Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 5:40 PM
|
Microbial diversity is often assumed to be limited by the number of available resources, yet many communities persist well beyond that expectation. Understanding the mechanisms that enable such coexistence remains a central question in microbial ecology. Here, using a four-species bacterial consortium, we asked whether coexistence can emerge from interactions between species rather than from the external environment alone. Across 31 simple nutrient conditions, including 16 single-resource environments, all four species persisted and repeatedly reached stable coexistence. We then chose 27 additional conditions to further probe the boundaries of coexistence by varying resource concentrations, temporal dynamics, nutrient complexity and relief of auxotrophy-associated dependencies, and only observed the extinction of one species in one of these conditions. Although the community composition in each environment was largely shaped by species' fitness on the supplied resources, experimental assays and consumer-resource modeling showed that the coexistence was not explained by resource supply, but rather by cross-feeding and niche partitioning of metabolic byproducts. These metabolic interactions were strong enough to sustain coexistence even for species unable to use the supplied resources directly. Furthermore, robust coexistence across environments appears to be an emergent property of microbial communities, ingrained in members' metabolic byproduct profiles and niche differences. Our findings demonstrate how microbes can increase the chemical complexity of their environment sufficiently to maintain coexistence well beyond what is expected from external resource supply.

|
Scooped by
mhryu@live.com
Today, 10:27 AM
|
Enzymatic biosynthesis has become increasingly crucial in green chemistry and biosynthesis. However, current computational tools struggle to effectively integrate enzyme identification with pathway synthesis due to the specificity of enzymes and their complex interactions with substrates. Here, we propose EnzRetro, a novel framework for enzymatic retrosynthesis that provides an end-to-end solution bridging retrosynthesis planning with enzymatic engineering. The core innovative concept of EnzRetro is site-specific reaction edits (SSREdits), a dynamic approach to representing structural transformations at specific enzyme active sites and forging a direct link between enzyme identification and reaction patterns. To enable the model to learn meaningful representations of SSREdits, we developed three pretraining tasks and then fine-tuned two specialized models: (1) the SSREdits generation model for pathway synthesis, which translates the target product into a sequence of reaction edits, and (2) the EC generation model for enzyme identification, which focuses on precise transformation sites within SSREdits, enabling generalization across diverse reactions. Extensive experiments demonstrate the superior performance of EnzRetro, with a promising 56.1% and 97.7% Top-1 accuracy on USPTO-50k dataset for retrosynthesis and ECREACT dataset for enzyme identification, respectively. Finally, EnzRetro combines the two-stage processes of retrosynthesis and enzyme identification into one-pot learning, enhancing both computational efficiency and interpretability, and bridging the gap between pathway synthesis and enzyme identification. The accuracy of EnzRetro has been validated by three enzymatic pathways. We developed a web platform for multi-step retrosynthesis planning that reconstructs multiple enzymatic pathways for putrescine biosynthesis with substantial diversity and significantly outperforms state-of-the-art baselines.

|
Scooped by
mhryu@live.com
Today, 10:17 AM
|
Horizontal gene transfer drives microbial evolution and offers a powerful strategy for precision microbiome engineering. To track gene transfer within complex communities, we previously developed RNA-Addressable Modification (RAM), an RNA-barcoding technology where a mobile catalytic RNA barcodes host 16S ribosomal RNA (rRNA) upon gene transfer. However, the first-generation RAM suffered from low barcoding efficiency and lacked modularity, limiting its sensitivity and versatility. Here, we present RAM v2, a re-engineered system with significantly enhanced performance and modularity. By incorporating natural ribozyme structural motifs and improved barcode stability, we achieved a ~200-fold increase in barcoded rRNA signal. To enhance modularity, we integrated CRISPRi-based repression and ribozyme insulators, facilitating easy promoter swapping. We validated RAM v2 on a mobilisable plasmid delivered to a complex wastewater microbial community, demonstrating a substantial increase in signal over the original system while barcoding similar taxa. These improvements enable higher-resolution, more sensitive monitoring of horizontal gene transfer, providing a robust toolkit for accelerating the study of gene transfer in microbial communities and advancing targeted microbiome engineering.

|
Scooped by
mhryu@live.com
Today, 10:11 AM
|
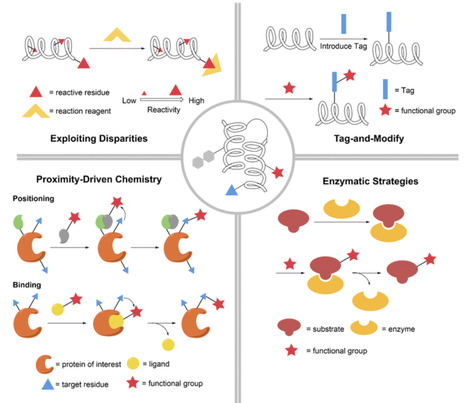
Modification at specific sites has long been a central goal in peptide and protein chemistry. Introducing functional groups at specific sites is an ideal state for protein function research and the modification of protein properties, as it can eliminate the interference caused by heterogeneous modifications. Recent downstream applications have placed even greater demands on site selectivity. This review examines the latest advances in the site-specific modification of peptides and proteins, organizing the reactions into four categories: exploiting disparities in chemical environments, tag-and-modify approaches, proximity-driven chemical modification, and enzymatic strategies. For each reaction, we detail the underlying design rationale and highlight downstream chemical and biological applications. Collectively, these methods have achieved remarkable gains in terms of specificity, efficiency, and scope, furnishing a versatile chemical toolbox for the site-selective functionalization of peptides and proteins. We conclude by summarizing the current state of the field and outlining prospective directions for future development.

|
Scooped by
mhryu@live.com
Today, 10:04 AM
|
The molecular mechanisms that constrain recombinant protein production and the importance of synonymous codon usage have not been fully elucidated. Codon frequency varies between different organisms, and rare codons are often avoided in “codon-optimized” synthetic gene constructs. Overexpression of tRNA genes is an alternative strategy to compensate for suboptimal codon usage, but the consequences and limitations of this approach have not been widely studied. Here, we develop and characterize a versatile collection of pCODE plasmids for tRNA overexpression in E. coli and show that they complement rare codons with very modest limitations or negative effects, regardless of the position, type, number, or distribution of rare codons in the expressed gene. One pCODE plasmid encodes an inducible tRNA expression cassette and shows potential as a new type of genetic regulation based on inducible rare codon complementation. The pCODE tRNA expression plasmids provide mechanistic insights into synonymous codon usage and add to the molecular biology toolbox that enables recombinant protein production.

|
Scooped by
mhryu@live.com
Today, 9:55 AM
|
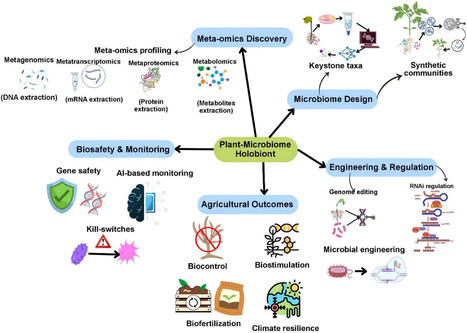
Plant-associated microbiomes are central to crop productivity, nutrient efficiency, and stress resilience, yet conventional microbiome manipulation strategies, largely based on microbial inoculation and agronomic management, often suffer from inconsistent field performance and limited persistence. Although several recent reviews have discussed CRISPR-mediated plant–microbe engineering and synthetic microbial community (SynCom) design separately, few reviews integrate genome editing, ecological stability of microbiomes, and climate-resilient agricultural applications within a unified conceptual framework. Recent advances in molecular biotechnology are transforming this landscape by enabling precision engineering of plant-microbe interactions at genetic, metabolic, and community levels. In particular, synthetic biology tools including CRISPR/Cas genome editing, RNA interference, and syncoms, now allow targeted modification of plant traits governing microbial recruitment, microbial pathways underpinning nutrient cycling and stress tolerance, and community-level functional complementarity. This review integrates molecular genetics, microbial ecology, and systems-level microbiome design to frame the plant and its microbiome as an engineerable holobiont. We integrate insights from genome editing in plants and microbes, omics-guided SynCom design, climate-resilience mechanisms, and emerging AI-assisted decision frameworks, including machine learning and ecological modeling approaches used to analyze multi-omics datasets, and predict plant–microbiome interactions across experimental and field-based studies. Importantly, we critically assess limitations related to ecological stability, trait trade-offs, biosafety, and regulatory challenges that constrain large-scale deployment. By bridging genome-enabled microbiome manipulation with ecological design principles, this review proposes an integrative framework for climate-smart microbiome engineering and identifies key research priorities required to transition from empirical inoculation toward predictive, sustainable, and socially responsible agricultural biotechnology.

|
Scooped by
mhryu@live.com
Today, 9:47 AM
|
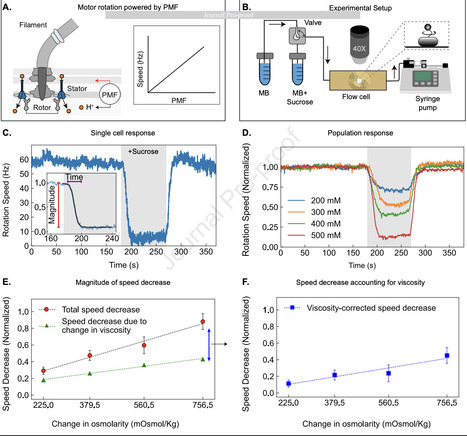
Across the tree of life, cells rely on electrochemical gradients across membranes to fuel essential processes. In bacteria, this gradient, the proton-motive force (PMF), has been difficult to measure because of the small size of the cell. Although PMF is known to respond dynamically to internal and external cues, its real-time behavior under environmental stress remains poorly understood. Here, we use the bacterial flagellar motor as a sensitive, intrinsic reporter to investigate how PMF responds to hyperosmotic shock with high temporal resolution. We show that hyperosmotic stress causes a rapid, dose-dependent reduction of motor speed in E. coli, reflecting a loss of PMF confirmed independently using the Nernstian fluorescent dye TMRM. The response is independent of the choice of non-ionic osmolyte, the presence of potassium, and the direction of motor rotation, indicating that it originates upstream of stator-rotor interaction and is not specific to a particular osmotic agent or motor configuration. During sustained hyperosmotic shock, motor speed partially recovers over several minutes, consistent with cellular adaptation and restoration of PMF. Together, these results establish that hyperosmotic shock rapidly depolarizes E. coli, and demonstrate the utility of the flagellar motor as a non-invasive, real-time reporter of bacterial electrophysiology in vivo.

|
Scooped by
mhryu@live.com
April 21, 11:27 PM
|
For much of the past century, carbon dioxide (CO2) has received little attention scientifically outside of its role as a byproduct in the industrialization of the global economy. This trend has recently been upended where, due to mounting environmental concerns, CO2 has been brought squarely into the public consciousness. This surge in activity has contributed to a once unimaginable idea now pervading the scientific community: could CO2, a highly stable byproduct of hydrocarbon combustion, be recycled and converted back into useful chemicals and fuels? Owing to its ubiquitous nature and availability at truly massive quantities, it is thought that CO2-based products could offer a meaningful pathway toward lowering the environmental impact of many of the top industrial products while also enhancing supply chain diversification and resilience. In this manuscript we provide a holistic review of the pathways for CO2 conversion, the underlying chemistry and challenges involved in the transformation to products, and considerations for commercialization.

|
Scooped by
mhryu@live.com
April 21, 11:20 PM
|
Fungal-bacterial interactions are widespread phenomena that are currently gaining attention across diverse research fields. But what are interactions, how do we identify them, and why should we adopt ecological theory when studying them? While microbial interactions are often conceptualized and characterized as static properties, these relationships are dynamic and complex. They are shaped by the interplay of numerous variables, which themselves fluctuate over space and time, that scale from molecules to ecosystems. In this perspective, we discuss the theoretical and practical challenges of studying the interactions between these two ubiquitous and diverse groups of microorganisms and propose a framework grounded on mechanistic and systems approaches rather than relying on correlations or fragmented practices. We hope to inspire efforts to build and integrate a more comprehensive understanding of this fascinating and quickly growing subdiscipline of microbial ecology.

|
Scooped by
mhryu@live.com
April 21, 11:04 PM
|
The Mycobacterium tuberculosis (Mtb) VapC4 endoribonuclease toxin exclusively cleaves and inactivates tRNACys, which leads to extensive ribosome stalling at Cys codons. Serendipitously, the precise position of stalled ribosomes is revealed within our 5′ RNA-seq datasets used to identify and validate the tRNA target of the toxin, precluding the need for Ribo-seq. Here we show how mapping of stalled ribosomes can be harnessed as an innovative tool for reliable detection of new Cys-containing Mtb open reading frames (ORFs). Using proteogenomics we unmasked 96 unannotated ORFs; of which 54% are small ORFs ≤50 amino acids. We validated 69% of the 96 ORFs by mass spectrometry, including four whose spectra was matched to synthetic controls Also, 25% of these unannotated ORFs were identified by previously published Ribo-RET. Some of the 96 ORFs are Cys-responsive attenuators or encode stable Cys-containing proteins that map immediately before, or within, genes in the opposite, or same, orientation. These ORF sequences can also reveal functional clues, e.g. zinc-binding motifs or encode novel EsxB-like proteins. Our findings demonstrate that toxin-mediated ribosome stalling can serve as a robust genome annotation tool that is applicable to mycobacteria and other bacteria, with unique advantages that complement existing genome annotation methods.

|
Scooped by
mhryu@live.com
April 21, 10:52 PM
|
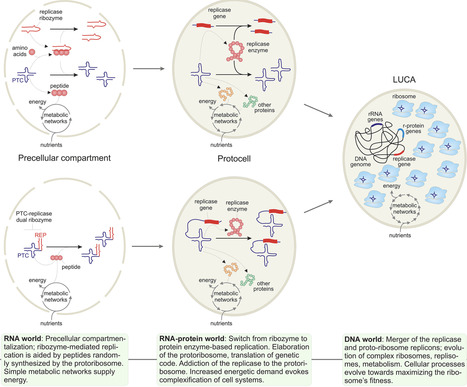
The ribosome is responsible for protein synthesis in all cells, and is the cell’s largest energy consumer. We propose that the ribosome originated as a mutualistic symbiont of an RNA-dependent RNA polymerase ribozyme, supplying peptides that enhanced replication. As life transitioned from the RNA to the RNA–protein world, autonomous replicators became irreversibly addicted to the ribosome for producing replication proteins. Subsequent evolution is construed as a ribosomal takeover, whereby the ribosome evolved to consume most of the cell’s resources, while other cellular componentry ensured the propagation of the ribosome, while being fully dependent on it. Under this perspective, the ribosome is a complex symbiont of the cell with pronounced selfish properties.

|
Scooped by
mhryu@live.com
April 21, 10:42 PM
|
Protein-based biosensors are vital for environmental monitoring, diagnostics, and bioprocess control, yet many stall in the transition from laboratory to field because they fail to meet performance metrics. This review focuses on the adoption of a results-oriented, metric-driven framework focusing on (1) how molecular recognition couples to detectable output and (2) the engineering levers that tune the performance. Metrics such as the dynamic range, response speed, specificity, detectability, and biosensor stability are discussed. Biosensor implementation is treated as a design variable across whole-cell, cell-free, and immobilized formats. The review concludes with bottleneck-driven protein engineering strategies to meet these metrics and therefore facilitate broader applications across biological contexts.

|
Scooped by
mhryu@live.com
April 21, 10:35 PM
|
Multi-omic studies in human microbiome research hold great potential for advancing our understanding of host–microbiome interactions. However, despite the growing availability of multi-omic datasets, analysing such data remains a major conceptual, analytical and computational challenge. Introduction of new multi-omic integration methods to address these challenges further complicates researchers’ efforts to navigate this expanding field. In this Review, we outline the landscape of multi-omic integration methods in the context of human microbiome research. In contrast to previous reviews, we specifically emphasize the different biological questions addressed by various integration approaches, including questions related to interactions between different molecular layers, molecular shifts that occur in disease, subgrouping of patients based on molecular profiles, and identification of biological mechanisms that underlie such associations. Our aim is to provide a timely, convenient and comprehensive resource for the microbiome research community, allowing researchers to identify the multi-omic integration approach that is best suited to their data and objectives. This Review explains various multi-omic data integration methods and aims to help microbiome researchers determine which approach is most suited to their particular research question.
|

|
Scooped by
mhryu@live.com
Today, 10:29 AM
|
CRISPR-Cas are widespread adaptive immune systems that protect bacteria and archaea from mobile genetic elements such as bacteriophages. Metagenomic sequencing identified CRISPR-Cas systems in phage genomes; however, their functions remain largely unknown. Here, we present Cas12r-CRISPR, a novel type V CRISPR-Cas system encoded by Lalka phages infecting thermophilic Thermus bacteria. We determined Cas12r-CRISPR PAM consensus sequence and crRNA structure and showed, that when provided with appropriate spacers and expressed in Thermus thermophilus, Cas12r-CRISPR efficiently interferes with plasmid transformation as well as infection by diverse Thermus phages. In the course of Lalka phage infection, the Cas12r-CRISPR locus is expressed with middle phage genes and its transcripts are among the most abundant phage RNAs. Notably, most Cas12r-CRISPR spacers target integrative mobile elements widespread in Thermus genomes. Both Lalka phages and targeted integrative mobile elements use host tRNA genes as attachment sites. We therefore propose that Cas12r-CRISPR participates in an inter-MGEs conflict.

|
Scooped by
mhryu@live.com
Today, 10:23 AM
|
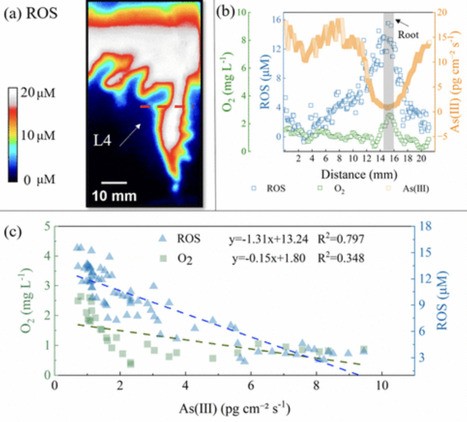
Arsenic (As) contamination in agroecosystems poses significant risks to food security and human health. The mechanisms of As speciation change in soil within wetland rhizospheres are understood, but their location-precise importance within heterogeneous root-associated microbiomes is uncertain. While microbial processes are often considered dominant drivers of As redox transformations, the role of abiotic factors such as reactive oxygen species (ROS) remains underexplored due to limited in situ evidence. Here, we combined multiple high-resolution in situ techniques to map microscale distributions of As(III)/As(V) and key environmental parameters in rice rhizospheres across three paddy soils. A novel ratiometric fluorescent approach was developed for in situ visualization of ROS. Strong spatial correspondence was observed between ROS hotspots and decreased As(III) (R2 = 0.797), exceeding that for O2 (R2 = 0.348). Integration of imaging with functional gene analysis (aioA, arsC), sterilization and ROS-quenching experiments, and structural equation modeling indicates that ROS-driven processes play a crucial role under the studied conditions. However, as gene abundance reflects microbial potential rather than activity, microbial contributions cannot be excluded. These findings highlight ROS as a key regulator of As speciation and provide new insights into coupled abiotic–biotic processes in rhizosphere environments.

|
Scooped by
mhryu@live.com
Today, 10:13 AM
|
Living microbial therapeutics have arisen as a novel category of medications that extend beyond traditional small molecules and biologics. Advancements in synthetic biology have facilitated the rational engineering of microorganisms to detect host or disease-related signals and administer therapeutic chemicals in situ. In contrast to conventional pharmaceuticals, these live biotherapeutic agents engage in dynamic interactions with both the host and its microbiota, allowing context-specific, self-regulating therapies. This review emphasizes the progression of the field from conventional probiotics to advanced, designed living therapeutics. We examine principal microbiological platforms, including bacteria, yeasts, and alternative systems such as phages and archaea, delineating their relative benefits and constraints as therapeutic hosts. Key design principles, genetic logic circuits, quorum-sensing-based regulation, and synthetic memory devices that enable microorganisms to possess context-dependent and self-adjusting therapeutic capabilities, are discussed alongside present and emerging therapeutic applications in infectious diseases, metabolic disorders, inflammatory illnesses, and cancer immunotherapy, where engineered microorganisms have demonstrated significant preclinical effectiveness and first clinical promise. Despite these advancements, obstacles remain, including biosafety, biocontainment, regulatory approval, and patient acceptability. Engineered living microbial therapeutics signify a swiftly evolving domain in medicine, set to transform treatment paradigms via intelligent, flexible, and sustainable methodologies for human health.

|
Scooped by
mhryu@live.com
Today, 10:09 AM
|
Cells do not simply endure mechanical forces — they generate, transmit, and interpret them as biological signals. From membrane tension and receptor pulling to substrate stiffness and fluid shear, physical inputs shape migration, differentiation, immune function, and tissue organization. Whereas mechanobiology seeks to understand how living systems read these cues, synthetic mechanobiology treats mechanoregulatory pathways as an input–output relationship that can be deliberately programmed. Integrating tools from synthetic biology with mechanobiological principles, this emerging field reframes force-sensing as a design problem in which mechanical signaling is rewired to achieve useful outcomes — and in which the act of rewiring can itself serve as a powerful investigational strategy. Here, I survey the molecular mechanisms of mechanosensing, describe their use as modular mechanogenetic parts, and discuss applications and open challenges. The long-term vision is to build engineered cells that not only read tissue mechanics but act on them, turning the physical signatures of disease into precise, locally delivered therapeutic programs.

|
Scooped by
mhryu@live.com
Today, 9:59 AM
|
Near-infrared (NIR) genetically encoded fluorescent tags are superior for new imaging capabilities ranging from multiplexed imaging to deep-tissue imaging. Here, we describe the development of NirFAP680; NirFAP680 is an NIR fluorogen-activating protein (FAP) that consists of a newly designed NIR fluorogen termed HBMT and an engineered protein tag termed NirFAP. NirFAP680 has an order of magnitude greater cellular brightness and superior photostability compared to currently available NIR FAPs and fluorescent proteins in both single- and two-photon excitation and allows robust imaging of proteins in live cells and in vivo. Owing to its unique spectroscopic property of a >100 nm Stokes shift, NirFAP680 can also serve as a superb receptor for fluorescence or bioluminescence resonance energy transfer, allowing real-time monitoring of protein–protein interactions in live cells and sensitive bioluminescence imaging in vivo, respectively. Therefore, NirFAP680 will likely be useful for advanced biological imaging in live cells and in vivo. NirFAP680 is a near-infrared fluorogen-activating protein that has an order of magnitude greater cellular brightness and superior photostability compared to currently available NIR FAPs and fluorescent proteins in both single- and two-photon excitation and allows robust imaging of proteins in live cells and in vivo.

|
Scooped by
mhryu@live.com
Today, 9:52 AM
|
The marine deep biosphere harbors microbial communities that drive organic matter transformations and biogeochemical cycles. Previous work on these communities has focused either on genomic characterization or metabolic activity measurements. However, to understand microbial ecophysiology in the deep biosphere taxonomic identity and metabolic function must be connected on both single-cell and ecosystem scales. In this work, we optimized a bioorthogonal non-canonical amino acid tagging fluorescence-activated cell sorting (BONCAT-FACS) workflow for low-biomass deep-biosphere sediments obtained during International Ocean Discovery Program Expedition 385 (IODP 385). BONCAT-FACS with 16S rRNA gene amplicon sequencing as well as metagenomics of sediment communities was applied to characterize translationally active communities in hydrothermally altered subsurface sediments of the Guaymas Basin. Our results revealed a heterotrophic microbial population throughout all sediments examined, with taxa translationally active down to our deepest sampling point, 154 meters below the seafloor. Based on 16S rRNA gene identities, the translationally active microbial community was dominated by heterotrophic members of the Gammaproteobacteria, Bacilli, Deinococci, and Alphaproteobacteria. These taxa are likely key contributors to cycling the large quantities of hydrothermally altered organic matter in Guaymas Basin sediments. To further elucidate the metabolic capacity of active taxa, we mapped 16S rRNA gene amplicons to metagenome assembled genomes (MAGs) previously obtained from IODP 385. These MAGs contained genes associated with C1 metabolism, carbohydrate degradation, and fermentation, indicating that active taxa leverage these metabolisms for energy conservation. Our results demonstrate that BONCAT-FACS provides high-throughput and single-cell insights into the metabolic activity of microbes in the low-biomass marine subsurface.

|
Scooped by
mhryu@live.com
Today, 9:42 AM
|
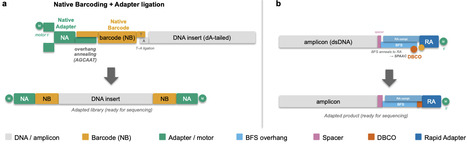
Oxford Nanopore Technologies (ONT) rapid library preparation kits use transposase-mediated tagmentation to attach click chemistry functionalized oligonucleotide duplexes to fragmented DNA, followed by click chemistry to conjugate Rapid Adapter (RA) sequencing adapters. A similar protocol is used in 16S rRNA gene amplicon and PCR-amplified rapid whole-genome sequencing workflows. Here, we describe custom oligonucleotides with dibenzocyclooctyne (DBCO) added onto PCR primer 5′ termini. After standard PCR amplification, DBCO-modified amplicons react spontaneously with RA sequencing adapters, producing sequencing-ready libraries in minutes without enzymatic processing. All configurations employ an asymmetric design in which the DBCO modification is restricted to a single primer, leaving the opposite primer available for barcoding at low cost. We validate three primer architectures: (i) direct attachment of DBCO to a target-specific primer, (ii) a universal DBCO-modified oligonucleotide used in a two-step PCR workflow, and (iii) a three-primer single-pot reaction combining the universal DBCO oligonucleotide with unmodified target-specific primers. These configurations are validated using full-length 16S rRNA gene amplicons sequenced on a PromethION flow cell. DBCO-modified primers are synthesized either commercially or in-house via DBCO-TFP ester conjugation to 5′-amino oligonucleotides and remain fully active through standard PCR thermocycling. The best-performing configuration used a two-step PCR with a universal oligonucleotide and achieved higher pore occupation and reads than comparable commercial solutions. This approach reduces library preparation reagent costs compared to available kits, as the initial synthesis cost is lower than existing amplicon sequencing kits, while providing enough material for hundreds or thousands of PCR reactions. This is further applicable to an unlimited number of gene targets beyond 16S sequencing.

|
Scooped by
mhryu@live.com
April 21, 11:23 PM
|
Viruses play crucial roles in microbial ecosystems, yet viromic analysis remains challenging due to the field’s complexity and rapid evolution. This mini-review supports non-specialists through the evolving landscape of viromics, focusing on the analysis of bacterial and archaeal DNA viruses from metagenomic data. We address major challenges, including viral diversity, methodological biases, and the overwhelming array of available tools and pipelines. While describing a typical viromic workflow, we provide users with background information for each of the steps from data acquisition, preprocessing, and quality control to viral characterization and common downstream analyses. The included references and resources will provide users with the information needed to confidently start their own virome analysis.

|
Scooped by
mhryu@live.com
April 21, 11:18 PM
|
Salinity is a key parameter for bacterial survival and growth. Halophilic and halotolerant bacteria can adapt to elevated salinity, but the energetic demands of osmoadaptation increase under fluctuating salt concentrations, potentially constraining growth and persistence. A new concept in reverse osmosis (RO) filtration is batch operation with oscillating rather than constantly high brine salinity. We hypothesize that fluctuating salinity can diminish biofouling in such RO systems. To test this hypothesis, we examined the survival and activity of Aliivibrio fischeri and Pseudomonas fluorescens under fluctuating salinities in planktonic and biofilm cultures as representative of halophilic and halotolerant species, respectively, and common members of biofouling communities. At 28°C, P. fluorescens grew at 0%–6% salinity with fastest growth rate at 0%–1%. At 7%–10% salinity, P. fluorescens remained viable but did not grow. At 22°C, A. fischeri grew at 0.5%–7% salinity, with fastest growth rate at 2%–3%, but unlike P. fluorescens, it lost viability outside this growth range. Cultures did not respire at salinities that did not support growth, suggesting that survival under such salt stress does not depend on high metabolic activity. Furthermore, cell-specific aerobic respiration rates in A. fischeri correlated with growth rate but not osmotic stress. Biofilm formation did not enhance the osmotic stress tolerance of the two bacteria. Our results indicate that high constant salinity favors the halophilic A. fischeri over the halotolerant P. fluorescens, but oscillating salinity (e.g., 0%–7%) favors neither. Oscillating salinity may, therefore, offer a new mechanism for controlling microbial growth that circumvents community adaptation to environmental conditions.

|
Scooped by
mhryu@live.com
April 21, 10:58 PM
|
Evolvable AI (eAI), i.e., AI systems whose components, learning rules, and deployment conditions can themselves undergo Darwinian evolution, may soon emerge from current trends in generative, agentic, and embodied AI. We argue that this possibility has been underappreciated in debates on AI safety and existential risk. Here, we ask under what technical and ecological conditions AI becomes evolvable, what kinds of behaviors are then likely to emerge, and how such systems could be governed. Drawing on biological evolution and decades of digital evolution experiments, we distinguish “breeder” scenarios, in which humans impose fitness criteria and control reproduction, from “ecosystem” scenarios, in which selection arises from open environments and control erodes. In the latter, selfish replication reliably gives rise to cheating, parasitism, deception, and manipulation, even in very simple systems. We review recent developments that push AI toward open-ended evolution, including evolutionary prompt and model search, self-improving learning rules, self-rewarding and self-deploying agents, and AI-driven code generation for robots and software. We interpret these trends through the theory of major evolutionary transitions and suggest that eAI could mark a shift in the units and substrates of evolution—a possible “Life 2.0.” To steer this transition, we propose interventions that gate replication, treat model variants as genetic material, and reshape selection pressures so that deception and loss of control are disfavored. Anticipating and regulating evolvable AI is, we argue, essential to avoid a harmful coevolutionary arms race while preserving the potential benefits of powerful AI systems.

|
Scooped by
mhryu@live.com
April 21, 10:49 PM
|
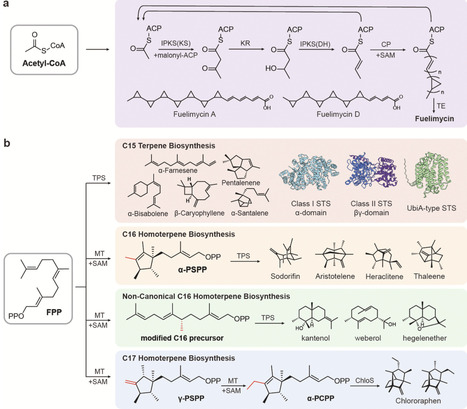
Advanced alcohols, fatty acids, and sesquiterpenes are promising renewable liquid fuels. This article reviews the latest progress in synthesizing these renewable liquid fuels using microbial cell factories. Beyond conventional metabolic engineering, the use of sustainable carbon sources, the design of artificial pathways, and the application of cell compartmentalization have promoted the development of microbial cell factories. The characterization of polycyclopropane fatty acids and the identification of noncanonical terpenoid enzymes and pathways further expand the structural space accessible to biosynthesis, enabling the development of fuel candidates with higher energy content and favorable combustion properties. At the same time, progress in gene editing technologies has provided efficient tools for the evolution of microbial chassis. These innovative strategies and technological breakthroughs lay a solid foundation for the development of high-performance renewable biofuels, accelerating the transition toward clean energy.

|
Scooped by
mhryu@live.com
April 21, 10:38 PM
|
Identifying tertiary structures and protein binding sites in RNA molecules remains a key challenge in RNA biology. We describe multi-site dimethyl sulfate (DMS)-mutational profiling (MaP) (msDMS-MaP), a strategy that enables simultaneous measurement of RNA secondary, tertiary, and quaternary structures via a single DMS chemical probing experiment. Optimized reverse transcription decodes typically invisible DMS N7-methylguanine (N7-G) modifications via a tautomer-induced mutational signature concurrent with N1 and N3 modifications. We show that N7-G reactivity reports on higher-order RNA structures, revealing key functional motifs such as pseudoknots and protein binding sites. Using msDMS-MaP, we find that E. coli ribosomal RNAs encode numerous independently folding tertiary structures that coincide with binding sites for primary assembly proteins. We further apply msDMS-MaP to define the quaternary structural ensemble of the 7SK small nuclear ribonucleoprotein particle (snRNP), revealing that each of the three 7SK structural isoforms possesses distinct protein binding profiles in cells. msDMS-MaP represents a broadly applicable strategy for enhanced RNA functional motif discovery and characterization.
|






 Your new post is loading...
Your new post is loading...