Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 4:54 PM
|
Land plants underpin civilization and planetary health, yet their genomic diversity remains largely uncharted. Current resources are unstandardized and scarce, lacking reference genomes for 95% of genera, 70% of families, and 51% of orders, impeding evolutionary and functional insight. We thus propose the PLANeT initiative, an international effort to generate high-quality, standardized genomes across the plant tree of life. Integrating artificial intelligence (AI) with genomics, we will decode conserved principles to advance fundamental plant biology, biodiversity conservation, crop improvement, and natural product discovery. Engaging around 100 labs to train 1,000 scientists, we will tackle pivotal questions for a sustainable future.

|
Scooped by
mhryu@live.com
Today, 4:46 PM
|
Integrons are genetic elements that drive bacterial adaptation by capturing and expressing mobile gene cassettes. They play a key role in dissemination of antimicrobial resistance (AMR) genes, particularly in Gram-negative bacteria. In addition to known AMR determinants, integron gene cassettes carry a vast reservoir of novel genes whose functions are largely uncharacterised, making it difficult to assess their full contribution to the resistome. Contributing to this are limitations in current sequence-based prediction methods which often lack the ability to identify unknown AMR or other adaptive genes with novel mechanisms. To address this, we developed a high-throughput gene cassette capture system that utilizes site-specific recombination activity of integrons and a counter selection strategy to capture and express gene cassettes from metagenomes. Coupling this platform with a functional screening approach allowed us to rapidly assay large libraries of environmental gene cassettes. Using this system, we recovered previously unknown AMR determinants while also providing insights into the prevalence of known clinical AMR genes in a range of environmental samples, including food and fertiliser. Here we provide experimental data on multiple novel bleomycin resistance genes and a stress response gene conferring gentamicin and tobramycin resistance. Our sequence analysis of the captured library also highlighted the diversity of the environmental cassette pool, with 656 unique cassettes recovered, the majority of which encoded proteins with unknown functions. The cassette capture system is a powerful tool for accessing hidden elements of the resistome and discovering novel adaptive genes that may go undetected using current sequence-based approaches.

|
Scooped by
mhryu@live.com
Today, 4:41 PM
|
Carboxysomes are self-assembling proteinaceous microcompartments that encapsulate ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco) and carbonic anhydrase within a semi-permeable shell, thereby elevating local CO2 concentrations around Rubisco to improve carbon fixation. Their inherent design principles, including programmable architecture, cargo encapsulation, semi-permeability, and modular assembly, position carboxysomes as powerful paradigms in synthetic biology and bioengineering, offering unprecedented opportunities to boost carbon assimilation and unlock novel biotechnological functions. This review summarizes current knowledge of the molecular mechanisms of carboxysome structure, assembly, and function, and highlights key recent breakthroughs and key challenges such as achieving precise control of shell permeability and efficient cargo encapsulation, integrating carboxysomes into heterologous hosts. We also outline emerging strategies and future perspectives for engineering carboxysomes as enhanced CO2-fixing engines and repurposing them as versatile nanomaterials in biotechnological applications. Together, these advances underscore the growing potential of carboxysome engineering to transform carbon-fixation pathways across diverse biological systems.

|
Scooped by
mhryu@live.com
Today, 4:32 PM
|
Plant-based materials provide a renewable and sustainable platform for tackling global challenges in energy, environment and healthcare. Among these, chronic diabetic wounds, currently affecting over 10% of the global population, remain a pressing public health crisis, largely owing to persistent hypoxia and unresolved inflammation. Here we report photosynthetic microspheres derived from the invasive aquatic plant Eichhornia crassipes (water hyacinth) as a green therapeutic strategy for chronic wound repair. Harnessing intact chloroplasts, these biogenic microspheres deliver sustained oxygen evolution for up to 3 weeks, alleviating local hypoxia and restoring redox balance. When integrated with the metabolic regulator metformin, the system couples exogenous photosynthesis and endogenous glucose metabolism, synergistically reprogramming the wound microenvironment. In both mouse and pig diabetic wound models, this approach suppresses inflammation, promotes angiogenesis and expedites re-epithelialization, leading to adaptive tissue regeneration. Our findings reveal that inexpensive, invasive aquatic biomass can be upcycled into advanced therapeutic biomaterials, offering a scalable and sustainable framework for next-generation regenerative medicine. This study showcases the upcycling of invasive water hyacinth into photosynthetic microspheres which provide sustained oxygen evolution and metabolic regulation, significantly accelerating the healing process of chronic diabetic wounds.

|
Scooped by
mhryu@live.com
Today, 4:16 PM
|
While nitrogen fertilizers are widely used in agricultural production, their application incurs significant environmental and energetic costs. In contrast, some crops are less dependent on these fertilizers because they engage in symbioses with rhizobia, nitrogen-fixing bacteria provide ammonium to the plant in exchange for carbon. However, the carbon cost associated with nitrogen fixation can negatively impact crop yields. Improving the efficiency of this metabolic process could alleviate this impact on crop productivity. Mathematical models can help us quantitatively explore metabolic behavior and identify potential targets for metabolic engineering. In this work, we developed a kinetic model of determinate root nodule metabolism, where this symbiotic exchange of carbon from the plant and nitrogen from the bacteria occurs. We used this model to evaluate how the predicted metabolic behavior differs between inefficient and efficient nodules, and to identify potential engineering targets for improving nitrogen fixation efficiency and rate. We show that the enzymes phosphoenolpyruvate carboxylase and pyruvate kinase have significant influence on the predicted rate and efficiency of nitrogen fixation, especially when their expression is varied in combination with oxidative Pentose Phosphate Pathway enzymes like glucose-6-phosphate dehydrogenase and 6-phosphogluconolactonase. The model predicts that pairing a 3-fold decrease in glucose-6-phosphate dehydrogenase activity along with either a 3-fold increase in phosphoenolpyruvate carboxylase activity or decrease in pyruvate kinase activity could increase nitrogen fixation rate by 5.51% while improving nitrogen fixation efficiency by 7.74%.

|
Scooped by
mhryu@live.com
Today, 4:10 PM
|
Many components of eukaryotic innate immunity originated from bacterial immune systems. However, it has been unclear how eukaryotes acquire these genes, why eukaryotes have sampled only certain families of bacterial proteins, and how these components become domesticated into eukaryotic physiology. Here, we discovered a recent instance of bacteria-eukaryote horizontal transfer and used it to characterize the genetic and biochemical changes that accompanied HGT. We focus on TIR domains, which are widespread yet potentially costly immune modules that are commonly associated with inflammation and/or cell death. By generating an atlas of TIR diversity across the tree of life, we phylogenetically categorized the domains and uncovered highly diverged TIR families found in eukaryotes. This analysis also allowed us to identify the TirBCD protein family of amoeba, which has been horizontally acquired and is closely related to the bacterial immune protein TIR-STING. Across their short eukaryotic history, the amoeba genes have acquired introns, evolved distinct patterns of gene expression, and engaged in evolutionary patterns of duplication and divergence typical of eukaryotic immune genes. While the TIR domain was transferred into amoebae, the genomic locus did not contain other components of a bacterial operon nor were regulatory domains transferred into the TIR protein. Nevertheless, TirC retains biochemical and physiological similarities to TIR-STING. TirC is a highly potent NADase, capable of spontaneously oligomerizing into large complexes and depleting cellular NAD even at very low protein concentrations. When expressed in yeast or E. coli, TirC is spontaneously active and highly toxic, illustrating the dangers of autoimmunity following TIR protein movement into novel hosts. In contrast, amoebae tolerated high TirC expression with no disruption in cell size, growth, or behavior. Single, double, and triple knock out mutants of amoeba tirBCD are viable and display modest defects in their ability to phagocytose bacteria, implying that the co-opted bacterial TIR domain may regulate eukaryotic host-microbe interactions. Overall, this study uncovers an informative example of recent eukaryotic TIR evolution that captures features of both bacterial and eukaryotic immunity. In addition, we expect that the TIR domain atlas will be useful to researchers in many model systems as they explore the vast diversity of TIR molecular and cellular functions.

|
Scooped by
mhryu@live.com
Today, 4:04 PM
|
The exploration of colocalized gene sets, such as Biosynthetic Gene Clusters (BGC) and symbiotic islands, is fundamental in modern genome mining. However, many existing prediction tools rely heavily on curated databases or predefined rules, inherently biasing detection toward known clusters. To address these limitations, we introduce Scan Cluster, a robust and flexible Python-based bioinformatic tool designed to identify user-defined, conserved gene clusters across diverse genomes without database-driven constraints. Scan Cluster leverages BLAST or HMMER to detect homologous proteins and evaluates their colocalization, accommodating complex evolutionary events such as orientation inversions, the insertion of alien genes, gene deletions, and the integration of insertion sequences. Beyond identification, the software performs progressive multiple cluster alignments and generates distance trees to assess cluster similarity, producing outputs ready for visualization in iTOL and Clinker. We validated Scan Cluster against standard tools like antiSMASH and DeepBGC, demonstrating high accuracy in delineating complete cluster boundaries. Its utility was further confirmed through the analysis of the nos and complex nod-nif-fix symbiotic gene clusters in rhizobia strains, successfully tracking genetic decay and grouping diverse architectures. Scan Cluster provides an accessible, low-resource framework to explore the evolutionary and functional dynamics of novel genetic clusters.

|
Scooped by
mhryu@live.com
Today, 3:55 PM
|
Here, we introduce the potential of plasma technology for future lunar agriculture. The use of discharge plasma to convert nitrogen, oxygen, and water into reactive species for nitrogen fixation is discussed. This study demonstrated the effectiveness of dinitrogen pentoxide as a nitrogen fertilizer for rice cultivation in lunar regolith simulants. This technology also shows promise in regulating plant growth and enhancing plant immunity, addressing the challenges of lunar agriculture.

|
Scooped by
mhryu@live.com
Today, 3:49 PM
|
GDP-Mannose transporters are Golgi-localized solute carriers that are essential for the virulence of pathogenic fungi, serving as critical components of fungal glycosylation pathways. However, the mechanism by which nucleotide sugars are recognised and transported across the Golgi membrane remains unclear, hindering efforts to develop effective inhibitors that could serve as distinct antifungal agents. Here, we present cryo-EM structures of the GDP-Mannose transporter, Vrg4, from Candida albicans in complex with nanobodies in both the cytoplasmic and Golgi-facing states. Structural comparisons between these two states, in addition to a GDP-mannose bound structure, demonstrate the importance of ligand movement during transport. Additionally, we demonstrate the ability of the nanobodies to specifically inhibit Vrg4, presenting proof-of-principle that nanobodies can be used as effective inhibitors of nucleotide sugar transport and glycosylation in cells. Invasive fungal pathogens have become a significant concern in healthcare systems worldwide. Here, authors demonstrate that nanobodies targeting the GDPmannose transporter can arrest fungal cell growth. The structures also reveal the complete transport cycle for an SLC35 nucleotide sugar transporter.

|
Scooped by
mhryu@live.com
Today, 3:43 PM
|
Sequence-based protein-protein interaction (PPI) site predictors typically analyze proteins in isolation, neglecting partner-specific context that is critical for interface specificity, particularly in transient and disordered interactions. We introduce SPPIDER-seq, a partner-aware PPI site prediction framework that combines pretrained ESM-2 embeddings with a cross-attention architecture to enable residue-level conditioning on interacting partners. We curated non-redundant protein-peptide interaction datasets from BioLiP and used them to train and benchmark two complementary models: a receptor-centric model optimized for structured interfaces and a peptide-centric model tailored to disordered, motif-driven binding. On blind benchmarks, SPPIDER-seq achieved AUROC values up to 0.797 and MCC values up to 0.269, outperforming AlphaFold3 on peptide-mediated and disordered interfaces while remaining complementary on globular complexes. Application to 341 TP53 interaction partners revealed coherent, partner-specific interface patterns across both structured and intrinsically disordered regions. Availability and Implementation: SPPIDER-seq models, datasets, and the Python code are freely available on the web at: https://github.com/aporollo-lab/SPPIDER-seq

|
Scooped by
mhryu@live.com
May 1, 5:04 PM
|
Microbial cell factories are vital for the sustainable production of chemicals, fuels, and pharmaceuticals, yet their performance depends on the precise control of gene expression to balance metabolic flux between cell growth and production. While the regulation of metabolic networks through constitutive gene expression can promote high-level production, it also imposes a metabolic burden that adversely affects cellular growth. Here, we developed a versatile and high-efficiency inducible platform for the methylotrophic yeast Pichia pastoris, which enables dynamic gene expression for 3-hydroxypropionic acid (3-HP) production from sole methanol. We first constructed an anhydrotetracycline (aTc) inducible promoter library of diverse strengths modified from the constitutive promoter PGCW14. The promoter library consisted of 32 inducible promoters with the activated strengths ranging from 2.5% to 84.7% of PGCW14. Second, we dynamically regulated key genes in the 3-HP biosynthesis pathway by aTc inducible promoters, including malonyl-CoA reductase gene (MCR) from Chloroflexus aurantiacus with strong promoters RM5 and RM6, endogenous malonyl-CoA carboxylase gene ACC1 with the very strong promoter RM1, and transmembrane permease PpMFS with the medium promoter RM12. The final engineered strain SHP18 produced 3.7 g/L 3-HP in a shake flask by using methanol as the sole carbon source. This modular aTc inducible promoter toolkit provided finely graded transcriptional control in P. pastoris, which offers a broadly applicable strategy for cell factory design.

|
Scooped by
mhryu@live.com
May 1, 4:13 PM
|
Water environments serve as critical sources and conduits for antimicrobial resistance (AMR). The complex water cycle facilitates the transmission of antibiotics, resistant bacteria and resistance genes that have been released through anthropogenic activities, thereby reshaping AMR dynamics across different environmental compartments. Among these resistance determinants, inactivating antibiotic resistance genes (inactivating ARGs) encode enzymes that degrade or modify antibiotics to reduce their bioactivity. They play a complex and dual role bridging environmental reservoirs and clinical risk. While they can threaten the therapeutic effectiveness of antibiotics when transferred horizontally to pathogens, they also act as cooperative traits that lower local concentrations of bioactive antibiotics and relax selection for resistance. In turn, reduced antibiotic exposure can enhance ecosystem functions and stability in both host-associated and environmental microbiomes through ecosystem resilience and community protection. Inactivating ARGs represent a crucial intersection between AMR challenges and microbial community stability. Here we discuss the risks and ecological benefits of inactivating ARGs, and present intervention strategies aiming to both protect ecological resilience and limit resistance development in pathogens. Water environments are pivotal in the spread of antimicrobial resistance, acting as conduits for antibiotics and resistance genes. Here, the authors explore the dual role of inactivating antibiotic resistance genes, highlighting their ecological benefits and risks, and propose strategies to enhance ecosystem resilience while curbing resistance in pathogens.

|
Scooped by
mhryu@live.com
May 1, 2:04 PM
|
Streptomyces and insects engage in complex interactions shaped by millions of years of evolution. While many beneficial relationships are well recognized, it remains unknown whether Streptomyces produce virulence factors targeting insects specifically. Here, through bioinformatic analysis, we identified diphtheria toxin (DT) homologues, which we named Streptomyces antiquus insecticidal proteins (SAIP), within a monophyletic lineage of Streptomyces that emerged more than 100 million years ago. SAIP is cytotoxic to insect cells and lethal to Drosophila melanogaster, suppressing neuronal activity and immune responses in vivo. Structural and functional studies validated that SAIP is homologous to DT and acts by ADP ribosylation of eukaryotic elongation factor 2. CRISPR–Cas9 screening identified the insect protein Flower as the SAIP receptor across a range of insects. Toxigenic Streptomyces can consume dead insects and produce bioactive secondary metabolites while growing on insect carcasses. These findings establish an insecticidal toxin in Streptomyces and demonstrate that Streptomyces have evolved highly specific virulence factors against insects. Bioinformatic, structural and functional analyses characterize Streptomyces antiquus insecticidal protein (SAIP) as a diphtheria toxin homologue that is lethal to insect cells and targets the Flower protein as a receptor
|

|
Scooped by
mhryu@live.com
Today, 4:53 PM
|
‘Structurally known’ and ‘Structurally unknown’ have long served as a binary classification of natural products. However, with the emergence of synthetic biology, the boundaries of this simple classification are being redrawn, because SynBio expands both what we can see and what we can build. To reflect this change, we propose a SynBio-centered framework that maps antimicrobial natural products along two axes, chemical novelty and engineering accessibility, yielding four quadrants: (i) known knowns, well-characterized natural products accessed mainly through pathway refactoring and producer strain optimization; (ii) unknown knowns, engineerable scaffolds with established diversification routes but uncertain chemical and functional outcomes; (iii) known unknowns, architectures with precedent but limited realization due to missing transferable engineering routes, often because the relevant coupling logic has not yet been converted into a practical engineering handle; (iv) unknown unknowns, unanticipated scaffolds that need to be recognized through minimal chemical, genetic, or mechanistic anchors before engineering. Since these quadrants are dynamic rather than fixed, compounds may shift across the framework as discovery, route-building, and outcome predictability improve. This two-axis map highlights complementary SynBio strategies, from optimizing existing antibiotics to accessing first-in-class entities, which may help guide efforts to combat current resistance challenges.

|
Scooped by
mhryu@live.com
Today, 4:43 PM
|
The systematic exploration of novel bioactive compounds with superior functional properties is critical for driving innovations in agriculture, healthcare, and related fields, thereby becoming essential for advancing sustainable biotechnological solutions. Nonprotein amino acids (NPAAs), functional amino acids not incorporated into proteins, exhibit unique physiological activities and provide distinctive advantages in nutritional enhancement, functional product formulation, and food/feed processing. These attributes challenge the conventional perception of proteins as mere nutritional carriers, positioning NPAAs as promising bioproducts for biosynthesis and functional applications in agriculture, food, and medicine. This review summarizes the classification of the available NPAAs based on their synthetic substrates for the first time and then outlines their diverse functional roles. A comprehensive analysis of recent advances in biosynthetic pathways, engineering strategies, and production level demonstrates their primary research progress in the laboratory phase. The further sustainable biomanufacturing of NPAAs is hampered by several challenges, including poorly elucidated biosynthetic mechanisms, limited robustness and low productivity of microbial strains, and difficulties in scaling up production for industrial applications. Addressing these bottlenecks will require innovative strategies and technologies to facilitate the translation of NPAA production from bench to industry. This review offers valuable insights into the potential of NPAAs in the development of next-generation bioproducts of nutrition, immune regulation, antioxidant defense, and intestinal homeostasis maintenance, suggesting a promising direction for microbial production of high-performance bioactive molecules in agricultural synthetic biomanufacturing.

|
Scooped by
mhryu@live.com
Today, 4:33 PM
|
CRISPR–Cas effectors typically rely on RNA guides to recognize target sequences. In Cas12a, the protospacer adjacent motif on DNA engages conserved protein residues, triggering target binding and nuclease activation. Here we reprogram Cas12a into a DNA-guided, RNA-targeting effector. Exploiting protospacer-adjacent motif-dependent interaction, we engineer synthetic CRISPR DNA that engages Cas12a to form a functional deoxyribonucleoprotein complex, while repurposing solely RNA as the programmable target. Structural, biophysical and biochemical analyses reveal the molecular basis of this DNA-guided, RNA-targeting configuration and support an activation pathway distinct from that of canonical RNA-guided systems. DNA-guided Cas12a enables direct RNA detection and efficient intracellular RNA knockdown, establishing a modular activation architecture for CRISPR–Cas12a and expanding the design space for programmable RNA manipulation. Synthetic DNA guides (crDNA) reprogram Cas12a nucleases for RNA targeting.

|
Scooped by
mhryu@live.com
Today, 4:18 PM
|
Metabolic interactions are central to the functioning of microbial communities. In spatially structured environments, such interactions typically require close physical proximity between partner cells. However, cell division drives spatial segregation of interaction partners, as the cells emerging from division remain adjacent and form clonal clusters, weakening these interactions. Here, we hypothesized that active cell motility can reduce this spatial segregation by enabling cells to leave clonal clusters and thereby enhance metabolic interactions. We tested this hypothesis by performing time-lapse single-cell imaging of a synthetic cross-feeding consortium growing in microfluidic chambers. We found that surface motility disrupted clonal clustering, enhanced spatial intermixing, and consequently increased the growth rates of individual cells and community productivity. Individual-based simulations further revealed that this effect is robust across motility modes and a wide range of ecological and physiological parameters. Together, our findings demonstrate that even non-directed, random motility, without chemotactic sensing, is sufficient to enhance metabolic interactions by separating cells from their clonal lineages and repositioning them in proximity to their metabolic partners, thereby acting as a key driver of community functioning in spatially structured microbial systems.

|
Scooped by
mhryu@live.com
Today, 4:13 PM
|
Accurate RNA secondary-structure prediction remains difficult despite decades of thermodynamics-based algorithms and the advent of deep-learning architectures (convolutional networks, Transformers, diffusion models). In fact, the datasets that pair RNA sequences with secondary-structure labels are often low-quality, noisy, and family-imbalanced, which limits out-of-distribution generalization and exacerbates catastrophic forgetting when new data regimes are introduced. We propose a continual-learning approach based on Lifelong Bayesian Optimization (LBO), RNAFoLBO, that treats each class of RNAs obtained from latent-space clustering as a sequential task and jointly orchestrates training and hyperparameter selection of heterogeneous models (UFold, RNA-FM, RNADiffFold), while preserving prior knowledge. Concretely, we apply LBO to 15 clusters obtained by clustering RNAStrAlign in the latent space of RNAGenesis, a model specialized in contextual representation learning and latent-space structuring, achieving a mean F1 per cluster of 0.931 (with a range of 0.177). These results surpass the strongest one-shot baseline and mitigate forgetting without full retraining. The gains persist as additional clusters are introduced. Overall, RNAFoLBO delivers higher and more stable performance and practical scalability for integrating new RNA clusters or families, enabling more robust and transferable RNA secondary-structure prediction.

|
Scooped by
mhryu@live.com
Today, 4:07 PM
|
Molecular phylogenetics based on primary sequence comparisons has been central to reconstructing protein evolution. However, structural evolution does not necessarily parallel sequence divergence, particularly in proteins combining ordered domains with intrinsically disordered regions (IDRs). Here, we introduce a quantitative secondary structure distance (S2D) metric that enables systematic comparison of protein secondary structure, including both ordered elements and IDRs. Using the MADS-box transcription factor family as a model, we show that structural divergence is domain-specific and only partially coupled to sequence-based phylogeny. Domain-resolved analyses reveal that the DNA-binding M domain remains structurally constrained, whereas the I and C domains exhibit extensive sequence divergence while retaining conserved intrinsic disorder. In contrast, the K domain contributes disproportionately to global structural variability. Integrating S2D with phylogenetic distance uncovers both convergent structural architectures among distantly related proteins and pronounced structural remodelling within closely related paralogs patterns not evident from primary sequence comparisons alone. Residue-level analyses further demonstrate that the structural impact of mutation depends strongly on amino acid identity and does not scale directly with substitution frequency or conservation metrics. Together, these findings indicate that secondary structural evolution provides an additional dimension of protein diversification beyond sequence divergence. By integrating phylogenetic and structural distances, this framework offers a complementary approach to interpreting protein evolution, particularly in families containing mixtures of ordered domains and intrinsically disordered regions.

|
Scooped by
mhryu@live.com
Today, 4:01 PM
|
Cancer immunotherapies are a promising method for treating cancers and directly enhance the antitumor capabilities of a patient's immune system. In particular, therapeutic mRNA cancer vaccines are an attractive strategy as they offer a specificity and safety level that other immunotherapies are unable to achieve. Incorporation of tumor-specific or tumor-associated antigens in mRNA vaccines aims to enhance tumor antigen-specific T cells. mRNA vaccine adjuvants often encode immune stimulating proteins such as proinflammatory cytokines, costimulatory molecules, and pattern recognition receptors or their corresponding agonists. This review summarizes prominent clinical and preclinical therapies utilizing mRNA tumor-associated or tumor-specific antigens and mRNA encoding immunostimulants delivered either as vaccine adjuvants or as monotherapies for the treatment of cancer. The limitations, challenges, and future directions of therapeutic mRNA vaccines are also discussed.

|
Scooped by
mhryu@live.com
Today, 3:52 PM
|
Cytoplasmic abundant heat-soluble (CAHS) proteins, a stress-responsive intrinsically disordered protein from tardigrades, have been discovered to form gel-like networks providing structural support during dehydration, thus enabling anhydrobiosis. However, the mechanism by which CAHS proteins protect the dehydrating cellular membrane remains enigmatic. Using giant unilamellar vesicles (GUVs) as a model membrane system, here we show that encapsulated CAHS12 undergoes a reversible structural transformation that reinforces membrane integrity and preserves encapsulated components, mimicking natural anhydrobiosis. CAHS12-containing GUVs demonstrated stability for weeks and mechanical robustness under dehydration, elevated temperature, and osmotic stresses. Molecular simulations suggest that CAHS12 forms a filamentous network within the vesicle lumen that mitigates membrane collapse and preserves compartmental architecture. Synthetic cells with cell-free transcription-translation capabilities withstand desiccation and recover biochemical activities, akin to the tun state of the tardigrade. This discovery opens up synthetic cell applications in bioengineering, cold-chain-independent biomanufacturing, and adaptive biointerfaces. Synthetic cells with tardigrade protein CAHS12 mimic desiccation survival by stabilizing membranes. This study uncovers how CAHS12 preserves cell structure and suggests new applications in biomanufacturing and adaptive biointerfaces.

|
Scooped by
mhryu@live.com
Today, 3:46 PM
|
Advances in artificial intelligence (AI) and synthetic biology are transforming biological research and biotechnology. These fields are for the first time enabling the design and development of human and bacterial cells that can serve as “living” drug delivery vehicles that perform sustained release of therapeutic cargo with spatiotemporal control. In recent years, human and bacterial cells have been engineered to deliver peptides, proteins, and biologics for treating a wide range of human diseases using synthetic biology approaches. To engineer effective living drug delivery systems, detailed knowledge is required about how to design receptors that can specifically sense the tissues targeted for drug delivery, signaling networks that can process signals from these receptors, and gene circuits that can control therapeutic cargo production and release. However, elucidating such receptors, signal processing and gene regulatory elements, and gene circuit compositions by traditional design-build-test-learn approaches is difficult and low throughput. Here we review how advances in AI and synthetic biology are meeting these challenges. We describe examples of how human cells and bacteria are engineered to become living drug delivery vehicles. We discuss how AI and synthetic biology approaches are being applied to discover the sequence-to-function design principles for engineering synthetic receptors, signaling proteins, and gene regulatory elements and the composition-to-function design principles for engineering synthetic gene circuits. We share an outlook on opportunities for AI and synthetic biology to synergize for creating next-generation living drug delivery systems.

|
Scooped by
mhryu@live.com
Today, 3:26 PM
|
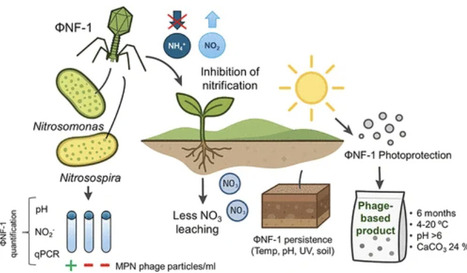
The intensive use of nitrogen fertilizers sustains crop productivity but causes substantial nitrogen losses, greenhouse gas emissions, and water contamination through nitrification-driven leaching and volatilization. Although chemical nitrification inhibitors reduce these effects, concerns over persistence, toxicity, and impacts on nontarget organisms limit their sustainability. As a biological alternative, bacteriophages can selectively suppress ammonia-oxidizing bacteria (AOB) responsible for nitrification. This study examines the virulent bacteriophage ΦNF-1, the first known phage infecting soil AOB of the genus Nitrosomonas. ΦNF-1 reduced nitrification in fertilized soils inoculated with Nitrosomonas europaea and, to a lesser extent, in fertilized soils containing native AOB communities. Beyond its previously reported hosts, ΦNF-1 also propagated in Nitrosomonas oligotropha and Nitrosospira multiformis, expanding its known host range. ΦNF-1 remained infectious for over six months in aqueous suspensions at 4–20 °C and neutral to alkaline pH, but declined rapidly under acidic conditions. In agricultural soils, infectivity persisted for up to six months, though recovery was lower in sandy soils. A calcium carbonate-based photoprotectant significantly enhanced phage survival under UV and sunlight exposure. Overall, ΦNF-1 shows strong potential as a stable, host-specific biological nitrification inhibitor to improve nitrogen use efficiency and reduce environmental impacts of fertilization.

|
Scooped by
mhryu@live.com
May 1, 4:23 PM
|
Directed evolution methods face trade-offs between the control of discrete approaches and the throughput of modern continuous systems. Here, we engineered a method called lytic selection and evolution (LySE) for near-continuous evolution of bacterial gene clusters while maintaining discrete checkpoints. We developed a hypermutagenic T7 DNA polymerase variant fused to a dual adenine-cytosine deaminase to install all possible transition mutations at similar frequencies. By relieving pressure from maintaining genome fidelity, we obtained mutation rates of 3.82 × 10−5 substitutions per base. For biocontainment, the T7 DNA polymerase was encoded on an accessory plasmid, while the target gene cluster was encoded on a T7 DNA polymerase-lacking T7 phagemid. Alternating cycles of lysis and transduction enable selective replication and mutagenesis of target genes, while off-target genomic mutations are discarded. LySE evolved a 25-fold increase in tetA-encoded tigecycline resistance in 5 cycles, and a 50.9% increase in endpoint biomass of a bacterial strain that uses the polyethylene terephthalate monomer, ethylene glycol, as its sole carbon source. Our method balances speed and control for directed bacterial evolution. The lytic selection and evolution (LySE) system uses a plasmid-encoded, hypermutagenic T7 DNA polymerase and T7 phagemid-encoded target genes to enable controllable, near-continuous directed evolution of large gene clusters.

|
Scooped by
mhryu@live.com
May 1, 4:11 PM
|
Phenotypic heterogeneity, a feature of both bacteria and eukaryotic cells, arises from inherent cell-to-cell variability. In eukaryotes, single-cell RNA sequencing has led to an explosion in understanding how heterogeneity impacts different cell types and states in organs and tissues. While single-cell RNA sequencing analyses in bacteria have lagged behind eukaryotic studies, recent technological advances now enable similar, high-resolution studies to be performed at scale in bacteria, yielding fundamental insights into how heterogeneity influences bacterial physiology, metabolism, antibiotic resistance, pathogenesis and interactions within complex microbial communities. Here we review recent advances in bacterial single-cell RNA sequencing, including the methods developed so far and what has been learned from their application. We also discuss technological and computational challenges going forwards, the need for standardization and how that could be achieved, and how this emerging field is now poised to revolutionize our understanding of bacterial physiology, infection biology and interactions within bacterial communities, such as the microbiota. This Review summarizes recent advances in bacterial single-cell transcriptomics approaches, while also discussing technological and computational challenges remaining to be met and the need for standardization going forwards.
|
 Your new post is loading...
Your new post is loading...