Your new post is loading...

|
Scooped by
mhryu@live.com
April 25, 11:56 PM
|
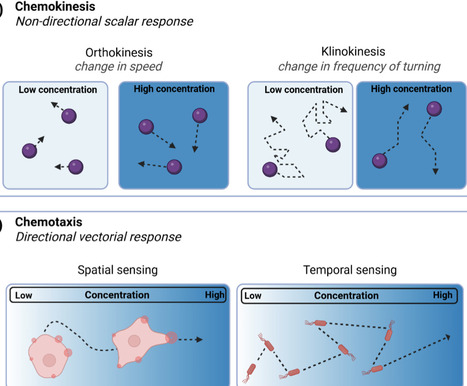
Chemotaxis is the directional motion of objects in response to chemical gradients, a process that drives navigation in micro/nanomotors, enabling a bio-inspired transition toward autonomous direction control. However, current studies lack consistent mechanistic justification, definition, and standardized experimental validation. This review summarizes key physical principles governing active chemotaxis, surveys experimental strategies for generating chemical gradients and quantifying responses, and examines how individual chemotactic mechanisms and long-range, anisotropic chemical interactions give rise to emergent collective behaviors, while outlining prospective applications. Together, these efforts establish a principled engineering framework for advancing chemotaxis as a robust functional navigation modality in synthetic micro/nanomotors. This Review Article examines artificial chemotaxis in micro/nanomotors, detailing mechanisms, experimental strategies, and collective behaviors. It highlights the need for standardized validation and explores applications in biomedical and industrial fields, emphasizing the challenges and potential of synthetic chemotaxis.

|
Scooped by
mhryu@live.com
April 25, 11:43 PM
|
Microbiomes perform critical functions for their hosts, and understanding microbiome variation is important for both basic and applied science. However, host traits alone cannot explain the entirety of microbiome variation, because, alongside host traits, microbiomes are shaped by multiple ecological processes. Researchers have thus turned to theories of island biology, conceptualising animal hosts as islands and animal microbiomes as metacommunities that assemble within and disperse between host islands. To develop realistic models, this host-as-island metaphor must be examined by explicitly comparing geological and host islands. Here, we critically examine the host-as-island metaphor by evaluating how microbiome variation is shaped by the four metacommunity processes that explain biodiversity on geological islands: local interspecies interactions, local selection, dispersal, and stochasticity. Key differences between host islands and geological islands include the complexity of microbiome transmission networks arising from host mobility and sociality and the capacity of hosts to evolve to control their microbiomes. We conclude with discussions of how eco-evolutionary dynamics differ between geological islands and host islands, and the reciprocal relevance of island biology and microbiome science.

|
Scooped by
mhryu@live.com
April 25, 11:29 PM
|
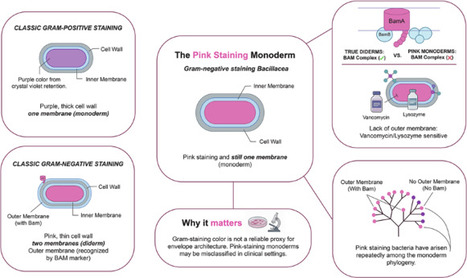
Gram staining has guided microbiology for over a century by coloring cells purple or pink, a read-out thought to distinguish monoderms (single membrane, thick peptidoglycan) from diderms (inner and outer membranes with thin wall). Here we show this rule fails repeatedly across Bacillaceae lineages historically deemed “Gram-positive”. By combining light and transmission-electron microscopy, antibiotic-sensitivity assays and comparative genomics across 57 strains, we identify “Gram-negative-staining monoderms” lacking outer membrane yet retaining thick peptidoglycan walls. These bacteria lack lipopolysaccharide- and β-barrel assembly-genes and remain highly susceptible to vancomycin and lysozyme (agents normally excluded by diderm envelopes), demonstrating functional monoderm status. Surprisingly, teichoic-acid biosynthetic pathways are patchily distributed and do not predict staining behavior. This discovery calls into question the textbook purple-or-pink dichotomy, decoupling stain color from membrane architecture. Clinically, misidentifying pink-staining Bacillaceae (including emerging pathogens such as Bacillus infantis) risks inappropriate therapy, whereas genome-guided diagnostics enable precise antibiotic stewardship. Genomic and microscopy evidence shows that the Bacillaceae, assumed to only stain Gram-positive, actually have some lineages staining Gram-negative, yet unexpectedly maintaining a thick wall and no outer membrane, challenging Gram staining assumptions.

|
Scooped by
mhryu@live.com
April 25, 11:20 PM
|
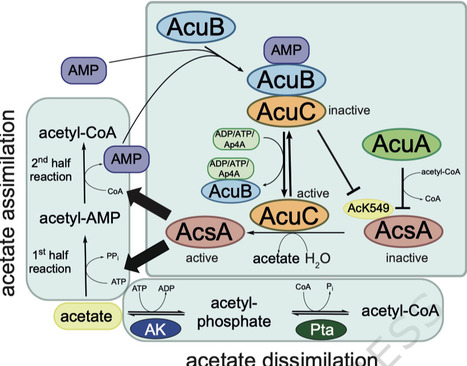
Bacteria adjust their metabolism to the cellular energy state. AMP-forming acetyl-CoA-synthetase AcsA generates acetyl-CoA from acetate, ATP and CoA. In Bacilli, including Bacillus subtilis and Geobacillus stearothermophilus, AcsA is reversely transcribed upstream of the acu-operon encoding for the proteins AcuA, AcuB and AcuC. Lysine-acetyltransferase AcuA uses acetyl-CoA to acetylate and inactivate AcsA, while AcuC re-activates AcsA activity by deacetylation. How the counteracting activities of AcuA and AcuC are regulated is not understood. Here, we close this gap of knowledge and perform a structure-function analyzes on AcuB. These reveal AcuB forming a scissor-shaped dimer with each monomer consisting of an N-terminal Bateman domain binding to adenine nucleotides and a C-terminal ACT domain. Structural and biochemical studies as well as molecular dynamics simulations support that AMP bound AcuB binds and inhibits AcuC. Our data describe another layer of regulation of AcsA activity in Firmicutes coordinating acetate assimilation and dissimilation by the energy sensor AcuB. Bacteria adjust their metabolism to the cellular energy state. Here, authors identify a layer of regulation of AMP-forming acetyl-CoA synthetase AcsA activity by AcuB acting as energy sensor inhibiting the AcsA deacetylase AcuC in presence of AMP.

|
Scooped by
mhryu@live.com
April 25, 11:15 PM
|
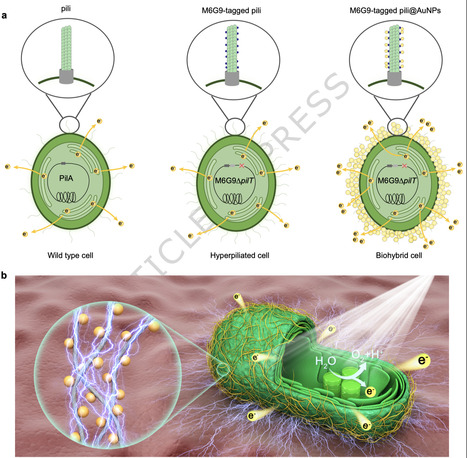
The efficient extraction of electrons from photosynthetic microorganisms remains a critical challenge in living biophotovoltaics (BPV). While nanomaterials can facilitate electron transport, their stochastic adsorption leads to inefficient material-wasteful interfaces. Here, we demonstrate a controllable approach to direct the targeted assembly of gold nanoparticles (AuNPs) onto the type IV pili of Synechocystis sp. PCC 6803 by using a genetically encoded gold-binding peptide. This approach creates a spatially precise conductive nano-bio interface on the cell envelope that serves as a dedicated electron conduit between photosynthetic electron transport chains (PETCs) and electrodes. This nano-bio interface enhances electron transfer through synergistic improvements in interfacial charge transfer and biofilm density, ultimately yielding a four-fold increase in photocurrent density, while using two orders of magnitude less gold than non-targeted strategies. Moreover, the AuNPs can be transferred from inactivated to fresh cells, indicating a potential pathway for long-term stability. This work establishes a generalizable strategy for the rational design of conductive interfaces on living cells, with implications for biophotovoltaics, microbial electrosynthesis, and next-generation biohybrid devices. Inefficient extracellular electron transfer of cyanobacteria impedes power output in biophotovoltaics. Here, the authors assemble a gold conductive layer on the cell surface by attaching gold-binding peptide to the pili structure, achieving a four-fold increase in photocurrent generation.

|
Scooped by
mhryu@live.com
April 25, 4:01 PM
|
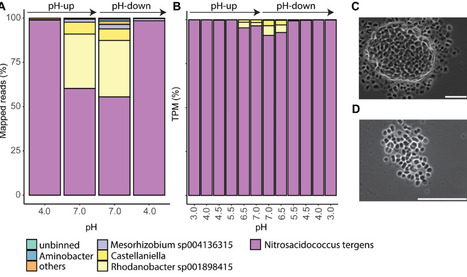
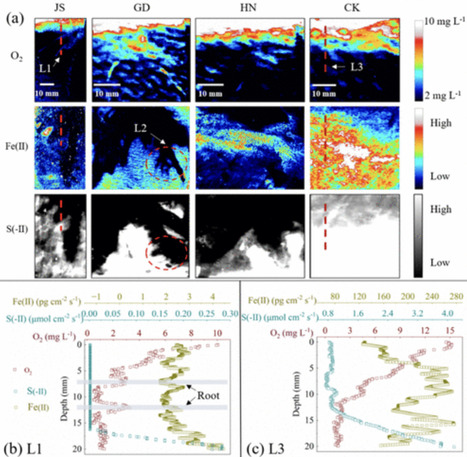
Acidophilic ammonia-oxidizing bacteria (AOB) have only recently been discovered. These organisms hold great promise for acidic wastewater treatment; however, their physiology remains poorly understood compared to that of neutrophilic AOB. Here, we investigated the physiology of the acidophilic AOB “Candidatus Nitrosacidococcus tergens” sp. RJ19 across a broad pH range (2.5–7.0) using a specialized bioreactor system. We monitored nitrogen (N) transformations combined with microbial community composition and transcriptomic profiles, focusing on nitrogen metabolism and proton stress responses. Our results show that above pH 6.0, “Ca. Na. tergens” performs complete and stoichiometric conversion of ammonium to nitrite, coinciding with isotopic fractionation effects specific for ammonia oxidation and increased expression of key ammonia oxidation genes. The apparent absence of nirK and cycA did not impede ammonia oxidation, suggesting that these genes are non-essential in this context. Below pH 6.0, nitric oxide and nitrate accumulated, and nitrous oxide (N₂O) levels, although negligible compared to the other N-compounds, peaked near pH 4.0. Stable isotope analysis, including the site-specific 15N-enrichment at the inner (α) and outer (β) nitrogen positions of the N2O molecule, indicated nitrifier-denitrification as the source of N₂O, supported by the highest norB expression at this pH. These findings provide new insights into the acid-tolerant physiology of “Ca. Na. tergens” and advance its potential application in engineered nitrogen removal systems under acidic conditions.

|
Scooped by
mhryu@live.com
April 25, 3:45 PM
|
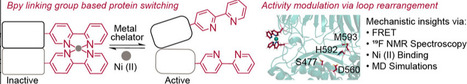
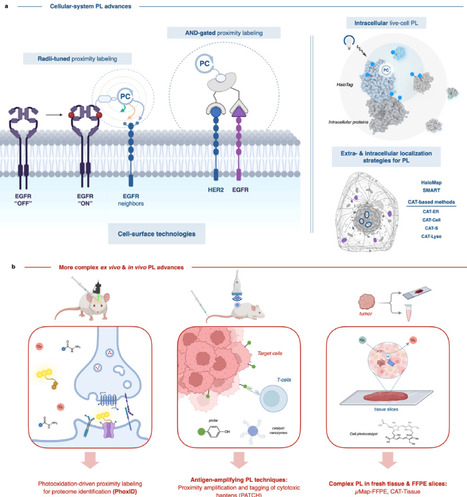
Protein–protein interactions (PPI) and spatially restricted molecular contacts govern cellular function, yet many are poorly captured by classical biochemical approaches that rely on cell lysis or stable complex isolation. Proximity labeling (PL) technologies have transformed interactome analysis by enabling covalent tagging of biomolecular neighborhoods directly within intact cells, tissues, and living organisms. By generating short-lived reactive species, PL provides spatially and temporally resolved snapshots of molecular organization under native conditions. Recent advances across enzymatic, chemical, and photocatalytic PL platforms have expanded control over labeling radius, kinetics, and activation, while reducing background and enabling microenvironment-specific targeting. Hybrid genetic-chemical and optogenetic strategies further extend PL beyond mapping toward proximity-based signal amplification and functional interrogation. This review focuses on the most significant methodological and conceptual advances in proximity labeling reported over the past two years, highlighting how these developments have enabled discovery of previously inaccessible interaction networks, including membrane assemblies, chromatin complexes, and in vivo protein microenvironments. We conclude by outlining key challenges and future opportunities for proximity labeling in interactome mapping and amplification.

|
Scooped by
mhryu@live.com
April 25, 3:36 PM
|
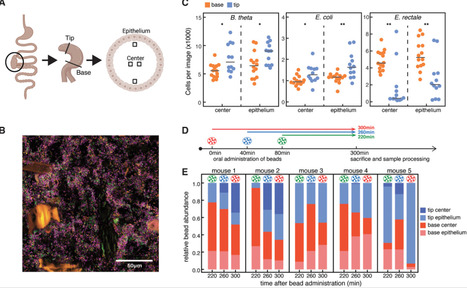
Spatial structure can functionally determine ecological interactions and evolution of microbial communities. The gut microbiota is known to be spatially structured longitudinally along the gastrointestinal tract, but micro-scale structure in the gut lumen has not been extensively explored. Here, we show that bacteria cluster within species in the cecum of gnotobiotic mice. We find that clustering is not driven by active swimming, antibody-mediated aggregation, or factors exclusive to the host, but likely due to bacterial growth in the matrix of gut content. In samples from mice and humans, we show that upper large-intestinal content behaves as a nonNewtonian fluid that changes its viscoelastic properties under the force of gut contractions. We argue that microbial growth in the gel-like structure of cecum content can lead to micro-scale bacterial clustering, which is periodically disrupted by peristalsis-driven shear thinning and clearance. Our study shows mechanistically how spatial structure in the gut emerges through the interplay between microbial and host physiology and highlights the possibility of host control over gut microbiota distribution through gut contractions.

|
Scooped by
mhryu@live.com
April 25, 3:24 PM
|
Microbiomes associated with both the human gut and plant root rhizosphere are essential for the maintenance of host health and function as holobionts where both the host and microbiome operate as an integrated unit. Though substantial differences exist in both host biology and environment, these systems share functional parallels: both are enriched by host-derived nutrients, undergo successional shifts during development and maintain core microbiomes that are taxonomically variable yet functionally redundant. Central to both systems is the balance that is maintained where beneficial microbes regulate nutrient cycling, modulate host immune response and suppress pathogens in the presence of biotic and abiotic influences that may serve to disrupt this equilibrium. When dysbiosis occurs, there is a disruption in the composition and/or function of the associated microbiome and a loss of beneficial functional guilds which results in a reduction in host fitness. These shared dynamics underscore dysbiosis as a cross-kingdom pathology that may be treated with similar interventions. Probiotics and prebiotics mirror microbial inoculants and organic amendments, synbiotics incorporate both biotic and abiotic factors, while fecal and soil microbiome transplants represent parallel strategies to restore a beneficial microbiome. By framing dysbiosis within a One Health perspective and illustrating the connectedness between human and plant health, this review advocates for microbial stewardship as a unifying strategy to mitigate disease, enhance resilience and ensure sustainable health across both systems.

|
Scooped by
mhryu@live.com
April 25, 2:24 PM
|
Information regarding the biodegradation of materials used in conservation ecology is limited. We evaluated the biodegradability of two polyurethane (PU) foams applied in wildlife management. The samples were buried in soil for 120 days to investigate their morphological and physicochemical changes. At the end of the study, the bacteria attached to the PU samples were isolated and identified. Weight loss analysis revealed stronger degradation in the aliphatic PU foam (11.70%) compared to the aromatic foam (1.85%). FTIR spectra indicated the breakdown of urethane and ester bonds, as shown by comparing peak area ratios between pristine and degraded samples. SEM analysis revealed surface alterations and microbial colonization. Eight bacterial strains isolated from the soil, identified as PU degraders, belonged to the genera Pseudomonas, Bacillus, and Klebsiella. All the strains had the ability to utilize Impranil DLN, an anionic PU dispersion, as a carbon source, indicating their potential for PU degradation.

|
Scooped by
mhryu@live.com
April 25, 2:14 PM
|
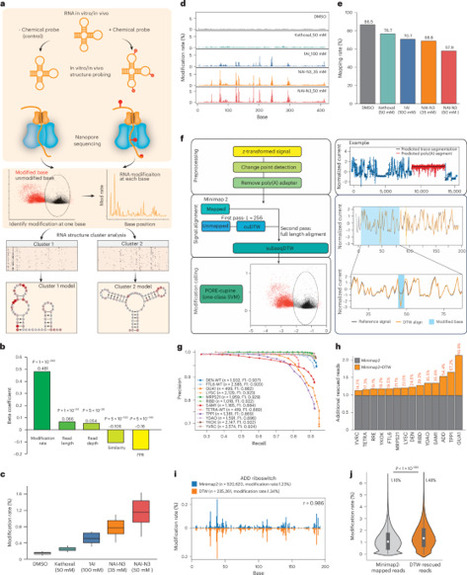
The extent to which an RNA folds into structure ensembles and how different structures in the ensemble regulate eukaryotic gene expression is not fully understood. Here, we coupled chemical probing with direct RNA sequencing to identify structure modifications along a single RNA molecule (sm-PORE-cupine). We used direct signal alignment in addition to base mapping to increase the percentage of mappable sequences and showed that Bernoulli mixture model clustering can separate structure ensembles accurately. We applied sm-PORE-cupine to identify isoform-specific structure ensembles along the SARS-CoV-2 genome and structure ensembles in the Candida albicans transcriptome. We observed that RNAs are more structurally homogeneous in vitro, at higher temperatures and in the 3′ untranslated regions of C. albicans. Structure ensembles are associated with changes in translation efficiency and decay in C. albicans, and we validated translation changes using reporter assays. sm-PORE-cupine expands the existing toolbox for studying RNA structure and function in diverse transcriptomes. sm-PORE-cupine combines SHAPE-based chemical probing with nanopore-based direct RNA sequencing to identify RNA structural ensembles in the SARS-CoV-2 genome and the Candida albicans transcriptome.

|
Scooped by
mhryu@live.com
April 25, 1:49 PM
|
Bacteria in fluctuating environments must balance the high metabolic costs of motility against risks of bacteriophage predation and immune clearance. While flagellar trade-off mechanisms are well-documented, regulation of type IV pilus (T4P) activity during environmental transitions remains unclear. We show that Pseudomonas aeruginosa uses an energy-dependent idling strategy to synchronize T4P-mediated surface motility with nutrient availability. In nutrient-depleted stationary phase, T4P transcription and protein levels remain constant, pre-assembled machines persist at the cell pole, yet cells produce only sparse, truncated pili that extend and retract slowly. Using a single-cell ATP biosensor, we show that T4P dynamics respond directly to cellular adenylate energy charge. Carbon source addition rapidly elevates intracellular ATP, reactivating pre-assembled T4P within minutes. This bypasses de novo protein synthesis, restoring pilus number, length, and extension/retraction rates. This rapid response drives opportunistic biofilm dispersal but, at the same time, creates an immediate tradeoff: reactivated T4P restore susceptibility to pilus-specific phages upon nutrient upshift. Thus, energetic gating of T4P enables P. aeruginosa to minimize exposure to phages during starvation while remaining poised for rapid reactivation. Importantly, T4P promote resistance to opsonization and phagocytosis by macrophages and neutrophils. Upon nutrient upshift, full T4P activity therefore supports dispersal and host colonization while conferring immune protection, revealing a fundamental dispersal-infection tradeoff at the host-microbe interface in fluctuating environments such as the lung and gut.

|
Scooped by
mhryu@live.com
April 25, 12:52 PM
|
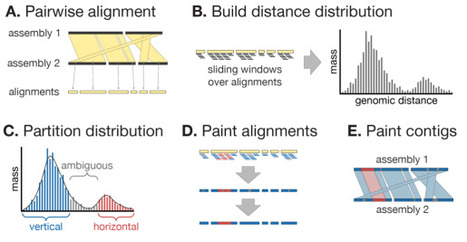
The inference and removal of horizontally acquired genomic regions is a crucial step in phylogenomics analyses for evolutionary studies. Existing tools perform well on clonal lineage-focused datasets on the scale of hundreds of genomes, but are limited in their ability to analyse larger or more diverse datasets. Here we present Verticall, a tool to identify recombinant regions in bacterial assemblies and generate recombination-free phylogenies, which scales to thousands of genomes from clonal to genus-level diversity. Verticall uses a non-parametric approach to assign genomic regions as horizontally or vertically related based on the distribution of pairwise genetic distances between genomes. Recombination-free phylogenetic trees may be inferred by either calculating a pairwise genetic distance matrix from vertical-only regions (distance-tree approach) or by pairwise comparisons of all genomes to a reference and then masking horizontally acquired regions in a pseudo-alignment to the reference (alignment-tree approach). We demonstrate Verticall's performance using four publicly available whole-genome sequence datasets of varying sample sizes (range: 154 - 4,857 genomes) and evolutionary scales (ranging from within-lineage to genus-wide diversity). Across all four datasets, Verticall showed comparable or superior performance to the established tools Gubbins and ClonalFrameML in terms of computational efficiency, plausibility of inferred phylogenetic trees, and recovery of temporal signal for molecular dating. Our results show that Verticall is a useful tool to more efficiently and accurately detect recombination, particularly applied to datasets for which existing tools are limited, including large datasets with hundreds to thousands of genomes and those that span entire species or genera. Verticall is available free and open source at https://github.com/rrwick/Verticall.
|

|
Scooped by
mhryu@live.com
April 25, 11:45 PM
|
Advances in live-cell fluorescence microscopy have enabled us to visualize single molecules (such as mRNAs and nascent proteins) in real time with high spatiotemporal resolution. However, these experiments generate large datasets that require complex computational processing pipelines to derive meaningful and quantitative information, which is a technical barrier for many researchers. Here, we introduce MicroLive, an open-source Python-based application for quantifying live-cell microscopy images. MicroLive provides an interactive Graphical User Interface (GUI) to perform key tasks, including cell segmentation, photobleaching correction, single-particle detection/tracking, spot intensity quantification, inter-channel colocalization, and time-series correlation analysis. As a ground-truth testing dataset, we used synthetic live-cell imaging data generated with the rSNAPed toolkit, demonstrating accurate extraction of biologically relevant parameters. Microscopy images of U-2 OS cells expressing a gene construct smHA-KDM5B-BoxB-MS2 were used to demonstrate the use of this software.

|
Scooped by
mhryu@live.com
April 25, 11:38 PM
|
Higher-order genetic interactions have profound implications for understanding the molecular mechanisms of phenotypic variation, yet they remain poorly characterized. Most studies focus on pairwise interactions because high-throughput screens over the vast combinatorial space are challenging. Here, we develop Dango, a computational method based on a self-attention hypergraph neural network, to predict higher-order genetic interactions among groups of genes. As a proof of concept, we provide predictions for over 400 million trigenic interactions in the yeast S. cerevisiae, greatly expanding their quantitative landscape. Dango accurately predicts trigenic interactions and reveals biological functions related to cell growth. We further incorporate protein embeddings and uncertainty estimation to improve biological relevance and interpretability. Moreover, predicted interactions serve as genetic markers for growth responses across diverse conditions. Together, Dango enables a more complete map of complex genetic interactions that shape phenotypic diversity.

|
Scooped by
mhryu@live.com
April 25, 11:27 PM
|
Freshwater lakes are globally significant sources of potent greenhouse gases (GHGs), but how their GHGs emissions respond to changing nutrient levels remains unclear. Here, we demonstrated that nitrous oxide (N2O) production pathways in lake sediments are tightly linked to trophic state, whereas methane (CH4) production appears to be multifactorial Through global metagenomics and controlled batch experiments. In eutrophic sediments, N2O is efficiently removed through complete denitrification, with nitrification serving as the main production pathway, whereas oligotrophic sediments produce N2O primarily via incomplete denitrification. By simulating nutrient transitions using a cross-inoculation experiment, we further revealed that lake sediments systematically shift between these N2O production pathways as their trophic state changes, from denitrification-driven to nitrification-dominated during eutrophication, with the inverse pattern during oligotrophication. Consequently, N2O emissions can be effectively mitigated by inhibiting nitrification in eutrophic lakes and restricting incomplete denitrification in oligotrophic ones. Our findings establish trophic status as a key driver of N2O production sources in lake sediments. Trophic status is a critical regulator of N2O dynamics in freshwater lake sediments, with implications for predicting GHG fluxes under global change scenarios.

|
Scooped by
mhryu@live.com
April 25, 11:15 PM
|
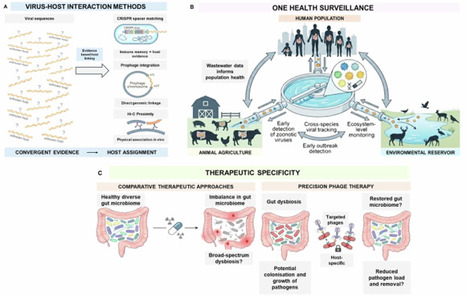
The Gut microbiome-virome dynamics and interactions Collection highlights gut viruses, mainly bacteriophages, as determinants of microbial community structure and host-relevant functions across human, animal, and environmental systems. The Collection welcomes studies that quantify virus-microbe interactions, evidence-based linking viruses to microbial hosts, characterise infection dynamics, and connect them to ecological or clinical outcomes. It prioritises methodological rigour and translational relevance in diagnostics, surveillance, and phage-based interventions.

|
Scooped by
mhryu@live.com
April 25, 4:03 PM
|
The gut-liver-brain axis is central to metabolic and neurological homeostasis and is mediated by host- and microbiota-derived metabolites. Disruptions in this axis contribute to complex disorders, underscoring the need for targeted, multi-metabolite interventions. Here, we engineered commensal Lactobacillus plantarum WCFS1 strains to specifically modulate metabolites dysregulated in hepatic encephalopathy (HE), a disorder driven by hyperammonemia and amino acid imbalance. One strain couples ammonia assimilation with branched-chain amino acid (BCAA) biosynthesis, whereas the other enhances L-glutamine utilization to suppress ammonia generation. In two preclinical HE models, these strains reduced systemic ammonia by up to 10-fold, restored BCAA and L-glutamine balance, and improved anxiety-like and cognitive behaviors. Notably, they outperformed rifaximin, a clinically used HE therapy, while preserving gut microbiota diversity. These findings establish engineered commensals as a modular, responsive platform for multi-metabolite modulation of host-microbiota metabolism, offering a programmable strategy to restore metabolic homeostasis in disorders of the gut-liver-brain axis.

|
Scooped by
mhryu@live.com
April 25, 3:50 PM
|
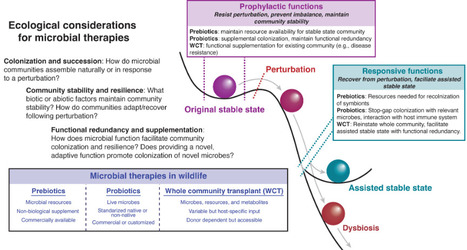
Microbial ecology is increasingly incorporated into human and animal medicine via the study and purposeful manipulation of host-associated microbiomes. Microbial therapies—treatments with the aim of beneficially modulating microbiomes—are a burgeoning area of research and industry. These microbial therapies include prebiotic dietary items, live probiotics, and whole microbiota transplants (e.g., fecal microbiota transplants). Although microbial therapies for humans and domestic animals are now widely produced for commercial use and application, evidence supporting the efficacy of commercial microbial therapies is mixed. We suggest that microbial therapies are most effective when paired with concepts from ecology and rigorous empirical research. This is particularly relevant for the development and use of microbial therapies in wildlife animal species, in which we see large-scale variation in microbial communities across hosts of varying ecologies. Identifying and developing microbial therapies that can simultaneously be accessible and effective in a variety of hosts poses a novel challenge for microbial ecologists, animal scientists, and human and animal medical professionals. In addition to pre- and probiotics, we suggest that whole microbiota transplants provide a method of microbial supplementation that may better align with species-specific microbial ecology. Moving forward, emerging methods used in human medicine such as machine learning, network analysis, and microbiome engineering using high-throughput culturomics will likely be key to identifying and applying functionally relevant (e.g., disease suppressive) microbial taxa for wildlife therapies.

|
Scooped by
mhryu@live.com
April 25, 3:42 PM
|
Gene expression analysis has evolved substantially over the past 25 years, from early transcript surveys using expressed sequence tags and microarrays to RNA sequencing, and more recently to single-cell and spatial transcriptomics. These successive waves have expanded measurement scale and resolution, enabling systematic discovery of transcriptional programmes, inference of gene regulatory networks, and increasingly direct links between transcriptomic insight and therapeutic strategies that modulate gene expression. In this Perspective, we synthesize major methodological milestones with bibliometric trends in leading bioinformatics journals to describe four revolutions that redefined gene expression analysis. We also map widely used computational tools onto a common timeline by analysing 70 78 831 open-access full-text articles, illustrating how enduring statistical frameworks coexist with rapidly growing end-to-end analysis ecosystems. We highlight current challenges and emerging directions in core bioinformatics approaches for gene expression analysis. Looking ahead, we argue that the next era will be defined less by generating new datasets and more by organizing, searching, and reusing transcriptomic and multimodal information at scale. We propose three future directions: consortium-scale searchable transcriptomic knowledgebases, foundation models for gene expression analysis, and programmable regulatory design for engineered control of gene expression. The landscape of gene expression analysis is shifting from descriptive measurement towards queryable, predictive, and programmable gene expression biology.

|
Scooped by
mhryu@live.com
April 25, 3:27 PM
|
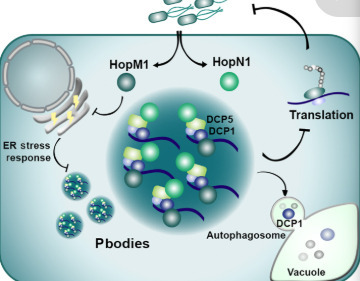
Pathogens use sophisticated strategies to modulate host protein homeostasis by targeting proteolytic pathways, but their impact on protein synthesis remains elusive. We report that pathogenic bacteria Pseudomonas syringae (Pst) targets ribonucleoprotein condensates, known as processing bodies (P-bodies), to attenuate host translation through two effectors with liquid-like properties. We uncovered a previously unknown link that Pst-mediated repression of the endoplasmic reticulum stress response is required for P-body assembly. Furthermore, we identify a functional link between P-bodies and autophagy, demonstrating that autophagic clearance of P-bodies is crucial for maintaining the balance between translationally active and inactive messenger RNAs. Together, our findings provide insights on how host translation is attenuated by bacteria to dampen plant immunity and uncover unknown connections between ER stress responses and autophagy with P-body dynamics.

|
Scooped by
mhryu@live.com
April 25, 2:33 PM
|
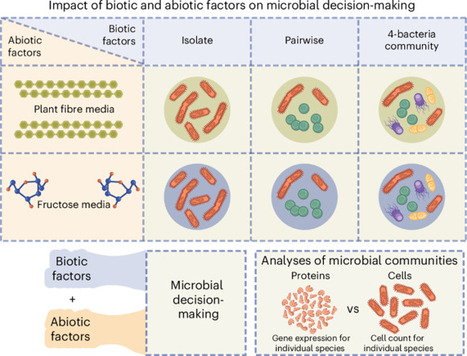
Microbial coexistence in complex communities requires mechanisms that minimize competition and optimize resource use. However, the mechanisms by which these ecological strategies are executed remain poorly understood. Here we show that bacteria modulate protein abundance in response to specific community members, reducing functional redundancy and promoting metabolic complementarity. Using synthetic gut-derived consortia exposed to distinct carbon sources, we systematically profiled proteomic responses of individual species across isolate, pairwise and 4-member communities. We found that biotic interactions, rather than abiotic conditions, were the dominant drivers of proteomic variation. These interactions led to reproducible, partner-specific expression shifts that significantly reduced functional overlap and were frequently associated with increased community productivity. Together, these findings highlight gene expression as a means by which microbes implement ecological strategies in community contexts. Through this regulatory plasticity, microbes dynamically reshape their realized niche through protein abundance modulation, enabling them to partition metabolic space and stabilize community structure. Biotic interactions modulate protein abundance, reducing functional redundancy and increasing productivity in complex bacterial communities.

|
Scooped by
mhryu@live.com
April 25, 2:19 PM
|
Microorganisms in the ruminant gastrointestinal tract play key roles in lignocellulose degradation and energy conversion. Prokaryote-infecting viruses play a pivotal role in shaping host abundance and metabolism. Despite their importance, host–virus prediction in this environment remains limited, partly due to the lack of specialized clustered regularly interspaced short palindromic repeat spacer datasets. Here, RuSpacer, a database of 181,023 clustered regularly interspaced short palindromic CRISPR repeat spacers extracted primarily from publicly available rumen-associated prokaryotic genomes, was established. Each spacer is annotated with the taxonomic identity of the genome from which it was derived. RuSpacer enables host–virus prediction via spacer–protospacer matching, particularly in the rumen ecosystem. It can also be integrated with existing publicly available spacer datasets and used for host–virus prediction in environments other than the rumen. Overall, this resource supports research on host–virus interactions, microbial ecology, and virus-based biocontrol strategies in livestock and other complex microbiomes.

|
Scooped by
mhryu@live.com
April 25, 2:11 PM
|
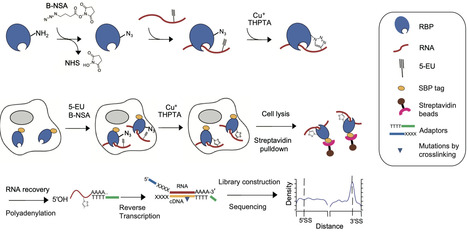
RNA binding proteins (RBPs) are multi-faceted proteins that interact with transcripts in various RNA driven processes and functions. However, in situ covalent capture techniques for screening authentic RNA substrates of RBPs remain challenging in terms of reproducibility, specificity, and sensitivity. Here, we developed CuCLIP-seq (CuAAC-Crosslinking and Immunoprecipitation Sequencing), an alternative in situ covalent capture sequencing method which utilizes CuAAC reaction to crosslink RBP-RNA between azido groups of RBPs and ethynyl groups of RNA molecules, followed by streptavidin-mediated enrichment of RNA substrates and high-throughput sequencing. We demonstrate the reliability of CuCLIP-seq by identifying substrate RNAs of several RBPs, including PTBP1, ADAR2, SRSF2, HNRNPA1, and PINX1, especially for capturing low-abundance targets. Additionally, this approach is authenticated by specifically resolving the alternative splicing transcripts targeted by PTBP1 and RNA targets by PINX1. Thus, this technique offers a sensitive and specific approach for detecting RBP substrates with high reproducibility, potential scalability, and wide applicability. Capturing RNA-protein interactions is vital for understanding cellular regulation but remains technically challenging. Here, the authors develop CuCLIP-seq, a sensitive, chemical-crosslinking method that accurately identifies diverse RNA substrates, including low-abundance targets.

|
Scooped by
mhryu@live.com
April 25, 12:55 PM
|
Protein-peptide interactions are important mediators of diverse biological processes. While deep learning has revolutionized protein structure prediction, comparative evaluation of these methods, specifically for protein-peptide complexes, remains an area of active investigation. Here, we present a systematic benchmarking of AlphaFold2 (AF2) and OpenFold3 (OF3) on a curated, non-redundant dataset of 271 protein-peptide complexes evaluated under CAPRI peptide criteria, partitioned into disordered (IDR) and structured (Non-IDR) peptide subsets. Results show that AF2 consistently outperformed OF3 across both subsets in overall success rate and proportion of high-quality models, while both methods exhibited comparable global fold prediction accuracy. We further demonstrate that AF2 exhibited memorization on a large set of protein-peptide complexes that were in its training data. Analysis of built-in and post-hoc confidence scores demonstrated that PAE-derived metrics, particularly pDockQ2, LIS, and ipSAE, provided the most reliable proxies for structural accuracy in AF2 predictions, whereas OF3's PAE distributions substantially diminished the discriminative power of its derived scores. Furthermore, we find that canonical DockQ threshold cutoffs for protein-protein complexes are not directly transferable to protein-peptide complexes, underscoring the need for method- and dataset-specific calibration. Peptide sequence composition and length were identified as potential modulators of prediction success, with glycine-rich short peptides and long receptors posing challenges to both methods. Collectively, these findings establish a peptide-specific evaluation framework and highlight the need for dataset/method-calibrated metrics to support the continued development of structure prediction tools for protein-peptide interactions.
|
 Your new post is loading...
Your new post is loading...