mutagenesis, and its functional validation in wheat protoplasts. We show that AvrPm4 directly interacts with and is phosphorylated by Pm4. Using genetic association and quantitative trait locus mapping, we further demonstrate that the evasion of Pm4 resistance by virulent mildew isolates relies on a second fungal component, SvrPm4, which suppresses AvrPm4-induced cell death. Surprisingly, SvrPm4 was previously described as AvrPm1a. We show that SvrPm4, but not its inactive variant svrPm4, is recognized by the nucleotide-binding leucine-rich repeat immune receptor Pm1a. These multiple roles of a single effector provide a new perspective on fungal (a)virulence proteins and their combinatorial interactions with different types of immune receptors. AvrPm4 is a chimeric powdery mildew effector that is recognized by the wheat kinase fusion resistance protein Pm4. The pathogen can evade Pm4 recognition by expressing the suppressor SvrPm4, an RNase-like effector with multiple roles.

|
Scooped by
The Sainsbury Lab
March 16, 7:52 AM
|
Increasing evidence suggests that lichens are not just a partnership between one fungus and one alga or cyanobacterium but may contain multiple interacting microorganisms. A recent study reveals the presence of a ubiquitous black fungus in a group of lichens, including their reproductive structures, suggesting it may be a previously unknown symbiont.

|
Scooped by
The Sainsbury Lab
March 9, 6:33 AM
|
Plant cell wall (CW) integrity signaling enables early detection of microbial invasion, yet the receptors involved and their spatial and temporal dynamics during infection remain largely unknown. We identify ATHENA (ATHE)/MEE39, a previously uncharacterized malectin‑like leucin-rich repeat receptor kinase (Mal-LRR-RK) that contributes to defense against the root vascular pathogen Fusarium oxysporum (Fo), particularly in outer root layers where colonization begins. ATHE abundance, localization, and endocytic trafficking are rapidly remodeled during infection, and loss of ATHE compromises basal immunity and early pathogen‑induced transcriptional reprogramming. ATHE responds to altered cellulose synthesis, cellulose‑derived oligosaccharides, mechanically induced CW perturbations, and the fungal secreted peptide Fo‑RALF. In most of these contexts, ATHE acts together with the LRR-RK MIK2, forming a pathogen‑strengthened RK complex that fine‑tunes root responses to Fo. This represents the first example of a receptor complex visualized subcellularly in vivo during a plant-microbe interaction. Although Brassicaceae‑specific, heterologous expression of ATHE enhanced tomato resistance to Fo, highlighting its functional relevance across plant lineages and its potential use for crop engineering. Our work reveals a previously unrecognized strategy by which plants decode microbial threats through dynamic CW‑integrity surveillance.

|
Scooped by
The Sainsbury Lab
February 26, 8:11 AM
|
Effective management of Phytophthora root rot in soybean is compromised by the rapid loss of efficacy of the most widely deployed resistance genes to Phytophthora sojae (Rps). However, some genes such as Rps3a and Rps6 are still offering a strong protection but are nonetheless rare in elite material, probably owing to the fact that they have never been properly characterized. In this study, we have employed RenSeq, and whole genome sequencing to unravel the nature of Rps6 as a complex locus composed of 10 NLR genes. In a cell death assay using soybean protoplasts, we show that one of the candidates from the cluster interacts robustly with Avr6. Transfer of the candidate gene into a susceptible root system confirmed its function and status as the bona fide Rps6. Through sequence comparison with other Rps differentials, we further discovered that Rps3c and Rps4, originally thought to be distinct genes on different chromosomes, are, in fact, the exact same resistance gene as Rps6, mediating recognition to the same effector, Avr6. These results clarify a long-standing confusion regarding the identity of some elusive Rps genes and offer the precise sequence and position of Rps6. At the same time, along with the recently exposed homology between Rps3b and Rps11, these findings raise some concerns about the apparent multiplicity of Rps genes. Indeed, there may be fewer sources of resistance than assumed, which should instill caution in using current Rps genes to ensure durable management of Phytophthora root rot.

|
Scooped by
The Sainsbury Lab
February 18, 4:58 AM
|
Plant cell surface pattern recognition receptors (PRRs) perceive non- or altered-self elicitors to induce immune responses. PRRs relay information across the plasma membrane and trigger downstream signalling via receptor-like cytoplasmic kinases such as BOTRYTIS-INDUCED KINASE 1 (BIK1). BIK1 associates with several PRRs and acts as a key executor of immune responses through the phosphorylation of substrate proteins. However, a comprehensive understanding of how BIK1 targets specific substrates and a full repertoire of these substrates are lacking. Here we defined the substrate specificity of BIK1 and used these data to predict candidate substrates in Arabidopsis. Using high-throughput biochemical and genetic screening of these candidates, we confirmed many as direct BIK1 substrates in vitro and novel regulators of plant immunity. Among the BIK1 substrates identified are MULTIPLE C2 DOMAIN AND TRANSMEMBRANE REGION PROTEIN 3, which we reveal regulates flagellin 22 (flg22)-induced plasmodesmata closure and immunity, and members of the largely uncharacterized CYCLIN-DEPENDENT KINASE-LIKE family, which we uncover as novel negative regulators of immunity. In parallel, we interrogated intracellular NUCLEOTIDE-BINDING LEUCINE-RICH REPEAT (NLR) immune receptors for potential BIK1 phosphorylation motifs and identified multiple NLRs as direct BIK1 substrates. We reveal that BIK1 phosphorylation regulates NLR oligomerization, thus controlling a key activation step for these immune receptors. Together, our unbiased biochemical screens shed light on the central role of BIK1 as a key kinase shaping multiple layers of plant immune signalling. Cell surface receptors perceive immunogenic elicitors, triggering downstream signalling via receptor-like cytoplasmic kinases such as BIK1. Here the authors define and use the phosphorylation motif of BIK1 to find novel substrate candidates.

|
Scooped by
The Sainsbury Lab
February 12, 4:54 AM
|
Single-nucleus omics technologies are increasingly applied in plant biology to investigate transcriptional and chromatin regulation at cellular resolution. A key requirement for these approaches is the isolation of high-quality nuclei, which can be particularly challenging in plant tissues due to cell wall complexity and residual debris. Here, I present a streamlined protocol for isolating nuclei from fresh Arabidopsis thaliana leaves infected with the bacterial pathogen Pseudomonas syringae. This method eliminates the need for density gradient separation and fluorescence-activated nuclei sorting (FANS), reducing hands-on time while maintaining compatibility with single-nucleus RNA and ATAC sequencing using the 10x Genomics platform.

|
Scooped by
The Sainsbury Lab
February 11, 5:34 AM
|
Plant nucleotide-binding leucine-rich repeat (NLR) immune receptors typically confer resistance through recognition of specific pathogen effectors. The Arabidopsis NLR WRR4A defies this paradigm by recognizing multiple sequence-divergent effectors from Albugo candida, conferring resistance to multiple pathogen races. Despite minimal sequence similarity, these effectors share a conserved N-terminal ferredoxin-like fold. Through cryo-EM structure determination of two WRR4A resistosomes bound to sequence-distinct effectors, combined with AlphaFold modelling, we reveal a shape-based recognition mechanism: WRR4A engages structurally conserved backbone features of the effectors in a mostly side chain-independent manner, enabling recognition of diverse effectors with similar three-dimensional architectures. These insights guided successful engineering of WRR4A to acquire novel recognition specificity. In addition, analysis of the monomeric WRR4A resting state reveals a distinct domain architecture characteristic of C-JID–containing TIR-NLRs and informs their activation mechanism. This work provides insights into NLR-mediated broad-spectrum recognition and the potential for structure-informed engineering of improved crop resistance.
|






 Your new post is loading...
Your new post is loading...