 Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 12:50 AM
|
Combinatorial mutagenesis is essential for exploring protein sequence-function landscapes in engineering applications. However, while large-scale machine learning benchmarks exist for protein function prediction, they are primarily limited to single-mutant libraries, leaving a critical gap for combinatorial mutagenesis. Here we introduce CombinGym, a benchmarking platform featuring 14 curated combinatorial mutagenesis datasets spanning 9 proteins with diverse functional properties including binding affinity, fluorescence, and enzymatic activities. We evaluated nine machine learning algorithms from five methodological categories (alignment-based, protein language, structure-based, sequence-label, and substitution-based) across multiple prediction tasks, assessing both zero-shot and supervised learning performance using Spearman's ρ and Normalized Discounted Cumulative Gain metrics. Our analysis reveals the substantial impact of measurement noise and data processing strategies on model performance. By implementing hierarchical dataset splits (0-vs-rest, 1-vs-rest, 2-vs-rest, and 3-vs-rest scenarios), we demonstrate the value of lower-order mutation data for empowering machine learning models to predict higher-order mutant properties. We validated this capacity through both in silico simulation (improving fluorescence brightness of an oxygen-independent fluorescent protein) and experimental validation (engineering enzyme substrate specificity), achieving a substantial increase in specific activity. All datasets, benchmarks, and metrics are available through an interactive website (https://www.combingym.org), facilitating collaborative dataset expansion and model development through integration with automated biofoundry platforms.

|
Scooped by
mhryu@live.com
Today, 12:43 AM
|
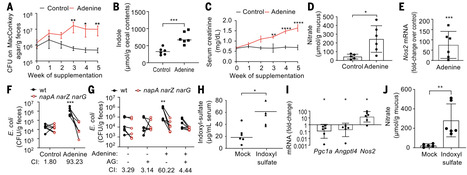
Chronic kidney disease (CKD) is linked to an elevated fecal abundance of Enterobacteriaceae, but the ecological drivers of this shift and its impact on disease progression remain unclear. The uremic toxin indoxyl sulfate is produced from microbiota-derived indole in the liver. Here, we found that in mice with adenine-induced CKD, impaired clearance of indoxyl sulfate elevated mucosal expression of the gene encoding inducible nitric oxide synthase (iNOS). The resulting rise in luminal nitrate levels promoted E. coli growth by means of nitrate respiration. Fecal microbiota from CKD patients generated more indole than feces of healthy controls during anaerobic culture, but only in the presence of nitrate. Nitrate enhanced indole production by E. coli, thereby worsening renal pathology in CKD mice, which was mitigated by iNOS inhibition.

|
Scooped by
mhryu@live.com
March 25, 11:49 PM
|
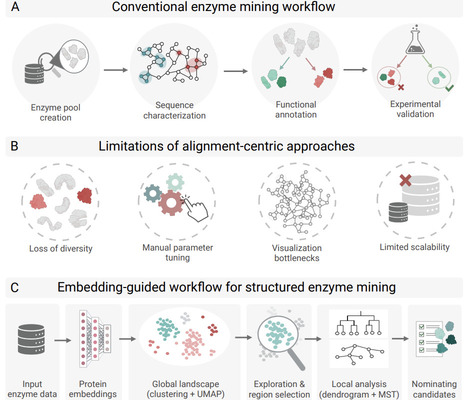
The rapid expansion of protein sequence databases continues to outpace functional characterization, limiting efficient enzyme discovery in large, heterogeneous, and sparsely annotated sequence spaces. Here, we present an embedding-guided framework for structured navigation of enzyme sequence space, implemented in SelectZyme. The workflow integrates protein language model embeddings with dimensionality reduction, hierarchical clustering, connectivity reconstruction, and quantitative dendrogram analysis to enable exploration without reliance on fixed sequence identity thresholds or predefined functional annotations. Across distinct case studies, we demonstrate that embedding-defined neighborhoods preserve fold-level coherence even when sequence identity resides in the twilight zone and that biologically meaningful functional organization can emerge under fully unsupervised conditions. In a large multi-family PETase landscape exceeding 100,000 sequences, the framework supports scalable, anchor-guided prioritization under sparse labeling and process-motivated constraints. By unifying latent representations with connectivity-aware and hierarchical interpretation, this approach reframes enzyme mining from threshold-driven similarity filtering into structured exploration, providing a scalable methodological foundation for hypothesisdriven biocatalyst discovery and reasonable starting points for downstream protein engineering.

|
Scooped by
mhryu@live.com
March 25, 11:03 PM
|
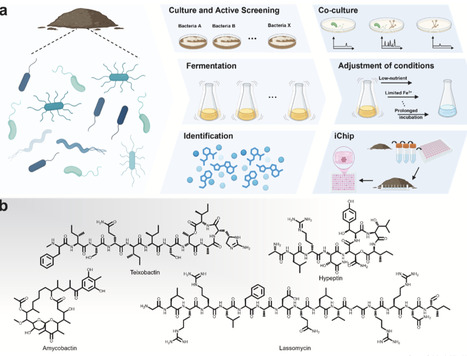
The discovery of antibiotics transformed modern medicine and extended the average human lifespan by decades. However, the initial golden era of antibiotic discovery has significantly waned, a decline aggravated by the relentless evolution of drug resistance among human pathogens, ultimately driving the current global antimicrobial resistance (AMR) crisis. The repeated rediscovery of known compounds from conventional soil-derived microbes underscores the urgent need for new strategies and ecological frontiers. Here, we review emerging directions in antibiotic discovery that collectively address this innovation gap. Refined cultivation techniques such as co-culture and iChip have reactivated rare taxa and yielded novel scaffolds like teixobactin. Multi-omics and synthetic biology approaches now enable culture-independent access to cryptic biosynthetic gene clusters from the vast uncultivated microbial majority. Most recently, artificial intelligence (AI) has expanded the search frontier to neglected taxa such as archaea and even evolutionary timeframes through paleoproteome mining. Overall, these innovations signal a new era of intelligent, data-driven antibiotic discovery. The integration of ecology, omics, synthetic biology, and AI provides a sustainable framework to replenish the antibiotic pipeline and mitigate the growing threat of AMR.

|
Scooped by
mhryu@live.com
March 25, 10:27 PM
|
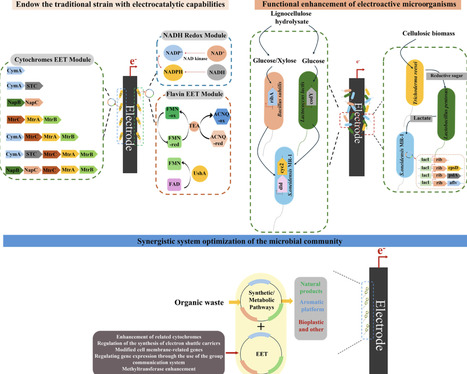
Bioelectrocatalytic conversion of organic waste uses renewable electricity to convert organic waste into high-value chemicals and fuels under mild conditions. It is regarded as a strategic link connecting waste refinement and the green economy, providing a sustainable approach for the high-value utilization of waste resources. This article systematically reviews: (1) the basic principles and reaction mechanisms of bioelectrocatalytic conversion of organic waste; (2) the development history and key breakthroughs of organic waste biological electrochemical conversion; (3) microbial engineering strategies based on genetic engineering and synthetic biology; (4) the directions for process and reactor optimization. This review uniquely integrates advancements in biological innovation and process engineering, offering a holistic perspective on synergistic optimization across molecular, microbial, and reactor scales. By explicitly bridging microbial carbon/electron flux engineering with bioreactor design and process coupling, it aims to establish a comprehensive roadmap for accelerating the industrial-scale application of bioelectrocatalytic technologies in organic waste treatment.

|
Scooped by
mhryu@live.com
March 25, 4:47 PM
|
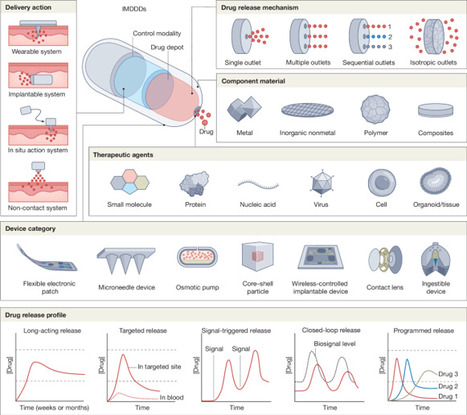
Advances at the intersection of biotechnology, artificial intelligence, electronics and materials science are reshaping how drugs can be delivered inside the body. Intelligent and miniaturized drug delivery devices (IMDDDs) leverage these technologies to achieve precise pharmacokinetics, targeted distribution and programmable release while minimizing toxicity and improving patient adherence. Unlike conventional approaches, IMDDDs can incorporate real-time sensing and adaptive control, enabling drug administration that is more precise and more responsive to dynamic physiological conditions. In this Review, we outline key categories and design principles, highlight artificial intelligence technologies for augmenting performance, discuss potential clinical applications across cancer, diabetes, cardiovascular disease, vaccination and beyond, and examine translation challenges and opportunities. By uniting engineering innovation with medical need, IMDDDs exemplify the next generation of drug delivery technologies. Intelligent and miniaturized drug delivery devices leveraging advances in biotechnology, artificial intelligence, electronics and materials science enable treatments with increased precision and responsiveness, with applications in cancer, diabetes, cardiovascular disease and other diseases.

|
Scooped by
mhryu@live.com
March 25, 4:17 PM
|
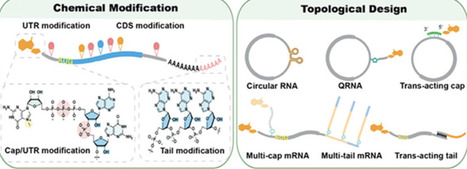
Messenger RNA (mRNA) has rapidly emerged as a transformative therapeutic modality, exemplified by its growing applications in infectious diseases, oncology, and genetic disorders. The chemical programmability of mRNA allows researchers to modulate its function by introducing synthetic modifications across the molecule─from the cap structure, untranslated regions (UTRs), coding sequence (CDS) to poly(A) tail and from base, backbone to ribose sugars. Beyond sequence-level design, recent advances have introduced a new dimension of control: topological engineering. Circular RNAs, branched structures, and synthetic lariat architectures are reshaping how we approach RNA stability, immunogenicity, and translation. This review surveys recent advances in the chemical and topological engineering of mRNA, emphasizing four key areas: (1) enzymatic, chemical, and hybrid methodologies that expand the repertoire of accessible mRNA modifications; (2) synthesis strategies for linear, circular, and branched mRNA topologies; (3) structure–activity relationships governing translation efficiency, decay, and immune activation; and (4) implications for next-generation mRNA-based therapeutics. By integrating chemical synthesis, synthetic biology, and RNA structural design, researchers are beginning to unlock the full therapeutic potential of engineered mRNA molecules.

|
Scooped by
mhryu@live.com
March 25, 3:52 PM
|
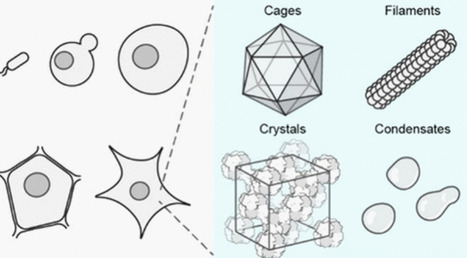
In cellulo protein assemblies, spanning protein cages, filaments, crystals, and biomolecular condensates, provide cells with modular strategies to package, organize, and regulate biomolecules and biochemical reactions. Their genetic encodability, structural diversity, and tunable material properties have also made them attractive biomaterials, where in cellulo fabrication has underpinned their precise assembly and broad applicability. This review surveys major classes of natural and engineered assemblies fabricated in cellulo, with particular emphasis on how their structure, chemistry, and material state shape functions. We compare diverse cellular reactors and outline how intracellular milieu, post-translational modifications, and folding/assembly machinery influence assembly outcomes. Engineering strategies for modifying the assemblies are summarized and mapped onto broad applications across fundamental biology, biomedicine, and nonbiological fields. Lastly, we highlight existing opportunities for engineering and designing in cellulo protein assemblies. Through this review, we hope to give a comprehensive overview of this exciting and rapidly growing field and share our perspective on the possible future directions.

|
Scooped by
mhryu@live.com
March 25, 3:33 PM
|
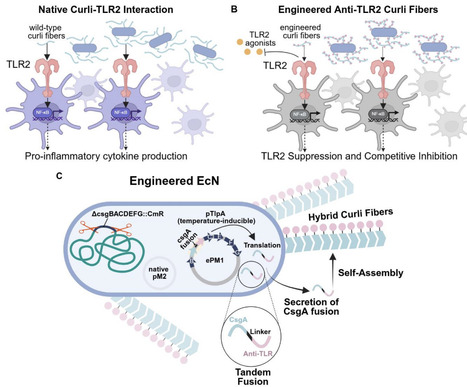
Curli fibers produced by E. coli are functional amyloids that activate Toll-like receptor 2 (TLR2), initiating innate immune responses at mucosal surfaces. While microbiome-derived curli contribute to host-microbe interactions, their intrinsic immunostimulatory activity limits their utility as programmable scaffolds for engineered probiotic systems, and dysregulated TLR2 activation has been linked to inflammatory bowel disease, systemic lupus erythematosus, neurodegeneration, and sepsis. Here, we engineered E. coli Nissle 1917 to produce modified curli fibers designed to inhibit TLR2 through two mechanistically distinct strategies: steric shielding via silk-elastin-like protein sequences, and direct receptor antagonism via a known TLR2 antagonist, staphylococcal superantigen-like protein 3 (SSL3). Both engineered variants assembled into structurally intact amyloid fibers and exhibited significantly reduced intrinsic TLR2-dependent NF-κB activation in reporter cells. In competitive inhibition assays against structurally diverse TLR2 agonists, the SSL3 fusion achieved near-complete inhibition maintained under rising agonist load, while steric shielding provided moderate, agonist-class-dependent inhibition. In primary human monocyte-derived dendritic cells, the SSL3 fusion robustly attenuated IL-8 secretion and transcriptional induction of IL-8, IL-6, and IL-1β, whereas steric shielding produced only partial attenuation that did not translate to broad inflammatory suppression. These results establish engineered curli as a tunable platform for receptor-specific modulation of innate immune signaling and highlight the broader potential of modular microbial amyloids as programmable interfaces for engineering host-microbe interactions at mucosal surfaces.

|
Scooped by
mhryu@live.com
March 25, 3:17 PM
|
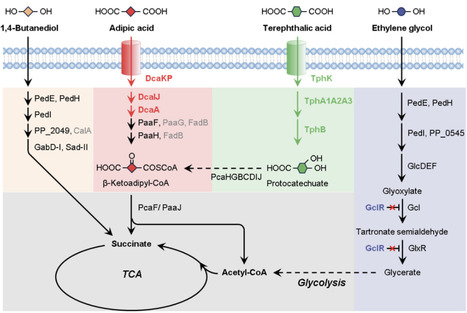
Current mechanical and chemical recycling strategies address less than 10% of global plastic waste, necessitating alternative valorization routes. Biological upcycling via enzymatic depolymerization combined with microbial conversion of the resulting monomers offers a promising pathway to transform mixed plastic waste into valuable alternatives. Here, we employed a single engineered Pseudomonas putida KT2440 for simultaneous co-utilization of five plastic monomers including ethylene glycol, terephthalic acid, adipic acid, 1,4-butanediol, and L-lactic acid, which can be derived from enzymatic hydrolysis of polyethylene terephthalate (PET), polybutylene adipate-co-terephthalate (PBAT), polyester-polyurethanes (PUs), and polylactic acid (PLA). Continuous fermentation over 21 days with alternating mixed-monomer feeds achieved steady state growth and complete substrate depletion, yielding adaptive mutations that informed iterative strain improvement. Further engineering enabled the biosynthesis of (R)-3-hydroxybutyrate (R-3HB), and 0.70 g/L R-3HB was produced directly from enzymatic hydrolysates of blended PET, PBAT, and TPU. These results establish a viable bio-based approach for upcycling realistic mixed plastics into value-added bioproducts.

|
Scooped by
mhryu@live.com
March 25, 3:03 PM
|
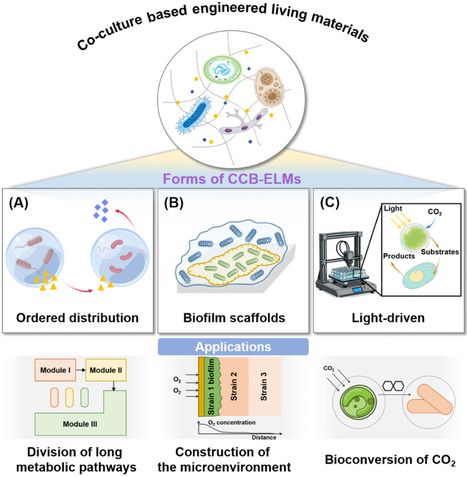
Coculture-based engineered living materials (CCB-ELMs) address the issue of poor stability in microbial coculture systems, which can be applied to biomanufacturing and CO2 conversion. This forum summarizes the latest research progress, core challenges, and future prospects of CCB-ELMs for sustainable biomanufacturing.

|
Scooped by
mhryu@live.com
March 25, 2:57 PM
|
Constructing and studying pangenome variation graphs (PVGs) supports new insights into viral genomic diversity. This is because such pangenomes are less prone to reference bias, which affects mutation detection. Interpreting the information arising from this is challenging, so automating these processes to allow exploratory investigations for PVG optimization is essential. Moreover, existing methods do not scale well to the smaller virus genome sizes and to facilitate analysis in laptop environments. To address this, we developed an easily deployable pipeline to facilitate the rapid creation of virus PVGs that applies a broad range of analyses to these PVGs. We present Panalyze, a computationally scalable virus PVG construction, analysis and annotation tool implemented in NextFlow and containerised in Docker. Panalyze uses NextFlow to efficiently complete tasks across multiple compute nodes and in diverse computing environments. Panalyze can also operate on a single thread on a standard laptop, and analyse sequence lengths of any size. We illustrate how Panalyze works and the valuable outputs it can generate using a range of common viral pathogens.

|
Scooped by
mhryu@live.com
March 25, 2:44 PM
|
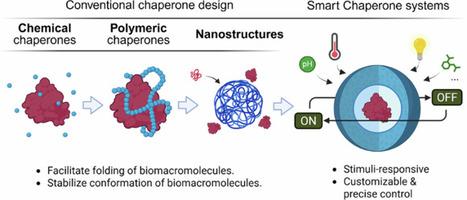
Chaperones are essential for the functions of many biomacromolecules as they facilitate folding, stabilize conformation, and prevent aggregation. Considering the high cost, low availability, and poor in vitro stability of natural chaperones, the development of artificial chaperones has been in high demand. Here, artificial chaperones that mimic the chaperoning functions on proteins and nucleic acids are reviewed. Aspects of compositions and structures of artificial chaperones, ranging from low-molecular-weight chemical chaperones to high-molecular-weight polymeric chaperones to complex nanostructures, are described. Furthermore, smart artificial chaperones that function in response to various external stimuli are described, pointing out their great potential to enable designs of more complex and functional systems for applications in biology, medicine, and industry. Molecular chaperones are proteins that assist in the proper folding of biomolecules, preventing misfolding and aggregation that can lead to diseases and economic losses in protein production. This study explores the development of artificial chaperones as cost-effective and customizable alternatives to natural chaperones. Researchers have designed chemical and polymeric chaperones that mimic natural mechanisms, using hydrophobic and electrostatic interactions to guide folding. For example, polymeric chaperones like PLL-g-Dex enhance DNA hybridization and protein folding by reducing repulsion and stabilizing structures. These findings are significant as they demonstrate the potential of artificial chaperones to improve protein stability and function in various applications. The study suggests that future research should focus on creating smarter chaperone systems that respond to environmental stimuli, enhancing their utility in biotechnology and medical treatments. This summary was initially drafted using artificial intelligence, then revised and fact-checked by the author. Artificial chaperones have been extensively developed as cost-effective, highly customizable, and stable alternatives to natural chaperones. This review firstly summarizes the conventional design of artificial chaperones in terms of compositional complexity from small-molecular chemical chaperones, large-molecular polymeric systems, to nanostructured assemblies. We further reviewed current advances in smart chaperone systems that can regulate chaperone functions in response to outer stimuli such as pH, temperature, and light. These progresses highlight the trending transition from material-based stabilization to programmable regulatory platforms with enhanced precision and expanded potential in complex biological, biomedical, and industrial applications.
|

|
Scooped by
mhryu@live.com
Today, 12:46 AM
|
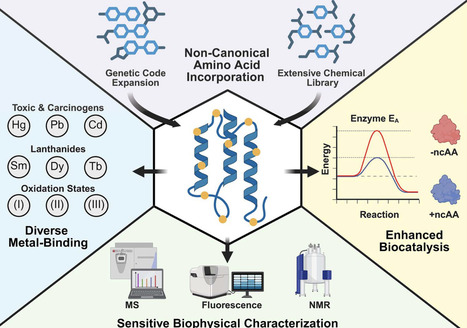
Non-canonical amino acids (ncAAs) are versatile molecular building blocks that can enhance nearly every aspect of protein engineering, from improving binding affinity to enabling precise quantitative analyses. This review highlights advances from 2020–2025 that demonstrate how expanding the amino acid repertoire unlocks new functionalities. We examine how genetically encoded ncAAs diversify and tune metal-binding properties, enabling programmable coordination and redox behavior in engineered enzymes. Further, we explore biocatalytic applications, including multi-fold activity enhancements in natural enzymes and the introduction of entirely novel reactivity in artificial systems. Finally, we discuss the growing use of ncAAs as intrinsic biophysical reporters, which support a wide range of spectroscopic methods for tracking structure, dynamics, and interactions at residue-level resolution. These capabilities establish ncAAs as essential tools that can be deployed at any stage of the protein design process, from constructing new catalytic centers to quantifying molecular behaviors in real time.

|
Scooped by
mhryu@live.com
March 25, 11:51 PM
|
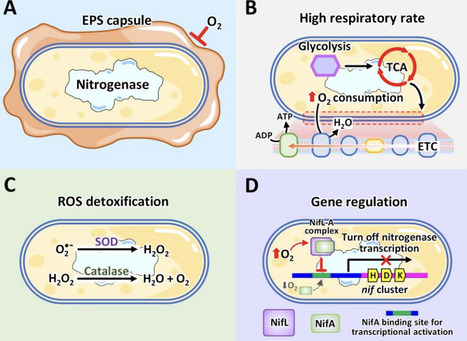
Ammonia (NH₃) is a critical molecule for agriculture, industry, and emerging energy applications. However, current industrial production by the Haber–Bosch process is energy-intensive and environmentally damaging, accounting for a significant portion of global CO₂ emissions. In contrast, biological nitrogen fixation (BNF) offers a sustainable alternative by converting atmospheric nitrogen (N₂) into NH₃ under ambient conditions through the action of nitrogenase enzymes. Among diazotrophic microorganisms, Azotobacter vinelandii stands out due to its ability to fix nitrogen aerobically, supported by unique physiological and genetic adaptations that protect its oxygen-sensitive nitrogenase. This review presents a comprehensive overview of NH₃ synthesis with A. vinelandii, highlighting its biochemical mechanisms, nitrogenase structure and function, and the protective strategies that enable aerobic nitrogen fixation. We further examine the diversity and advantages of Azotobacter strains, with a focus on A. vinelandii’s potential for engineered NH₃ production through synthetic biology, metabolic engineering, and emerging bio-based technologies such as photobiocatalysis and bioelectrochemistry. Recent innovations aimed at improving nitrogenase expression, cellular stability, and overall system efficiency are discussed in the context of advancing A. vinelandii as a robust chassis for industrially scalable, carbon-neutral NH₃ synthesis. The review emphasizes the importance of free-living nitrogen fixers in addressing the challenges of sustainable NH₃ production and provides insights into future directions for research and application.

|
Scooped by
mhryu@live.com
March 25, 11:24 PM
|
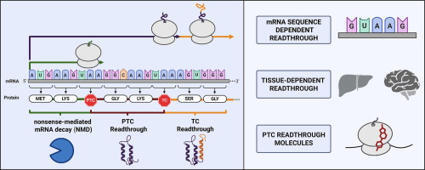
Accurate translation termination is essential for proteome integrity and is primarily governed by the release factors eRF1 and eRF3, which ensure precise recognition of stop codons and efficient release of nascent polypeptides. However, proteome integrity is challenged by mutations that generate premature termination codons (PTCs), leading to truncated, nonfunctional proteins and degradation of the aberrant transcript via nonsense-mediated mRNA decay (NMD). Collectively, these events account for ∼1800 human genetic diseases. Translational readthrough, the process by which near-cognate tRNAs decode stop codons and allow ribosomes to continue elongation beyond the stop codon, represents a possibility to suppress PTCs and restore full-length protein synthesis. Initially discovered in viruses as a mechanism to expand coding capacity, readthrough is now recognized as a regulated feature of eukaryotic gene expression influenced by both cis-acting sequence elements and trans-acting factors. Recent evidence highlights the remarkable context dependence of readthrough, revealing variation across transcripts, tissues, and developmental stages. In this review, we examine the molecular determinants that define stop codon recognition and readthrough efficiency, with particular emphasis on nucleotide context. We further discuss the mechanisms and binding sites of small molecules that promote PTC readthrough, and summarize the clinical development landscape of readthrough-inducing compounds for the treatment of diseases caused by nonsense mutations.

|
Scooped by
mhryu@live.com
March 25, 10:39 PM
|
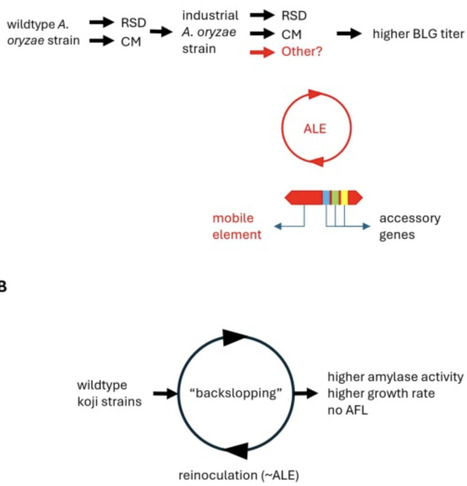
Precision fermentation promises to transform the food supply, but achieving cost parity with traditional production requires unprecedented protein titers. Unlike pharmaceutical proteins, food proteins demand exceptionally high yields, making strain optimization critical. Aspergillus oryzae, optimized over decades, provides a robust platform for milk protein expression, including beta-lactoglobulin (BLG). Rational strain engineering and adaptive laboratory evolution accelerate titer improvements, resembling traditional koji fermentation, where domestication of A. oryzae selected strains with enhanced amylolytic activity and faster growth. Domestication — including transposon and amylase gene expansion — highlight similarities between historical and modern strain optimization. We describe BLG strain optimization and discuss how mimicking domestication principles may fast-track strain improvement. We discuss recent advances and challenges in scaling precision fermentation to supply affordable, nutritious proteins. Our work on BLG illustrates how industrial biotechnology is reaching cost parity with dairy, underscoring the potential of evolution — natural and engineered — at the speed of fermentation.

|
Scooped by
mhryu@live.com
March 25, 4:53 PM
|
CRISPR–Cas9 is a powerful genome-editing tool, but genome-wide off-target activity can hinder therapeutic applications. Negative supercoiling ((−)SC) has been implicated in off-target activity, but a molecular-level understanding is lacking. Here, using (−)SC DNA minicircles, we observe supercoiling-driven structural defects in the DNA that are resolved by Cas9 binding. Cryo-electron microscopy structures of Cas9 bound in both the on-target and off-target configurations highlight that the Cas9 HNH domain is poised in a more catalytically competent conformation. New DNA–RNA mismatch geometries are accommodated across the protospacer and structural plasticity in the protospacer adjacent motif distal region of the protospacer is topology dependent. Together, our study reveals the molecular basis for (−)SC-induced Cas9 targeting and provides a framework for the design of next-generation high-fidelity CRISPR effectors with topological context. Cas9 structures explain topology sensing and off-target activation.

|
Scooped by
mhryu@live.com
March 25, 4:45 PM
|
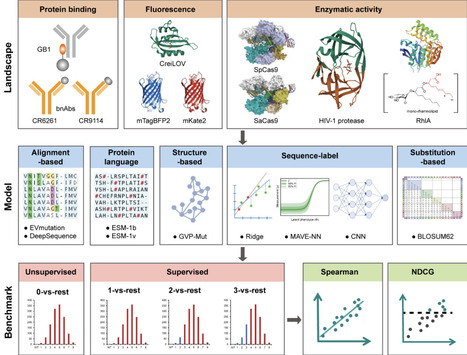
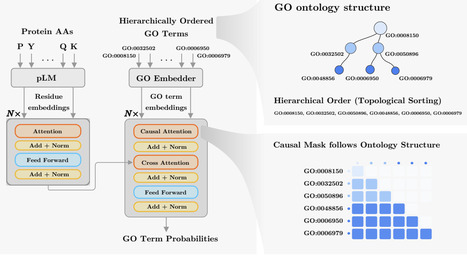
Accurate prediction of protein function is essential for elucidating molecular mechanisms and advancing biological and therapeutic discovery. Yet experimental annotation lags far behind the rapid growth of protein sequence data. Computational approaches address this gap by associating proteins with Gene Ontology (GO) terms, which encode functional knowledge through hierarchical relations and textual definitions. However, existing models often emphasize one modality over the other, limiting their ability to generalize, particularly to unseen or newly introduced GO terms that frequently arise as the ontology evolves, and making the previously trained models outdated. We present STAR-GO, a Transformer-based framework that jointly models the semantic and structural characteristics of GO terms to enhance zero-shot protein function prediction. STAR-GO integrates textual definitions with ontology graph structure to learn unified GO representations, which are processed in hierarchical order to propagate information from general to specific terms. These representations are then aligned with protein sequence embeddings to capture sequence–function relationships. STAR-GO achieves state-of-the-art performance and superior zero-shot generalization, demonstrating the utility of integrating semantics and structure for robust and adaptable protein function prediction.

|
Scooped by
mhryu@live.com
March 25, 3:57 PM
|
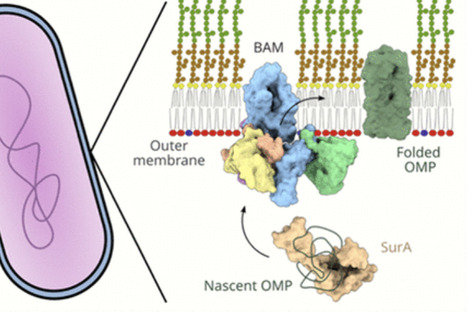
Assembly of the outer membrane (OM) of diderm bacteria is coordinated by the essential β-barrel assembly machinery (BAM) and is critical for cellular survival and pathogenicity. BAM operates in a membrane environment that is highly rigid and spatiotemporally organized, and functions without ready access to an energy source. In addition, BAM interacts with many other proteins to efficiently fold outer membrane proteins (OMP), assemble complexes in the OM, and maintain cell envelope homeostasis. In recent years, great strides have been made toward understanding the molecular mechanism of BAM-mediated (OMP) folding, with structural biology used to visualize the different stages of the pathways of OMP folding and membrane insertion. The conformational cycling of BAM and its ability to transiently form hybrid barrels with substrate OMPs facilitates their folding. Both these mechanistic features appear to be well conserved and are attractive targets for antimicrobials.

|
Scooped by
mhryu@live.com
March 25, 3:49 PM
|
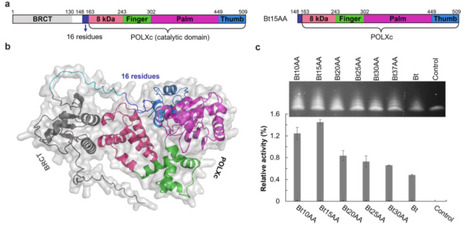
The 21st century has witnessed rapid advancements in synthetic biology, with DNA synthesis emerging as a foundational technology. Conventional phosphoramidite-based methods face significant limitations, including short DNA elongation lengths (<300 nt), hazardous chemical waste, and low stepwise incorporation efficiency. Enzymatic DNA synthesis using terminal deoxynucleotidyl transferase (TdT) offers a promising alternative, enabling kilobase-scale assembly with greater efficiency and minimal environmental impact. Here, we identified Bos taurus TdT (BtTdT) through UniProt database mining as a catalytically active scaffold for natural and 3′-modified dNTPs. Comprehensive characterization of BtTdT's enzymatic properties—including pH, temperature, metal ion dependence, and substrate specificity—revealed its optimal conditions. Truncation of the BRCT domain generated variants with enhanced activity compared to wild-type BtTdT. Guided by AlphaFold3-predicted structural models, we engineered a quintuple mutant (M5: Bt15AAR336L/K338G/L397M/E456S/D395G) optimized for 3′–ONH2–dNTP incorporation. M5 exhibited 30-fold activity enhancement relative to the triple mutant M3 (Bt15AAR336L/K338G/L397M) and achieved stepwise incorporation efficiency exceeding 98% in de novo synthesis of 10-nt ssDNA, demonstrating its potential for scalable enzymatic DNA synthesis. This work establishes a rational framework for TdT engineering through rational domain truncation and computational design, showing potential toward industrial-scale enzymatic DNA manufacturing.

|
Scooped by
mhryu@live.com
March 25, 3:20 PM
|
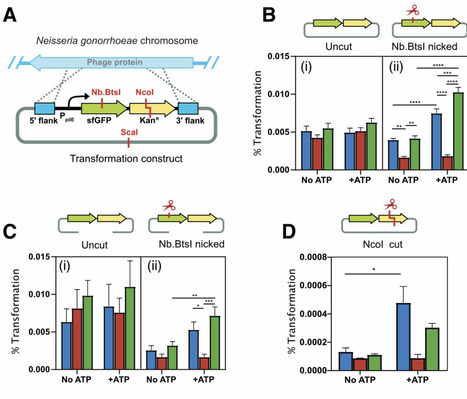
Lig E is a periplasm-targeted ATP-dependent DNA ligase found in many Gram-negative bacteria including Neisseria gonorrhoeae. Although LigE has been shown to have a role in biofilm formation, many Lig E-possessing bacteria are also naturally competent, suggesting a possible function in transformation with extracellular DNA. Here, we demonstrate that Lig E participates in bacterial competence by increasing transformation with nuclease-damaged extracellular DNA that contains single-stranded or cohesive breaks. We show that increased transformation with this restricted DNA is ATP-dependent, and that the ATP concentration increases in the extracellular milieu during maintenance of N. gonorrhoeae in liquid culture.

|
Scooped by
mhryu@live.com
March 25, 3:14 PM
|
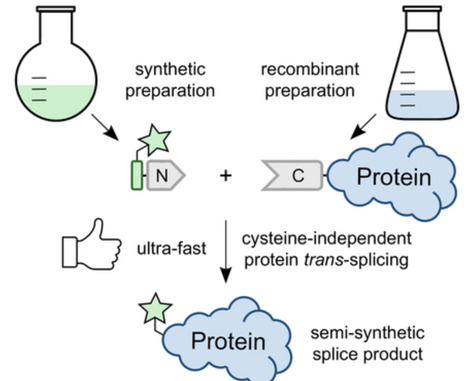
Cysteine-less split inteins have recently emerged as valuable addition to the protein labeling and modification toolbox as they can perform the protein trans-splicing (PTS) reaction under oxidizing conditions. Furthermore, their use is compatible with the chemical labeling and different redox states of cysteines in the extein sequences and hence the protein of interest. However, a rapidly splicing cysteine-less split intein with one short precursor fragment easily amenable to solid-phase peptide synthesis was still missing. A chemically synthesized split intein precursor allows for semisynthetic PTS, attractive to introduce fully synthetic sequence segments into the protein of interest. Here, we generate an optimized variant of the highly efficient split CL (cysteine-less) intein that splices with an ultra-fast rate on the second scale, comparable to the best performing split inteins. We achieved the nine- to 16-fold increase of the reaction rate by optimizing the artificial split site and the immediately flanking extein residues. Following chemical synthesis of the N-terminal precursor with an intein fragment of only 26 amino acids, we demonstrated protein semisynthesis by transferring a short fluorescently labeled peptide tag to a protein's N terminus with ultra-fast kinetics. The optimized split CL intein will thus further expand chemical protein labeling approaches.

|
Scooped by
mhryu@live.com
March 25, 3:01 PM
|
Protein engineering modifies protein molecules to achieve specific biological or technological functions. Protein design forms the core methodology, and in recent years, artificial intelligence (AI)-driven approaches have enabled more precise trait design in plants. This review highlights the convergence of protein structure prediction, generative sequence modeling, and function optimization to create synthetic proteins with improved specificity, stability, and activity in plant systems. We trace the development of protein design, from rational design to semi-rational strategies and AI-driven platforms that integrate structure prediction, sequence generation, and de novo design. We discuss eight application areas relevant to plant physiology and breeding: enhanced disease resistance via engineered immune receptors; insect resistance through optimized insecticidal proteins; abiotic stress tolerance through metabolic enzyme stabilization; improved nutrient use via transporter redesign; variant mining for trait fine-tuning; genome editing system optimization; environmental sensing with synthetic biosensors; and programmable regulatory circuits for plant factories (including controlled environment agriculture). Across these areas, we summarize design principles, advances, and translational considerations, emphasizing how AI expands sequence space and improves candidate prioritization. We also address current bottlenecks, including domain shift, generative model reliability gaps, limited portability, the genotype-to-phenotype gap, and design-to-validation workflow constraints. Finally, we propose a staged roadmap for AI-driven plant trait design, outlining milestones and requirements for translation into breeding.

|
Scooped by
mhryu@live.com
March 25, 2:45 PM
|
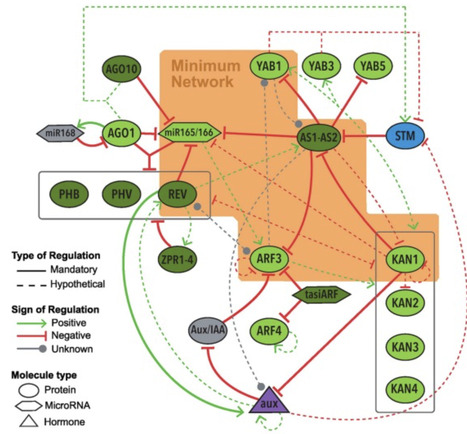
Biological processes are complex. Their behavior cannot be understood from the sum of their parts. Modeling, which allows for articulating hypotheses as equations and rules, has become indispensable for studying biological complexities. When written as computer code, mathematical models can be evaluated to test multiple potential outcomes. Modeling is useful for identifying essential system components and revealing new interactions. However, implementing models in biological research requires a solid understanding of computational and mathematical vocabulary, which typically requires formal training. Furthermore, communication barriers often hinder collaborations with computational modelers. To address this, we present an annotated glossary of terms associated with modeling gene regulatory networks. Aimed at biologists, we explain common computational modeling terms and frameworks using examples from published studies.
|
 Your new post is loading...
Your new post is loading...



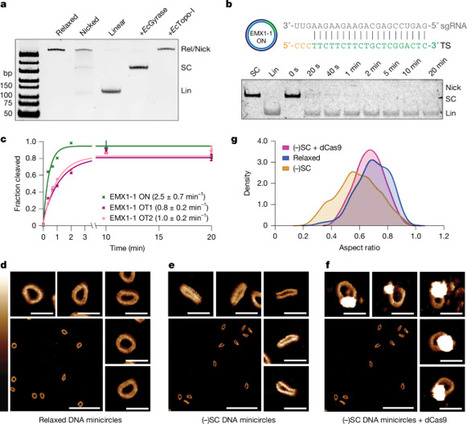








ha tk, Our (−)SC off-target structures show mismatch accommodation across the protospacer, including several in the seed region, with to date undescribed R-loop base-pair geometries. R-loop flexibility in the PAM distal region substantially increases the tolerance for mismatches and reduces the R-loop requirement for cleavage to just 14 bp.