Your new post is loading...

|
Scooped by
mhryu@live.com
April 28, 11:12 PM
|
Rare sugars are monosaccharides and their derivatives that occur only in trace amounts in nature. Owing to their low caloric value and diverse physiological activities, they have attracted increasing attention as functional ingredients in the food, pharmaceutical, and health-related industries. However, their limited natural abundance and the inefficiency of traditional production methods remain significant challenges for large-scale application. Biosynthesis has emerged as a promising platform for rare sugar production due to its mild reaction conditions, environmental compatibility, and high catalytic specificity. With advances in synthetic biology and metabolic engineering, multiple biosynthetic routes have been developed. Further improvements in productivity and industrial feasibility increasingly depend on the coordinated integration of multi-level engineering strategies. In this review, we first briefly introduce the physiological functions and applications of representative rare sugars, including D-allulose, D-tagatose, D-allose, L-ribose, L-ribulose, and L-xylulose. We then provide a systematic overview of recent advances in biosynthesis, with a focus on metabolic engineering, enzyme engineering, biocatalysis, and fermentation process optimization. Finally, we discuss the key challenges and future directions for developing efficient microbial cell factories for rare sugar production.

|
Scooped by
mhryu@live.com
April 28, 10:40 PM
|
Accurate enzyme function annotation remains a major bottleneck in genome analysis despite the rapid expansion of available protein sequence and structure data. Most existing methods rely on sequence similarity or machine-learning representations, which often perform poorly for proteins with low sequence identity or convergent evolutionary histories. Because enzymatic activity is determined by the three-dimensional arrangement of catalytic and binding-site residues, structure-based approaches offer a mechanistically grounded alternative. However, their broader application has been constrained by the limited size and coverage of curated active-site reference databases. To address this challenge, we developed ActSeekN, a structural-motif-based functional annotation pipeline that combines the ActSeek active-site search algorithm with a newly constructed large-scale reference database derived from AlphaFold-predicted structures, UniProt annotations, and curated catalytic residue information. This framework enables rapid and scalable identification of conserved catalytic motifs across structurally related proteins, allowing function to be transferred on the basis of local three-dimensional catalytic geometry rather than global sequence similarity. In this way, ActSeekN overcomes a central limitation of previous structure-based methods by expanding the searchable space of catalytic motifs while retaining mechanistic interpretability. Benchmarking against state-of-the-art machine-learning approaches demonstrates competitive or superior performance. Applications to yeast, human, and Trichoderma reesei proteomes refine existing annotations, complete partial EC assignments, and identify previously unrecognized enzymatic functions, highlighting ActSeekN as a powerful tool for genome annotation and biotechnology.

|
Scooped by
mhryu@live.com
April 28, 9:38 PM
|
Synthetic genomics is advancing from microbial toward multicellular organisms. However, current manual methods for DNA and genome assembly remain inadequate for the efficient, large-scale production of long DNA constructs. Here, we present Programmed DNA Assembly via Cas9 and Conjugative Transfer (PACT), a method that integrates a linear vector system, bacterial conjugation, and programmable Cas9-mediated cleavage to achieve highly efficient, iterative assembly of large DNA fragments. PACT enhances assembly efficiency by ~30-fold compared to conventional circular vector strategies, enabling one-step assembly of DNA up to 80 kb. We engineered four single guide-RNA-Marker donor cassettes to support iterative assembly workflows. PACT can utilize low-recombination E. coli strains as hosts to efficiently assemble Arabidopsis thaliana mitochondrial genome with high repeat units. Integrated with an automated robotic platform, we developed an unattended, high-throughput pipeline (aPACT) toward large-scale parallel DNA assembly. Using aPACT, we successfully assembled three large DNA constructs: a 210 kb digital DNA, the designed chloroplast (120 kb) and mitochondrial (350 kb) genome of A. thaliana. This automated system offers a powerful tool for scalable assembly of large DNA molecules, accelerating synthetic genomics research toward complex multicellular organisms.

|
Scooped by
mhryu@live.com
April 28, 9:14 PM
|
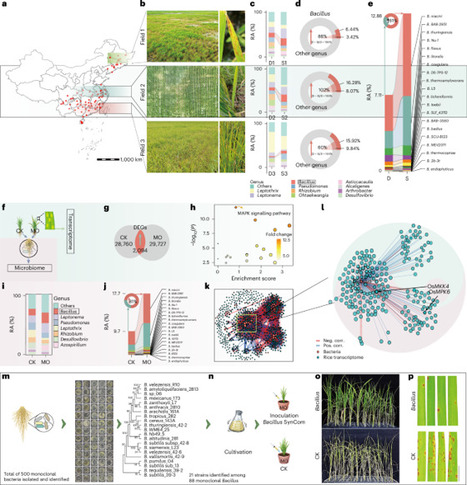
Plants actively recruit beneficial rhizosphere microorganisms to enhance immunity, though the molecular mechanisms are largely unclear. Here we identified a complete OsMAPK–OsWRKY pathway of rice rhizosphere Bacillus recruitment via root secretion of heptadecanoic acid upon rice blast occurrence. In field trials, enrichment of Bacillus in their rhizospheres was associated with higher disease resistance of rice. Integrated omics analyses demonstrated a strong correlation between the activation of the OsMKK4–OsMPK6 cascade and Bacillus recruitment. Loss-of-function mutants of OsMKK4 and OsMPK6 reduce rhizosphere Bacillus abundance by 82% and 79%, respectively. As the substrates of OsMPK6, OsWRKY24/53/70 transcription factors activate OsKCS2, which encodes a β‑ketoacyl-CoA synthase involved in heptadecanoic acid biosynthesis. Mutations of OsWRKY24/53/70 or OsKCS2 reduce root-secreted heptadecanoic acid by 15–28% and impair Bacillus colonization. Heptadecanoic acid application recapitulated Bacillus recruitment and disease resistance. Collectively, our results demonstrate that the MAPK pathway regulates heptadecanoic acid synthesis, which controls Bacillus recruitment as well as rice blast disease resistance. This study reveals that rice roots secrete heptadecanoic acid to recruit beneficial Bacillus for defence against blast fungus, a process regulated by a MAPK pathway.

|
Scooped by
mhryu@live.com
April 28, 9:08 PM
|
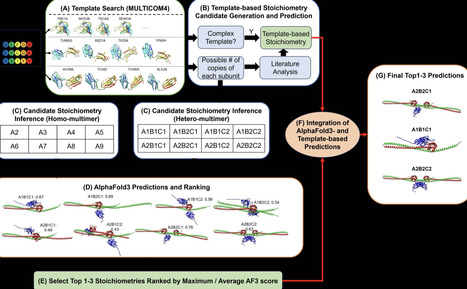
Protein complexes carry out many essential functions in cells, but predicting their structures requires knowing their stoichiometry, meaning how many copies of each subunit are present. This information is often unavailable, making stoichiometry prediction crucial for complexes with unknown stoichiometry. Despite its importance, few computational methods address this challenge. Here we show that combining AlphaFold3 structure prediction with information from related known protein complexes enables accurate prediction of protein complex stoichiometry. Our method generates plausible subunit combinations, builds structural models for them using AlphaFold3, ranks them using AlphaFold3 scores, and further refines predictions with template-based information when available. In the 16th community-wide Critical Assessment of Techniques for Protein Structure Prediction, our method identifies the correct stoichiometry as the top prediction for 71.4% of targets and among the top three predictions for 92.9% of targets, outperforming other methods overall. This demonstrates the complementary strengths of AlphaFold3- and template-based predictions and highlights the applicability of our approach to uncharacterized protein complexes lacking stoichiometry data. Integration of AlphaFold3 and homologous template information enables accurate prediction of protein complex stoichiometry for uncharacterized multimers in the CASP16 experiment.

|
Scooped by
mhryu@live.com
April 28, 8:44 PM
|
Salidroside exhibits a diverse range of biological activities and is extensively utilized in the fields of biomedicine and cosmetics. Given the rising global demand, there is a critical need to develop more sustainable production methods for salidroside. This study implemented metabolic engineering strategies to enhance the biosynthesis of salidroside. The aromatic aldehyde synthase (AAS) from Petroselinum crispum was introduced to synthesize tyrosol from endogenous l-tyrosine. Competing metabolic pathways were suppressed, and endogenous alcohol dehydrogenase enzymes were screened. Additionally, a quorum-sensing-based dynamic regulation system was used to optimize salidroside production. The enzymes phosphoglucomutase (pgcA) and uridine diphosphate glucose pyrophosphorylase (gtaB) were overexpressed to balance the precursor supply. Finally, the engineered strain BS18 produced salidroside at 661.4 mg/L. This was achieved in shake-flask fermentation using potato starch as the carbon source. The study shows the potential for the large-scale production of salidroside in Bacillus subtilis. It provides a foundation for industrial applications and advances synthetic biology in natural product biosynthesis.

|
Scooped by
mhryu@live.com
April 28, 7:46 PM
|
l-Tryptophan, an essential amino acid, serves as a critical precursor for the biosynthesis of numerous bioactive compounds. In this study, we engineered tryptophan synthase (TrpS) to develop an efficient cell factory for l-tryptophan production. Initially, EcTrpS from E. coli K-12 was expressed in E. coli BL21 (DE3) to construct strain BS, which produced 836.43 mg/L of l-tryptophan. To alleviate substrate inhibition, we constructed a mutant library of 104–105 variants through two rounds of error-prone PCR and utilized a biosensor employing a green fluorescent protein to report l-serine consumption for screening TrpS mutants. Mutant C10 showed a 375% increase in enzyme activity compared with strain BS. Computational analysis was used to explore an improved catalytic mechanism. Finally, through systematic process optimization and catalytic strategy, strain C10 produced 94.2 g/L of l-tryptophan, the highest titer via whole-cell transformation. This study provides new perspectives and strategies for TrpS engineering and the industrial production of l-tryptophan.

|
Scooped by
mhryu@live.com
April 28, 11:33 AM
|
As for sustainable food security, plant genetic engineering has emerged as a transformative technology offering innovative solutions. This review comprehensively examines recent advances in plant genetic engineering, from technical foundations to technological innovations, and to multifaceted applications. They transcend the constraints of traditional breeding, including its long cycle and narrow genetic base, showing remarkable potential in crop improvement. By modulating key genes governing plant height, branching, leaf morphology, and root structure, plant architecture can be optimized to enhance light utilization and lodging resistance. Targeted manipulation of genes related to disease and pest resistance, and tolerance to drought, salinity, and temperature extremes, substantially improves resilience to biotic and abiotic stresses. Additionally, by fine-tuning yield determinants and by engineering photosynthetic pathways, yield potential can be effectively increased. Beyond productivity, genetic engineering facilitates nutritional fortification, improved sensory quality, and enhanced processing characteristics, paving the way for novel crop varieties that integrate nutrition with palatability. Looking forward, coordinated multi-gene editing, utilization of wild germplasm, strengthened field adaptability testing, and exploration of controllable epigenetic regulation represent key directions for the future. Collectively, these advances will drive plant genetic engineering toward greater precision, efficiency, and intelligence, providing a robust foundation for sustainable agricultural development.

|
Scooped by
mhryu@live.com
April 28, 11:25 AM
|
Co-translational protein folding is shaped by the vectorial nature of translation, which causes residues to emerge sequentially from the ribosome. As a result, residues whose native interaction partners lie downstream in sequence cannot immediately form their native contacts and remain transiently unsatisfied until those partners are synthesized. These unsatisfied residues are vulnerable to non-native interactions and often require the engagement of co-translational chaperones. We previously developed the Native Fold Delay (NFD) metric to quantify the time lag between the synthesis of a residue and the point at which it can form all its native contacts. Here, we present the FoldDelay web server, a freely accessible platform that extends the NFD concept into a more comprehensive framework for analyzing native residue–residue contact formation during translation. Starting from user-submitted AlphaFold or PDB structures, the site identifies all N- to C-terminal residue–residue contacts, estimates their earliest possible formation times, and integrates domain annotations to distinguish between intra- and inter-domain contacts. The server provides a suite of linked interactive visualizations that allows users to explore native contact formation dynamics and detect transiently unsatisfied regions. The FoldDelay web server is freely accessible at https://folddelay.switchlab.org.

|
Scooped by
mhryu@live.com
April 28, 2:16 AM
|
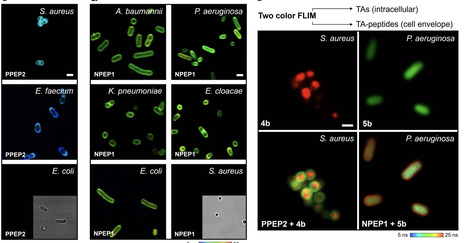
The rise of multi-drug resistant bacteria constitutes a global challenge with direct impact on millions of patients worldwide. The identification of bacterial species is essential to avoid unnecessary use of antibiotics and to minimize the emergence of additional resistance; however, there are few chemical strategies to identify resistant species in a rapid and unbiased manner. Cross-reactive arrays combine multiple sensor components to produce distinctive fingerprints for similar targets; herein, we present a cross-reactive sensing array built with long lifetime organic fluorophores. To the best of our knowledge, we synthesize the largest collection to date of bioconjugatable triangulenium fluorophores by including heteroatom and side-chain diversification for spectral and lifetime diversity, as well as water-soluble and orthogonal moieties for peptide tagging and compatibility with biological samples. After optimization, we evolve a four-fluorophore sensor array that correctly assigns all seven ESKAPEE bacterial species by combining their optical signatures. Lifetime chemical sensor arrays will open avenues for concentration-independent, reliable and sensitive optical detection without the need for prior knowledge of molecular targets. Infections caused by multi-drug resistant bacteria present a global health challenge, so the identification of bacterial species is essential to avoid the unnecessary use of antibiotics and to minimize the emergence of additional resistance. Here, the authors present a lifetime cross-reactive sensor array that correctly assigned all seven ESKAPEE bacterial species by combining their optical signatures.

|
Scooped by
mhryu@live.com
April 28, 2:07 AM
|
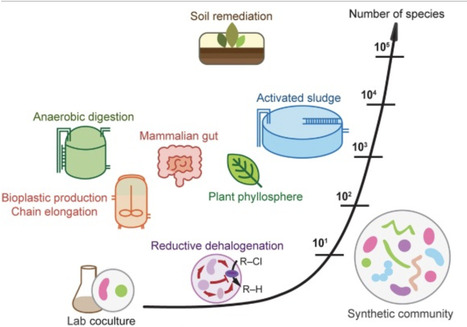
Microbial interactions hold vast potential for improving sustainability, including waste upcycling, greenhouse gas mitigation, contaminant bioremediation, and host performance. However, attempts to manage interactions in complex, open microbial communities often fall short of expectations, in part because interactions are highly context-dependent, with net effects arising from multiple component interactions, higher-order interactions, and rapid evolution—features largely revealed from studies with simplified communities. Here, we propose how these insights can be translated to complex communities by prioritizing the scales and disturbances that matter, leveraging the ecological context that constrains interactions, targeting function-critical behaviors and traits, and accounting for eco-evolutionary feedbacks. We further propose coupling top-down, trait-based, and coarse-grained approaches with bottom-up synthetic ecology approaches to diagnose failures and design robust microbial processes.

|
Scooped by
mhryu@live.com
April 28, 2:02 AM
|
The plant immune system engages cell-surface receptors that detect microbe-associated molecular patterns to initiate pattern-triggered immunity (PTI), and intracellular receptors that sense microbe-secreted effectors to activate effector-triggered immunity (ETI). Whether additional modes of microbial detection exist remains unclear. Here, we define membrane mechanosensing immunity (MSI), a third layer of cell surface immune signaling. A bacterial lipid, the main diffusible signal factor (DSF) from Xanthomonas campestris pv. campestris, acts as a membrane-active molecule that alters plasma membrane biophysical properties, activates Mechanosensitive Channel of Small Conductance (MscS)-like (MSL)-dependent immune signaling, and triggers a broad transcriptional reprogramming that overlaps with PTI and ETI. MSI modulates PTI signaling and requires both PTI and ETI components for effective disease resistance. These findings establish the sensing of metabolite-induced membrane perturbations as a mechanism of microbial detection.

|
Scooped by
mhryu@live.com
April 28, 1:52 AM
|
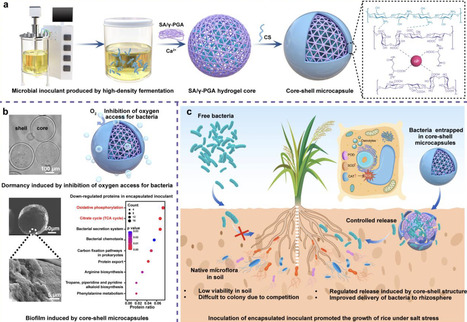
Non-spore-forming bacteria can enhance crop salinity tolerance via various strategies, but poor viability during storage and field application limits their use. Inspired by the robust structure of spore dormancy, a core-shell microcapsule consisted of sodium alginate, poly (γ-glutamic acid), and chitosan (APC) is proposed. Using Pantoea alhagi NX-11 as a model, we found that APC encapsulation significantly enhanced bacterial survival during room-temperature storage compared to free cells or alginate beads. This protective effect was further confirmed using two other non-spore-forming strains. The survival mechanism was mainly attributed to the APC-induced metabolic dormancy via suppression of the TCA cycle and oxidative phosphorylation pathways. Furthermore, inoculation with APC-encapsulated NX-11 increased dry weight of rice plant by 24.2% under salt stress in greenhouses and increased grain yield by 15.8% in saline fields, attributing to the enhanced root colonization of NX-11. Overall, this bio-inspired encapsulation strategy provides an effective approach for developing robust microbial inoculants to improve crop resilience in saline soils. Poor storage stability and soil survival limits agricultural application of non-spore forming bacteria. Here, the authors develop a hydrogel microcapsule which improves storage and soil survival, demonstrating application in improving plant salt tolerance under field conditions.
|

|
Scooped by
mhryu@live.com
April 28, 10:47 PM
|
In the context of global energy transition and carbon neutrality goals, converting lignocellulosic biomass into high-value products is essential for a circular bioeconomy. This review systematically explores technological advancements in modern biorefineries, from feedstock deconstruction to precise valorization. Pretreatment strategies-physical, chemical, and integrated pretreatment strategies-effectively overcome lignocellulose recalcitrance, achieving high-purity fractionation of cellulose, hemicellulose, and lignin. Advanced conversion technologies, including electrocatalysis, photocatalysis, biological funneling, and bio-photoelectrochemical hybrid systems, offer superior selectivity, atom economy, and energy efficiency in biomass upgrading. Addressing economic and stability challenges in scale-up, the integration of artificial intelligence and digital twins promises intelligent, carbon-neutral biorefinery systems. This review provides theoretical guidance and technical roadmaps for closed-loop resource utilization and sustainable development.

|
Scooped by
mhryu@live.com
April 28, 10:33 PM
|
Rare Earth Elements (REEs) are critical components in the low-carbon economy and high-tech industries. However, rapid depletion of high-grade reserves and geopolitical concerns have triggered a supply crisis, driving the need for innovative technologies to extract REEs from unconventional sources. Phytomining, which employs hyperaccumulator plants (“REE crops”) to concentrate REEs from soil into harvestable biomass, offers a sustainable supplementary source. However, its commercial deployment faces two major barriers: inadequate yields of high-REE biomass and the lack of viable processes to recover REEs from the complex biomass. Herein, we propose a strategic framework to advance REE phytomining. Firstly, our approach involves selecting location-specific REE crops such as the fern Dicranopteris linearis, which is abundant in (sub)tropical regions and does not require deliberate planting or intensive agronomic management to maximize phytoextraction efficiency. Secondly, we identify the up-cycling of REE-enriched biomass into products (e.g., fertilizers and catalysts) as a lever to improve the commercial viability of phytomining, beyond the conventional route of hydrometallurgical REE separation. Environmental and economic analyses indicate that this route could produce millions of tons of low-carbon, plant-based REE materials annually. Realizing this potential requires a transition from labs to fields and from research to application, necessitating multidisciplinary, cross-sector collaboration. Location-specific hyperaccumulator plants for rare earth elements that require no deliberate planting, or intensive management can maximize phytoextraction, while converting enriched biomass into products improves commercial viability.

|
Scooped by
mhryu@live.com
April 28, 9:28 PM
|
The first wave of alternative proteins and plant-based, precision-fermented and cultivated meat has moved from scientific curiosity to commercial reality. Products are on shelves, companies are publicly traded and regulators in multiple jurisdictions have begun to define pathways for approval. Yet the next decade will determine whether alternative proteins remain niche innovations or whether they mature into a resilient pillar of the global food system. The most immediate challenge is economic. Despite technical advances, cost parity with conventional animal protein remains elusive for many categories. Second, regulatory harmonization remains uneven. Public perception itself is the third axis of challenge. Sustainability claims also require scrutiny. A fourth challenge lies in biological complexity. Finally, the industry must confront its own consolidation dynamics. Venture-driven growth fuelled rapid expansion, but recent capital contraction has exposed fragile business models. . The next phase, however, will not be defined solely by molecular breakthroughs. It will depend on infrastructure, regulation, cultural legitimacy and disciplined execution.

|
Scooped by
mhryu@live.com
April 28, 9:10 PM
|
Chemical modifications on RNA represent an additional regulatory layer of gene expression analogous to epigenetic marks on DNA and histones. Over the past decade, the fundamental mechanisms of N6-methyladenosine and other mRNA modifications have been extensively characterized, establishing their roles in nearly all aspects of RNA metabolism with broad implications for physiological and pathological processes. These advances lay the groundwork for future therapeutic approaches targeting mRNA modification pathways. By contrast, emerging evidence indicates that RNA methylation on chromatin-associated RNAs (caRNAs) intersects with chromatin regulators to modulate chromatin state and transcription, adding a new dimension to epigenetic regulation with biological significance. Here we summarize established principles of post-transcriptional RNA modifications and their therapeutic potential, highlight the rapidly developing chromatin-related functions of RNA modifications as regulatory elements on caRNAs and discuss future directions that emphasize the need to investigate additional regulatory elements on these RNAs in shaping gene expression programs. This Perspective introduces the recent development of RNA modifications on chromatin-associated RNAs, discusses how they engage chromatin regulators to modulate chromatin states and direct cellular function and proposes future directions for studying RNA regulatory elements to control gene expression.

|
Scooped by
mhryu@live.com
April 28, 9:05 PM
|
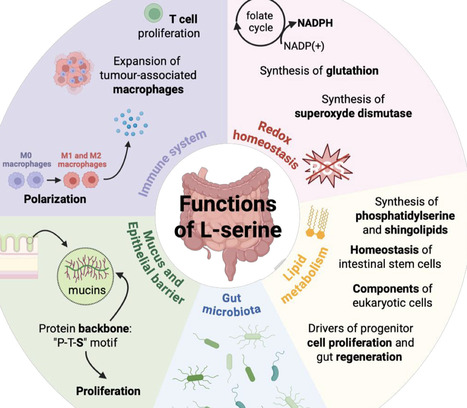
L-serine is an amino acid involved in maintaining intestinal mucosal homeostasis through its role in protein synthesis but also in redox balance, immune function and lipid metabolism. It also contributes to mucus production, the epithelial barrier integrity and gut microbiota composition. Prokaryotic cells, including Escherichia coli, have developed mechanisms to exploit L-serine to support their growth, virulence, inflammatory and/or carcinogenic properties. In this review, we summarize current findings on L-serine metabolism in the intestinal mucosal homeostasis, its interactions with microbiota, and its modulation in digestive diseases. Finally, we highlight directions for future research to target L-serine as a promising therapy. In this review, we summarize current scientific findings on L-serine metabolism in the intestinal mucosal homeostasis, its interactions with the gut microbiota, and its modulation in digestive diseases. Finally, we highlight directions for future research to target L-serine as a promising therapeutic strategy.

|
Scooped by
mhryu@live.com
April 28, 8:15 PM
|
Cell surface LPxTG motif protein (LMP) plays a crucial role in adhesion and biofilm formation during lactic acid bacteria (LAB) colonization. LMP anchors to the cell wall via its C-terminal LPxTG motif and contains functional domains that perform specific tasks. By integrating bioinformatics analysis with existing literature, this review systematically evaluates, for the first time at the domain level, the diversity of LMP in LAB and constructs a functional domain framework for understanding their roles in adhesion and biofilm formation. Annotation analysis reveals domain diversity across LAB strains: mucin-binding and extracellular-matrix-binding domains are widespread, while others exhibit strain-specific characteristics. We also speculate that certain LMP domains may form pilus-like adhesion structures via sortase C-mediated polymerization. Overall, this domain-level elucidation of LMP probiotic mechanisms provides a theoretical basis for probiotic strain screening and functional optimization, and offers new insights into structural and functional localization of LMP on the LAB surface.

|
Scooped by
mhryu@live.com
April 28, 6:58 PM
|
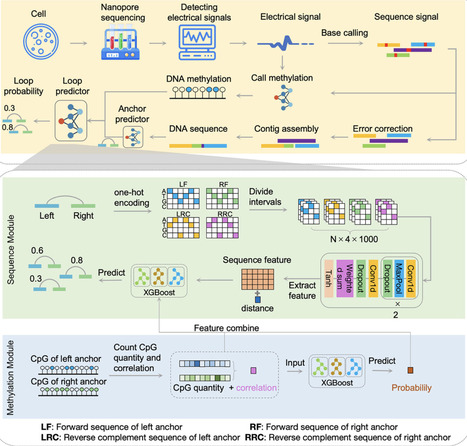
Chromatin loops play a crucial role in gene regulation and cellular function, providing key insights into understanding the 3D structure of the genome and its impact on cellular homeostasis. Nanopore sequencing technology, with its advantages in simultaneously detecting sequences and methylation patterns, brings new opportunities for studying 3D genome structures. We introduce NanoLoop, the first algorithmic framework attempting to predict genome-wide chromatin interactions using Nanopore data. In experiments across four human lymphoblastoid cell lines, NanoLoop demonstrated excellent predictive performance and cross-cell line generalization capabilities. We also discovered four distinct methylation patterns at loop anchors that influence histone modification levels and determine various loop types. NanoLoop further predicted previously uncharacterized long-range chromatin loops, highlighting the potential link between DNA methylation and 3D genome organization and providing new insights into the complex regulatory relationships between epigenetic modifications and 3D genome organization.

|
Scooped by
mhryu@live.com
April 28, 11:30 AM
|
Mycobacterial pathogens remain major global health threats, exacerbated by both rapid acquisition of antibiotic resistance and the formidable drug diffusion barrier presented by the rigid mycomembrane. These challenges have renewed interest in antimycobacterial peptides (AMyPs), a diverse class of short amphiphilic sequences capable of rapidly killing both drug-sensitive and drug-resistant mycobacteria. Beyond their intrinsic potency, AMyPs can synergize with existing antibiotics and exhibit markedly slower resistance development relative to conventional small molecules. In this review, we synthesize recent advances spanning natural bioprospecting, mechanism-guided rational design, and chemical optimization strategies that have yielded increasingly potent and selective AMyP candidates. We further highlight the rapid emergence of artificial intelligence–driven discovery platforms, which leverage machine-learning models trained on curated, mycobacteria-specific data sets to predict and refine novel AMyPs with growing accuracy. Together, these technologic, biologic, and computational advances outline a rapidly expanding landscape for AMyP-based therapeutic development and establish a foundation for next-generation antimycobacterial drug design. amp

|
Scooped by
mhryu@live.com
April 28, 2:33 AM
|
Here, we report selective covalent assembly of Gram-negative bacteria and synthetic polymers into functional living materials. We discovered that triblock polymers decorated with vinyl sulfone (VS), a motif that forms a stable covalent bond with cysteine residues on surface proteins, yielded stable covalent assembly with bacterial cells. Notably, we found that these assemblies were uniquely cell-type-specific, occurring only in Gram-negative bacteria, highlighting differences in surface structures and the macromolecular diffusion barrier across bacterial species. We also demonstrated that assembling engineered cells into materials results in situ melanin production from living materials, with robust biocontainment and mechanical reinforcement. Spontaneous enrichment of tyrosine-derived red pigments in the supernatant showcases the effect of confinement in a complex biochemical pathway. This work establishes a platform for encoding complex, engineered, and evolved functions of Gram-negative bacteria into synthetic materials, enabling the development of a wide range of material-based bioreactors.

|
Scooped by
mhryu@live.com
April 28, 2:11 AM
|
Uncovering what drives select biomolecules to form phase-separated condensates in vivo and identifying their physiological significance are topics of fundamental importance. Here, we show that nitrogen-starved E. coli produces long-chain polyphosphates, which scaffold the RNA chaperone Hfq into high molecular weight complexes, which eventually phase separate together with components of the RNA translation and processing machinery. The presence of polyphosphate within these condensates controls Hfq function by selectively stabilizing polyadenylated RNAs involved in transcription and protein translation and by promoting interactions with translation- and RNA-metabolism-associated proteins involved in de novo protein synthesis. Lack of polyphosphate significantly impairs condensate formation, increases cell death, and hinders recovery from N-starvation. In functional analogy, we demonstrate that polyP contributes specifically to the formation of Processing (P)-bodies in human cell lines, revealing that a single, highly conserved and ancestral polyanion serves as a modulator for functional phase-separated condensate formation across the tree of life.

|
Scooped by
mhryu@live.com
April 28, 2:05 AM
|
Competition for a single limiting resource is expected to lead to competitive exclusion, yet diverse microbial communities persist even in nutrient-poor environments. Cross-feeding of essential metabolites is one mechanism that can promote coexistence between species, but its contribution is difficult to pinpoint experimentally. Here, we studied a prototroph-auxotroph pair growing on a single carbon source in chemostats. In minimal medium, the prototroph Comamonas testosteroni (Ct) supplied thiamine to the thiamine-auxotroph Ochrobactrum anthropi (Oa), allowing stable coexistence in agreement with consumer-resource theory. Contrary to our expectation that supplying thiamine would remove the dependency and lead to exclusion of Ct, coexistence persisted even when thiamine was supplemented. Our theoretical analysis showed that coexistence between competitors can be maintained by trace concentrations of an additional metabolite if it is taken up at sufficiently high affinity by the weaker competitor. Consistent with this prediction, targeted metabolomics and spent-medium assays identified growth-enhancing compounds at micromolar concentrations in Oa spent medium and as residues in fresh medium. Model analysis further showed that such weak positive effects can qualitatively change coexistence outcomes in chemostats while remaining undetected in standard batch interaction assays. Together, our results show that trace metabolites and subtle positive effects can reshape coexistence outcomes and should be incorporated into ecological models and interaction measurements.

|
Scooped by
mhryu@live.com
April 28, 1:58 AM
|
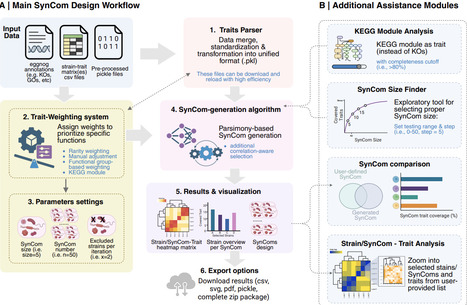
Synthetic microbial communities (SynCom) are essential tools for dissecting the causal mechanisms in host-microbiota interactions. To date, however, SynCom design suffers from a lack of standardization, typically oscillating between arbitrary strain selection and computational pipelines that misalign with experimental design. As microbiome research transitions toward functionally defined community systems with reproducible experimental outcomes, there is a strong need for a user-friendly platform that integrates multi-dimensional genomic and/or biological data into a standardized and tailored SynComs design. Here, we present SynCom101, a web-based platform that democratizes the design of reproducible, hypothesis-driven SynComs. SynCom101 accommodates diverse input formats including genomic annotations and laboratory-obtained phenotypic traits, allowing users to customize their design criteria with high flexibility. The platform utilizes a parsimony algorithm to ensure computational scalability for large datasets, complemented by an optional correlation-aware mode to account for microbial compatibility and co-occurrence patterns when ecological interactions among strains are available. A core innovation of SynCom101 is its suite of trait-weighting modules, which empowers researchers to strategically guide the selection algorithm toward maximal functional trait coverage, the emulation of natural community architectures, or the enrichment of positively correlated microbial assemblages to enhance community stability. We showcase the functionalities of the platform by in silico design of communities from different datasets, demonstrating its capacity to generate concise, functionally prioritized SynComs aligned with targeted design objectives. By providing a transparent, parameter-documented workflow, SynCom101 ensures that community design is no longer a "black box" but a reproducible scientific record. This platform establishes a necessary standard for in silico community assembly, facilitating the transition from descriptive microbiome studies toward high-throughput, predictive functional screening and cross-study comparability. https://syncom101.bioinformatics.nl/ The datasets used for case studies are available on Zenodo (https://doi.org/10.5281/zenodo.18310451). The source code is available at Git (https://git.wur.nl/jiayi.jing/syncom101).
|
 Your new post is loading...
Your new post is loading...