Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 3:02 PM
|
Long before nature was ‘red in tooth and claw’, it was sundered by nano-spears and seeping poisons. Microorganisms were the first predators, and predation has since deeply shaped all major branches of life — from individual traits to collective systems, community dynamics and major evolutionary transitions. Yet, we have only begun to understand how microbial predation influences the genetics, morphology, behavior, ecology and evolution of microorganisms in natural communities and, in turn, the macroscopic biosphere. With the field advancing rapidly on diverse fronts, integrative conceptual frameworks, questions and research approaches are needed to promote synthetic development of the field. In this Review, we explore the remarkably diverse forms of microbial predation that have evolved so far, considering organismal traits and their molecular foundations alongside the evolutionary ecology of predator–prey interactions in community contexts. Building on a process-based definition, forms of microbial predation are conceptualized along gradients, including gradients of evolutionary adaptedness for predation and of privatization of prey-derived nutrients. Important future research themes include predation origins and early stages of predatory adaptation, effects of diverse forms of predation on community diversity and stability, predator–prey co-evolution in complex communities, and multi-approach development of unicellular predators as biocontrol agents. In this Review, Vasse and Velicer explore the phylogenetic and functional diversity of predators of microorganisms, conceptualizing the forms of microbial predation along gradients, including gradients of evolutionary adaptedness for predation and privatization of prey-derived nutrients.

|
Scooped by
mhryu@live.com
Today, 2:48 PM
|
The probiotics field, a historically popular yet scientifically debated discipline, is moving beyond a decades-long promotion of ‘first-generation’ food-derived strains towards the development of ‘next-generation probiotics’ (NGP) or ‘precision probiotics’, natural and engineered strains featuring improved human colonization, clinical efficacy and safety profiles. In this Review, we outline the evolution of NGP and means by which their development is designed to tackle challenges of live bacterial therapy related to colonization resistance, in-host evolution, long-term safety and insufficient understanding of therapeutic and off-target mechanisms of activity. We showcase how a variety of emerging strategies enable the identification of NGP strains and define consortia featuring therapeutic potentials in metabolic, immune and oncological diseases. Finally, we discuss how computational and artificial intelligence (AI) advances can reshape NGP development, including AI-based discovery of strains and bioactive compounds; computational-driven design of engineered microorganisms and multi-kingdom consortia; and AI-assisted structural and metabolic network-based modelling predicting personalized NGP function, interactions and therapeutic impacts. In this Review, Kern, Tofield, Frame and Elinav discuss recent advances in the design of next-generation probiotics, from identification of candidates to therapeutic applications across diverse disease contexts, and highlight major challenges and the potential of artificial intelligence to develop effective, personalized probiotics with therapeutic functions.

|
Scooped by
mhryu@live.com
Today, 1:56 PM
|
Chitin and its deacetylated derivative, chitosan, are natural polymeric polysaccharides derived from crustaceans, fungi, and insects with antimicrobial, anti-inflammatory, antioxidant, and biocompatible bioactivities. As a complement to crustacean biomass, fungi provide a sustainable source of chitin/chitosan. Recent biotechnology and materials science advancements have stimulated significant research interest in applying fungal-derived chitin and chitosan for wound healing materials. This paper comprehensively reviews fungal chitin/chitosan-based materials for wound healing applications, encompassing innovative forms, such as nanoparticles, membranes, hydrogels, sponges, microfibrous nonwovens, and nanofibrous scaffolds. Moreover, this Review elucidates the limitations of using fungal chitin and chitosan in wound healing applications, including low purity and extraction yield, inadequate quality control measures, suboptimal biocompatibility, and insufficient evaluation of action mechanisms. To advance the innovation and commercialization of fungal-derived chitin- and chitosan-based wound healing materials, it is imperative to enhance multidisciplinary collaboration among microbiology, chemistry, materials science, and clinical medicine.

|
Scooped by
mhryu@live.com
Today, 1:50 PM
|
The accelerating global crisis of antimicrobial resistance (AMR) demands rapid and accurate methods for antibiotic susceptibility testing (AST). Conventional phenotypic assays remain the gold standard but are hindered by long culture times, while genotypic tests cannot reliably predict phenotypic resistance. In recent years, metabolism-based AST has emerged as a promising alternative, enabling the rapid detection of bacterial responses to antibiotics through shifts in metabolic activity. These approaches bridge molecular speed with phenotypic precision, allowing susceptibility determination within hours, or even minutes, without requiring cell proliferation. In this review, we summarize the latest advances in metabolism-based biomarkers for rapid AST. First, we discuss how antibiotics influence bacterial metabolism, linking resistance mechanisms to metabolic activities. We then summarize emerging metabolic biomarkers, categorized by their physiological underpinnings: nutrient uptake, respiratory activity, metabolic reprogramming, and enzymatic function. Finally, we list key challenges and future directions toward deployable metabolism-based AST platforms.

|
Scooped by
mhryu@live.com
Today, 1:33 PM
|
mRNA-based therapeutics have revolutionized the treatment and prevention of infectious, neurological, and cancer diseases. However, their linear topology makes them susceptible to rapid degradation in vivo, which limits their therapeutic efficacy. Engineered circular RNAs (circRNAs) due to their closed ends and high stability are emerging as a promising alternative to linear RNA therapies. Engineered circRNAs are also increasingly used to mimic naturally occurring circRNAs in functional studies. Both applications, however, depend on production of precise circRNAs with homogenous sequences to enable accurate interpretation of biological outcomes. To address this, we developed and optimized methods for generating precise circRNAs. We employed enzymatic ligation of linear RNAs rather than autocatalytic splicing to produce circRNAs to minimize extraneous nucleotides remaining from the ribozymes. We carefully designed the DNA transcription template to maintain sequence and structural integrity. A permuted DNA template leveraging three internal guanosines (Gs) was synthesized and amplified using a reverse primer containing two 2′-O-methyl groups. This approach optimally produced the linear precursor RNA with correct 5′ and 3′ ends. After testing multiple workflows, we found that GMP-primed in vitro transcription, T4 RNA ligase 2-mediated circularization, and urea–polyacrylamide gel electrophoresis (PAGE) gel extraction produced the highest fidelity circRNAs.

|
Scooped by
mhryu@live.com
Today, 11:20 AM
|
Natural products, as metabolites from microorganisms, animals or plants, exhibit diverse biological activities, making them crucial for drug discovery. Nowadays, existing deep-learning methods for natural products research primarily rely on supervised learning approaches designed for specific downstream tasks. However, such one-model-for-a-task paradigm often lacks generalizability and leaves substantial room for performance improvement. Additionally, existing molecular characterization methods are not well-suited for the unique tasks associated with natural products. Here, to address these limitations, we pretrained a foundation model for natural products (NaFM) based on their unique properties. Our approach employs a pretraining strategy specifically tailored to natural products. By incorporating contrastive learning and masked graph learning objectives, we emphasize evolutional information from molecular scaffolds while capturing side-chain information. NaFM achieves state-of-the-art results in various downstream tasks related to natural product mining and drug discovery. We first compare taxonomy classification with synthetic molecule-focused baselines to demonstrate that current models are inadequate for understanding natural synthesis. Furthermore, by diving into a fine-grained analysis at both the gene and microbial levels, NaFM demonstrates the ability to capture evolutionary information. Eventually, our method is applied to virtual screening, illustrating informative natural product representations that can lead to more effective identification of potential drug candidates. Ding et al. present a scaffold-aware foundation model for small-molecule natural products leveraging masked objectives and contrastive learning to enhance taxonomy classification, genome mining and virtual screening in drug discovery.

|
Scooped by
mhryu@live.com
Today, 9:20 AM
|
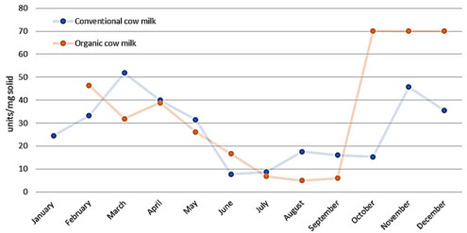
Milk is one of the most vital foods worldwide, valued not only for its nutrient-rich composition but also for its diverse range of bioactive compounds. In addition to its nutritional importance, milk contains a variety of proteins that serve significant biological functions. Among these are lactic acid bacteria (LABs), which can produce antimicrobial peptides and organic acids. The antimicrobial effects of milk are primarily attributed to its bioactive proteins, including whey proteins and caseins. Whey proteins, such as lactoferrin, lysozyme, and immunoglobulins, as well as peptides derived from these proteins, exhibit significant antimicrobial activity, particularly against Gram-positive and Gram-negative bacteria. These peptides are released during proteolysis, either through enzymatic digestion or fermentation, and can interact with bacterial membranes, destabilising them and preventing microbial growth. The concentration of antimicrobial proteins varies across mammalian milks, with higher levels often observed in species such as sheep and goats, reflecting adaptations to specific environmental and immune challenges. Despite the reduction in antimicrobial efficacy following heat treatments like ultra-high temperature (UHT) or pasteurisation, fermented dairy products such as yoghurt and cheese retain significant antimicrobial properties, mainly due to the presence of bioactive peptides and increased acidity. These antimicrobial activities underscore the potential of milk-derived compounds as natural alternatives to antibiotics, particularly in food safety and therapeutic applications. Further research into milk’s bioactive peptides could expand their use in the prevention and treatment of microbial infections.

|
Scooped by
mhryu@live.com
Today, 1:10 AM
|
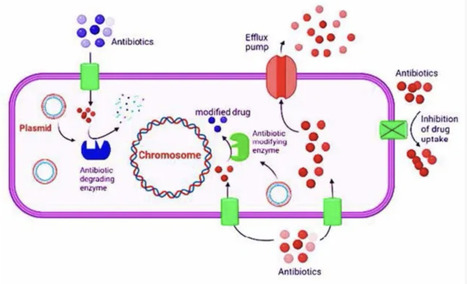
Bacteria display an incredible genetic plasticity, which enables them to adapt to various environmental stressors, such as antibiotic compounds that may imperil their existence. Bacterial resistance mechanisms include degradative enzymes, inactivation of antibiotics, antibacterial target site mutation, change in target, altered cell wall permeability to antibiotics, bypass of metabolic pathways, and efflux pumping of antibiotics across the cell membrane. These mechanisms are encoded by genomic changes ranging from point mutation via genetic elements assembly to horizontal transfer of genes from the environment. Antibiotic resistance in bacteria can be inherited or acquired. Antibacterial resistance genes may accumulate mobile elements, leading to multi-drug-resistant phenotypic transfer via a single genetic event. The resistance to antibiotics has been frequently increasing in clinical settings, which drives scientists to research alternative antibacterial medicines to prevent the growth and spread of drug-resistant bacteria. Technological advancements and the discovery of innovative drug moieties with targeting potential have led to the development of novel drug compounds with diverse therapeutic properties. This includes alternative cellular, physiological, and metabolic patterns of bacteria that may be potential pharmacological targets for the next generation of antibiotics. It is beneficial to characterize antibiotic resistance genotypes and phenotypes causing antibiotic bacterial resistance. An understanding of mechanisms that lead to the development and spread of antibiotic resistance will help clinicians in making appropriate decisions regarding antibiotic usage in a wide range of circumstances. The current review has highlighted the mechanism of drug resistance in bacteria, and has enlisted the antibiotic resistance genes (ARGs) and their importance in aggravating the resistance phenomenon.

|
Scooped by
mhryu@live.com
Today, 12:15 AM
|
Engineered protein chimeras enable new biological functions but remain difficult to design due to context-dependent constraints on insertion tolerance and the need to preserve host protein function. Here, we report CRISPR-guided protospacer adjacent motif (PAM) scanning in yeast to map chimera-permissive sites in living cells. We apply this approach to peptide and reporter insertions. In the first application, we generated 91 insertion chimeras encoding a defined protease cleavage sequence across six components of a model G protein-coupled receptor (GPCR) signaling pathway. Sixty-three percent of sites retain signaling, identifying positions that preserve host function and reveal broad, position-dependent tolerance. Coupling insertional scanning with cognate proteases enables site-resolved mapping of in-cell accessibility, distinguishing protected and exposed regions and defining EX1- and EX2-like regimes. These chimeras are responsive to proteolysis and pharmacological inhibition, enabling reversible control of protein activity. In a second application, we scanned 32 positions in yeast Ste2 and human A2A and MTNR1A receptors to engineer bi-functional chimeras that retain native function while incorporating reporter activity. Together, these results establish PAM scanning as a scalable, protein-agnostic framework for mapping insertion tolerance, interrogating protein accessibility in vivo, and enabling scalable ground-truth benchmarking of predictive chimera engineering.

|
Scooped by
mhryu@live.com
May 7, 11:42 PM
|
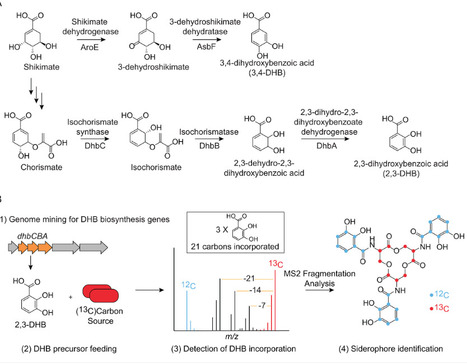
Bacteria produce high-affinity, iron-chelating secondary metabolites called siderophores to access insoluble Fe(III) in their environments. Genome mining has revealed many predicted siderophore biosynthetic gene clusters (BGCs) in bacterial genomes; however, the structures of their siderophore products remain mostly undetermined. This limits our molecular-level understanding of how bacteria acquire iron. Here, we apply inverse stable isotope labeling (InverSIL) to rapidly connect predicted siderophore BGCs to their products. With InverSIL, bacteria are grown on 13C-substituted carbon sources and then fed predicted biosynthetic precursors at their natural isotopic abundance to identify BGC products by mass spectrometry, removing issues with the availability of isotopically substituted precursors. We use InverSIL to determine the structures of the siderophore products of predicted BGCs from the methylotrophic genera Methylophilus and Methylorubrum, as well as the siderophores produced by the opportunistic pathogen Chromobacterium violaceum, which were previously shown to be essential for virulence yet remained structurally uncharacterized. We next use this approach to reveal the unexpected production of enterobactin by the genera Kushneria and Paracoccus, which was difficult to predict from genome sequences due to the distributed nature of the biosynthetic genes within the genomes. Finally, we use InverSIL to discover new siderophores, the cellulochelins, from the cellulose-degrading plant symbiont Cellulomonas sp. strain Leaf334. These findings demonstrate the utility of InverSIL for functional BGC characterization and expand our molecular understanding of bacterial iron acquisition strategies.

|
Scooped by
mhryu@live.com
May 7, 11:18 PM
|
Many microbiological outcomes are shaped by the determinants of community composition, including the factors that allow pathogens to invade healthy microbiota and the processes that maintain the diversity that underpins soil function. Community ecology provides a rich conceptual toolset to investigate patterns of coexistence in ecosystems that can be adapted to explain and manage these outcomes. However, these concepts have complex histories of controversy and debate that must be considered when applying them to the microbial context. Microorganisms also have distinctive characteristics that must be accounted for when applying ideas that were originally developed to describe macroscopic ecosystems. Here, we provide a concise overview for microbiologists to five key frameworks from community ecology: Niche theory, Trophic levels, Keystone species, Succession, and Metacommunities. We discuss the historical context and controversies surrounding each framework and outline existing and potential applications to microbial systems. This work therefore provides a practical guide for microbiologists who wish to apply community ecology for understanding and manipulating microbial community composition.

|
Scooped by
mhryu@live.com
May 7, 10:44 PM
|
Pollution of natural ecosystems is a global concern, with industrialization contaminating millions of soil and water sites. These contaminants threaten human health, agricultural productivity, and ecological balance. Traditional bioaugmentation strategies, while cost-effective and sustainable, face challenges including slow degradation rates and environmental constraints on microbial efficacy. This mini review examines phage bioaugmentation, which uses bacteriophages to enhance microbial pollutant degradation and support a circular economy. We focus on lysogenic phages that integrate auxiliary metabolic genes into bacterial hosts to improve degradation capacity, synthesise current knowledge, identify challenges, and propose a conceptual workflow for implementation. This mini review introduces phage bioaugmentation, a strategy to enhance soil bioremediation by using lysogenic bacteriophages to transfer pollutant-degrading genes to bacteria.

|
Scooped by
mhryu@live.com
May 7, 4:37 PM
|
Cells in many naturally occurring organisms routinely cooperate to control their extracellular pH in a dynamic and reversible manner, but this capability has been underexplored in synthetic biology. Here, we sought to engineer a microbial system that switches between two states —high and low extracellular pH— with minimal human intervention. We accomplished this by combining: (1) a genetic circuit that produces recombinant urease under the control of a light-inducible promoter; (2) a degradation tag on urease to accelerate the high-to-low pH transition; and (3) optimization of several environmental factors, including media composition, replenishment rate, and light exposure patterns. The system raises the pH when urease is produced and hydrolyzes urea in the media to produce ammonia; it lowers the pH as a byproduct of the cell's native metabolism when urease production ceases. We demonstrate that the optimized system cycles continuously for up to 14 days with minimal performance loss. Overall, our system demonstrates synthetic pH control in an engineered living system and highlights challenges and potential solutions for using such systems outside of the context of typical laboratory manipulation.
|

|
Scooped by
mhryu@live.com
Today, 3:00 PM
|
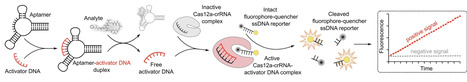
Analyte detection through aptamer-induced signal generation by CRISPR-Cas enzymes has rapidly emerged as a popular biosensing approach. Here, we investigated the implementability and analytical performance of this setup for the detection of diverse small molecule analytes. We selected nine aptamers from the literature targeting seven analytes and tested a commonly used assay design whereby analyte binding by the aptamer liberates a short complementary ssDNA strand, which in turn activates Cas12a to generate a fluorescence signal. After extensive optimization, the assay functioned for only two of the seven analytes, and several previously reported results could not be reproduced. While Cas12a fluorescence ssDNA detection was robust, the low success rate is likely due to aptamers not functioning reliably, underscoring the need for careful aptamer validation. Overall, our study provides a critical assessment of aptamer-Cas12a assay performances and discusses potential strengths, limitations, and pitfalls of this biosensing strategy.

|
Scooped by
mhryu@live.com
Today, 2:01 PM
|
Nisin, a natural, nontoxic antimicrobial peptide, has been widely studied for its broad-spectrum antibacterial activity against Gram-positive bacteria and its applications in food preservation and pharmaceuticals. However, the low nisin titer of Lactococcus lactis poses an obstacle to its large-scale industrial production. Here, we studied the metabolic basis of increased nisin production under aerobic conditions and proposed a high-efficiency fermentation strategy. Aerobic cultivation of L. lactis CF6 significantly increased intracellular ATP and the NAD+/NADH ratio, thus redirecting metabolic flux toward nisin production. Furthermore, cysteine availability and the supplementation of hemin and antioxidants effectively reduced oxidative stress caused by high dissolved oxygen, leading to a significant improvement in nisin titer. Through multimodule synergistic regulation and process optimization, a nisin titer of 18200 IU/mL was achieved in a 5 L bioreactor. In summary, this study lays the foundation for the metabolic engineering of L. lactis and provides valuable insights into the large-scale biomanufacturing of natural antimicrobial peptides.

|
Scooped by
mhryu@live.com
Today, 1:53 PM
|
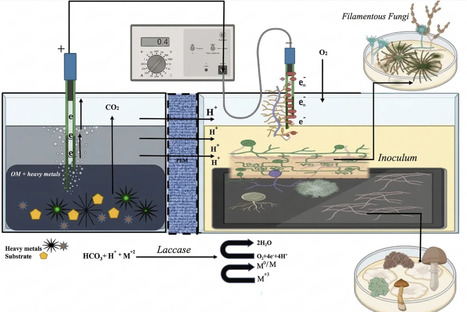
Heavy metal contamination of soil and water is an escalating global environmental issue driven by industrialization and poor waste management. This problem is no longer confined to urban areas, as even small towns are grappling with severe heavy metal contamination, posing substantial threats to human health and aquatic ecosystems. To address these issues, fungal fuel cells have become one of the most promising and environmental friendly technologies. This technology is at the forefront of efforts to combat heavy metal contamination. This method utilizes the unique properties of fungi at the biocathode to treat contaminants and remove heavy metals from soil and water. In addition to reducing pollution, this technology has the capacity to generate electrical current which can serve as an alternative to conventional remediation methods. This review aims to provide a general overview of the fungal fuel cells as a method of bioremediation to remove toxic heavy metals and while simultaneously generating electricity. This review is based on a critical analysis of the recent peer-reviewed publications focusing on the development, operation, and application of fungal fuel cells in heavy metal remediation and bioelectricity production. By exploring the potential of fungal fuel cells, the review provides insight into a future where heavy metal pollution is effectively curtailed while contributing to sustainable energy production, thereby fostering a cleaner and healthier environment.

|
Scooped by
mhryu@live.com
Today, 1:38 PM
|
Developing engineered genetic switches that can respond to specific stimuli in mammalian systems has great potential to advance next-generation therapies. This study introduces a novel tetracycline-inducible artificial riboswitch that regulates gene expression at the translational level via a modified −1 programmed ribosomal frameshifting (−1 PRF) platform. Our dose-response analysis reveals an EC50 of 0.41 μM, indicating exceptional sensitivity compared to other tetracycline-dependent artificial riboswitches developed for mammalian applications. By varying the precise spacing and composition of the tetracycline aptamer as the stimulatory structure on the −1 PRF platform, a series of switches has been constructed that enable robust induction of gene expression upon ligand addition. To attain a translational gene control platform with broader applicability, we developed a concept that allows it to be integrated upstream of a gene of interest (GOI), enabling precise control without requiring extensive re-engineering and expressing the GOI as a fusion protein. We validate the platform’s effectiveness in several genetic contexts, confirming its modularity. As optimization efforts continue to develop artificial riboswitches with optimized properties in mammalian systems, our findings contribute to the development of highly sensitive and efficient regulatory devices with beneficial characteristics, such as reversibility, modularity, and compatibility with RNA-based delivery therapies.

|
Scooped by
mhryu@live.com
Today, 1:30 PM
|
PockFlex is a web server designed to analyze pockets across protein structural ensembles and support the reconstruction, characterization, and prioritization of recurrent binding site organizations. Applicable to ensembles derived from molecular dynamics simulations, multiple experimental structures, or protein structure predictions, PockFlex detects pockets independently in each conformation, retains those overlapping a user-defined region of interest, and groups them across the ensemble by residue-level similarity. This residue-centred clustering framework identifies recurrent binding site clusters, quantifies residue recurrence and variability, and distinguishes persistent from transient binding site regions across the ensemble. Pocket-level druggability, predicted using the PockDrug workflow, is summarized at the cluster level to support binding site prioritisation under conformational variability while preserving access to individual pocket scores. The web application provides interactive, residue-level insights into pocket organisation, variability, and druggability in structural ensembles. The web server is free and open to all users, without login requirement, at https://pockflex.rpbs.univ-paris-diderot.fr/.

|
Scooped by
mhryu@live.com
Today, 9:25 AM
|
A non-fluent English speaker struggles to navigate language barriers in academic publishing. such programmes are rare and don’t always have capacity to meet the demand, notes evolutionary ecologist Andrew McAdam at the University of Colorado Boulder. He helped to create an online platform called Peer Edits that enables authors to solicit feedback on their manuscripts from early-career volunteers working in the same field. Another online resource, Rising Scholars, which provides support for scientists based in low- and middle-income countries, also offers free workshops and mentoring programmes for scientific writing. Laxman Gnawali, the president of the Nepal English Language Teachers’ Association in Kathmandu, adds that he’s found the online university Coursera’s course on academic writing helpful.

|
Scooped by
mhryu@live.com
Today, 1:15 AM
|
Hyperuricemia has emerged as the fourth most prevalent metabolic disorder, necessitating the development of safer and more effective therapeutic strategies. In this study, we constructed a recombinant probiotic strain expressing the PucL and PucM enzymes, which demonstrated a uric acid degradation rate of 65% in vitro. To enhance this activity, we performed modular optimization by employing three ribosome binding sites (RBSs) of different strengths—RBS 29, RBS 31, and RBS T7—to tune the expression levels of pucL and pucM, resulting in highly efficient uric acid degradation. Further improvement was achieved by overexpressing the uric acid transporter gene ygfU and the hydrogen peroxide–degrading catalase gene katG, leading to significant uric acid degradation. Furthermore, the engineered E. coli Nissle 1917 strain was evaluated in a mouse model of hyperuricemia; treatment with the optimized probiotic reduced serum uric acid levels to 39.11 mg/L, representing a 15.98% decrease compared with the control group. Further analysis revealed that this engineered bacterium ameliorates hyperuricemia by modulating the Firmicutes-to-Bacteroidetes ratio, increasing microbial diversity, and promoting the growth of beneficial genera. Collectively, this study establishes an engineered probiotic cell factory for uric acid degradation and demonstrates a proof-of-concept for the microbial remediation of hyperuricemia.

|
Scooped by
mhryu@live.com
Today, 12:52 AM
|
Because all known living organisms are made from at least 20 canonical amino acids, the feasibility of life using a more simplified alphabet remains unclear. In this work, we leveraged computational design and synthetic biology to explore building a cell from a 19–amino acid alphabet. Initial analyses suggested that isoleucine (Ile) may be dispensable, which we confirmed by directly replacing Ile residues in essential proteins in E. coli. Critically, protein language models and structure-based models were necessary to redesign functional Ile-less proteins in most cases. We systematically replaced all 382 Ile residues from the ribosome and combined 21 redesigned subunits at a native genomic locus to produce a viable, evolutionarily stable cell. This work provides a roadmap to create the first 19–amino acid organism since early evolution.

|
Scooped by
mhryu@live.com
Today, 12:00 AM
|
Bio-based materials are known for their excellent biodegradability and, in some cases, their potential to fix carbon dioxide. Owing to these properties, they are increasingly being utilized as environmentally friendly alternatives across various applications. In this study, we focused on using living cells themselves as material components, aiming to evaluate their potential as substitutes for conventional plastic-based thermal insulators. We selected two types of cells, photosynthetic purple non-sulfur bacterium Rhodovulum sulfidophilum and tobacco BY-2 plant suspension cells. After optimizing solidification conditions through the addition of pectin and cellulose nanofibers, we measured the thermal conductivity of the solidified cells under atmospheric pressure. The results showed that R. sulfidophilum exhibited 0.0553 W/m·K, while BY-2 exhibited a thermal conductivity of 0.043 W/m·K. Both values indicate relatively low thermal conductivity compared to existing bio-based materials, suggesting high insulation performance. Among the solidified cells, the solidified BY-2 cells showed minimal variation in thermal insulation performance under pressure changes, and had a low thermal emissivity as revealed by FT-IR analysis. Based on these findings, we propose that cell-derived materials can serve as potentially biodegradable bio-based thermal insulation materials.

|
Scooped by
mhryu@live.com
May 7, 11:30 PM
|
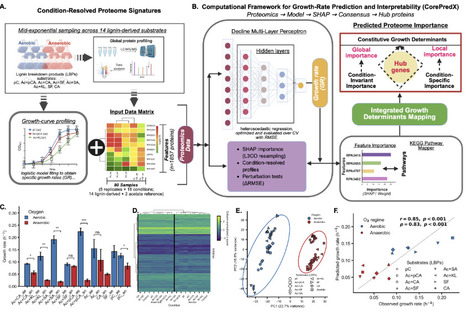
Lignin depolymerization generates mixtures of aromatic compounds that are promising carbon sources for microbial bioconversion, yet the constraints governing microbial growth on these substrates remain unclear. Here, we investigated how Rhodopseudomonas palustris CGA009 organizes metabolism to grow on chemically distinct lignin-derived aromatics under aerobic and anaerobic conditions. Quantitative proteomics across 14 substrate-oxygen environments revealed extensive oxygen-dependent proteome remodeling, consistent with shifts between respiratory and photoheterotrophic programs. Despite this reorganization, predictive modeling showed that growth rate is encoded by a relatively small subset of proteins whose abundance tracks physiological performance across environments. Using CorePredX, a machine-learning framework that combines global importance analysis with dependence-aware redundancy filtering, we identified 118 high-confidence proteins whose variation made conditionally non-redundant contributions to growth-rate prediction among 1,857 quantified proteins. These proteins are organized into a hierarchical architecture comprising a cross-condition core, adaptive regulators, substrate-specific specialists, and conditional hubs. The cross-condition core linked translational capacity, sulfur assimilation, redox buffering, and carbon storage cycling, highlighting conserved proteomic features associated with growth across oxygen regimes and substrate chemistries. Notably, an uncharacterized cystathionine beta-synthase domain protein, RPA3416, emerged as a strong predictive component of this core, raising the hypothesis of an adenylate-responsive process associated with aerobic growth on lignin-derived aromatics. Together, these results suggest that the metabolic versatility of R. palustris arises from flexible regulation layered onto partially conserved proteome-encoded growth-associated constraints, providing candidate targets for lignin bioconversion engineering and metabolic model refinement.

|
Scooped by
mhryu@live.com
May 7, 11:06 PM
|
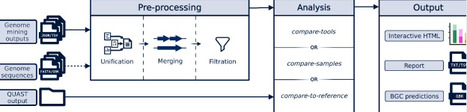
Biosynthetic gene clusters (BGCs) encode microbial natural products, many of which have important ecological and biomedical roles. Genome mining tools enable large-scale BGC prediction, but their outputs differ substantially, complicating comparison and interpretation. We present BGC-QUAST, a framework for evaluating and comparing BGC predictions across three analysis modes: comparison across samples, assessment of BGC recovery in draft assemblies relative to reference genomes, and comparison of predictions from different tools using overlap analysis. BGC-QUAST provides standardized metrics, interactive visualizations, and integrated outputs for joint inspection of predictions, enabling the comprehensive comparison of genome mining results and facilitating sample prioritization based on biosynthetic potential. https://github.com/gurevichlab/bgc-quast

|
Scooped by
mhryu@live.com
May 7, 10:40 PM
|
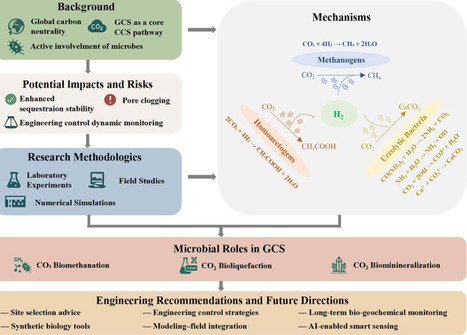
Geological carbon sequestration (GCS) is a key option for climate change mitigation, and subsurface microorganisms can alter CO2 behavior through biomethanation, bioliquefaction, and biomineralization. This review summarizes the major microbial processes involved in GCS and evaluates their effects on carbon stabilization, resource reutilization, and storage risk. We compare microbial distribution and metabolic functions across representative geological reservoirs and synthesize laboratory, numerical, and field approaches into a multi-scale framework for studying microbially mediated GCS. We also discuss engineering regulation strategies, site selection, monitoring, and risk control, together with current technical challenges and future research priorities. Overall, this review provides an integrated perspective on microbial mechanisms and practical guidance for safer and more effective GCS deployment.
|
 Your new post is loading...
Your new post is loading...