 Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 1:17 AM
|
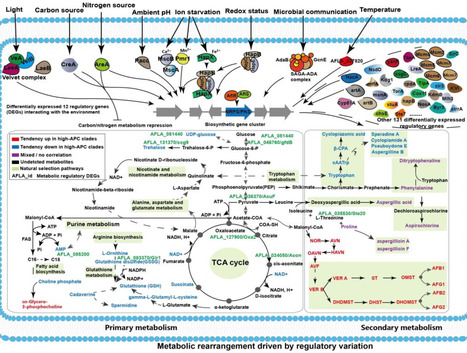
Chemical innovation is essential for fungi to adapt to ever-changing ecological environments. However, the environmental and evolutionary drivers of fungal metabolic differentiation remain ambiguous. Here, we show the phylogeographic diversity of 1052 Aspergillus flavus strains across four continents, as conducted through phylogenetic and biogeographical analysis, including 544 newly sequenced strains from China. These strains exhibit varying levels of population-specific mycotoxin production, as determined by population metabolomics analysis. We report a toxigenic subpopulation from China, identified through comparative population genomics analysis. Pan-metabolome analysis reveals strong phylogeographic metabolic patterns associated with specific ecological niches. Low-mycotoxin production clades harbor distinct uncharacterized biosynthetic gene clusters and produce different specialized metabolites instead. This discrepancy is only partially explained by variation in biosynthetic pathway genes, and changes in regulation and primary metabolism appear to mainly drive differentiation of specialized metabolite profiles across fungal populations, as indicated by pangenome profiling, metabolites-genome-wide association study, genotype-environment association study, pan-transcriptome analysis, and gene knockout experiments. Altogether, our results reveal how environmental shifts drive the fungal metabolic evolution, and provide insights for predicting the risk of harmful fungal outbreaks and for biogeographically-informed, precise control measures. Drivers of fungal metabolic diversity are incompletely understood. Here, the authors conduct a global genomics study of over 1,000 pathogenic fungi to show that geography shapes the metabolic diversity in Aspergillus flavus revealing how climate drives fungal chemical adaptive evolution.

|
Scooped by
mhryu@live.com
Today, 12:50 AM
|
Primary amines are the industrially important chemicals widely used in the manufacture of dyes, pharmaceuticals, and agrochemicals. In this study, we developed metabolically engineered E. coli strains for the efficient production of two-carbon primary amines, ethylamine and ethanolamine. For ethylamine production, metabolic pathways were optimized by enhancing flux toward the precursor l-alanine and by increasing the expression of a valine decarboxylase variant, VlmDV94I. Subsequent optimization of fed-batch fermentation resulted in the production of 36.3 g/L of ethylamine with a productivity of 0.432 g/L/h and a yield of 0.184 g/g glucose. For ethanolamine production, the l-serine biosynthetic pathway was engineered by introducing a feedback-inhibition-resistant enzyme and deleting l-serine utilization pathways. Additional deletion of the iclR and ygeA genes further boosted precursor supply by activating the glyoxylate shunt and preventing the conversion of l-serine to d-serine. We then screened serine decarboxylases and identified Atsdc from Arabidopsis thaliana as the most effective enzyme. Under optimized fermentation conditions, the final strain produced 36.9 g/L of ethanolamine with a productivity of 1.04 g/L/h and a yield of 0.184 g/g glucose. These titers and productivities represent the highest reported values for bio-based production of two-carbon primary amines, demonstrating the strong industrial potential of microbial cell factories.

|
Scooped by
mhryu@live.com
Today, 12:27 AM
|
Antimicrobial resistance (AMR) is a critical global health challenge. In this study, we developed a platform based on chromosome-free and nonreplicating simple cells (SimCells, size 1 to 2 µm) and mini-SimCells (size 100 to 400 nm) for targeted pathogen elimination. Engineered with surface-displayed nanobodies, SimCells and mini-SimCells selectively bind bacteria expressing specific antigens (e.g., OmpA in E. coli). The selective interactions facilitate close SimCell-pathogen proximity, enabling two antimicrobial mechanisms: direct injection of toxic effectors into bacterial cytoplasm via a heterologous expression of type VI secretion system (T6SS), and enzymatic conversion of aspirin into catechol by engineered salicylate hydroxylase, leading to sustained local production of hydrogen peroxide (H2O2). Our results demonstrate that both reprogrammed SimCells and mini-SimCells can eliminate target E. coli with high specificity and efficiency. Multidose reprogrammed mini-SimCell treatment led to a 103-fold selective reduction of targeted bacteria in mixed microbial communities, with minimal disruption to nontarget bacteria. We demonstrate that reprogrammed mini-SimCells, engineered with nanobody targeting outer membrane protein OmpA of the clinically relevant multidrug-resistant pathogen E. coli ST131, achieved elimination efficiencies over 97% at 24 and 48 h. This modularized “plug-and-play” antimicrobial platform provides a highly specific, efficient, and adaptable solution for combating diverse AMR pathogens.

|
Scooped by
mhryu@live.com
March 17, 11:53 PM
|
Microbial consortia, considered low-risk pesticides (LRPs), appear to be valuable tools for reducing our dependence on chemical pesticides. However, their use is limited by inconsistent product efficacy and registration difficulties. Artificial intelligence (AI) and machine learning (ML) offer solutions for designing and evaluating synthetic microbial communities (SynComs), predicting their compatibility, ecological stability, and biocontrol efficacy. The transition from laboratory discovery of SynCom-based LRPs to field application and commercialization could be significantly accelerated. Here, we review the methods and steps necessary to establish reliable SynComs and describe how AI and ML approaches could improve the construction and validation of SynCom-based LRPs to obtain more specific results that can contribute to their risk assessment.

|
Scooped by
mhryu@live.com
March 17, 11:14 PM
|
For the first time, the AlphaFold protein-structure database will include predictions of complexes of proteins — with the addition of 1.7 million ‘homodimers’ comprising two interacting strands of the same molecule. Since its release in 2021, this repository has become a bedrock in discovery and a first port of call for research projects that try to understand life at the molecular level. In the coming weeks, the AlphaFold database will also include complexes called heterodimers, which are made of two different proteins.

|
Scooped by
mhryu@live.com
March 17, 6:51 PM
|
Diverse phage-bacteria communities coexist at high densities in environmental, agricultural, and human-associated microbiomes. Phage-bacteria coexistence is often attributed to coevolutionary processes mediated by complex, pairwise infection networks. Here, using in vitro experiments and mathematical models, we explore how higher-order interactions function as a complementary, ecological feedback mechanism to stabilize phage-bacteria communities. To do so, we examine an environmentally-derived, synthetic phage-bacteria community comprised of five marine heterotrophic bacteria (Cellulophaga baltica and Pseudoalteromonas strains) and five associated phage. We used Bayesian inference to reconstruct free phage production in one-step growth experiments and then forecasted pairwise phage-bacteria community dynamics over multiple infection cycles. In contrast to model predictions of rapid bacterial population collapse, each bacterial strain persisted in the community. We hypothesized and then experimentally validated the relevance of infection attenuation at relatively high viral densities. We extended models into a community context, corroborating complex coexistence of all phage and bacteria. Life history traits inferred in community fits often differed from those inferred in a pairwise context, implicating higher-order interactions as an additional, ecological stabilization mechanism. Follow-up experiments confirm that phage traits (including burst size) can shift when infecting single vs. multiple strains. More broadly, these findings suggest that complex community coexistence of phage and bacteria may be more common than anticipated when including feedback mechanisms outside of the growth-dominated regimes of fitted pairwise models that do not reflect the full scope of ecologically relevant contexts.

|
Scooped by
mhryu@live.com
March 17, 2:36 PM
|
Colonization of plastic surfaces by microbial biofilms offers a promising starting point for engineering efficient biodegradation systems. However, most studies to date focus on characterization or prevention of biofilms on plastics in diverse environments and the potential biotechnological application for these systems has been underexplored. To address this, we report the efficient adhesion of E. coli cells to a range of plastic surfaces through overexpression of two key determinants of bacterial biofilm formation; curli and Antigen 43 (Ag43). A general trend of higher total biomass was observed from curli-mediated adhesion, but more uniform adhesion from Ag43 overexpression. We further demonstrate application of this technology through inducible adhesion of E. coli to polyethylene terephthalate (PET) surfaces and concurrent secretion of the PET depolymerase PHL7. Co-overexpression of curli fibres and secreted PHL7 resulted in 5.6-fold increase in terephthalic acid release in comparison to the non-adherent control. These methods offer a general approach to programmable adhesion of genetically tractable cells to plastic surfaces and concurrent secretion of degradative enzymes, and are anticipated to be broadly applicable across the field of plastic bioremediation technologies.

|
Scooped by
mhryu@live.com
March 17, 1:58 PM
|
High-throughput sequencing has generated vast genomic repositories that remain under-annotated, hampering enzyme discovery. We present an integrated pipeline that (i) builds a high-resolution, cross-kingdom phylogenetic database, (ii) mines candidates via multilocus phylogeny, (iii) predicts activities using an evolutionary-scale protein language model, and (iv) removes false positives through multilevel residue–atom contact rescoring. When applied to the r-BOX pathway, this approach uncovered numerous previously undocumented FadB, BktB, Ter, and YdiI homologues. Our activity model achieved R2 = 0.68 and reduced the RMSE on high-value targets by 11% compared to the prior SOTA (UniKP). Contact scoring improved early enrichment (EF1%) by 16-fold. Experimental validation targeting FadB increased titers from 0.65 g/L (shake flasks) to 1.7 g/L, reaching 10.2 g/L in a fermentation process. Together, these results establish a robust, generalizable framework for discovering scarce, high-value enzymes and prioritizing functional variants at scale.

|
Scooped by
mhryu@live.com
March 17, 1:50 PM
|
Lactopontin (LPN) plays a critical role in the growth and development of infants. Caprine milk and bovine milk are the primary raw materials for infant formulas. However, the differences between caprine and bovine LPN have not been studied. In this study, the structure, digestive characteristics, and active peptide composition of LPN from different sources were compared. Two N-glycosylation sites, Asn101 and Asn208, were identified in bovine LPN, while a single N-glycosylation site, Asn79, was found in caprine LPN. The in vitro infant digestion simulation results indicated that caprine LPN released a greater quantity of small peptides and amino acids. The intestinal digestion products were subsequently analyzed. The digestive peptides derived from caprine LPN may possess various potential biological functions. These findings provide insights into optimizing protein digestion and nutrient absorption in infant formula.

|
Scooped by
mhryu@live.com
March 17, 1:39 PM
|
Type III CRISPR–Cas systems confer antiviral immunity via cyclic oligoadenylate (cOA) signaling. Here, we elucidate a cooperative bacterial defense strategy involving two cOA-activated CRISPR-associated Rossmann fold (CARF)-containing effectors, adenosine deaminase CAAD and ribonuclease Csx1, in Thermoanaerobaculum aquaticum. Genomic analyses indicate widespread co-occurrence of CRISPR-associated adenosine deaminase (CAAD) with ancillary CARF-containing effectors in type III CRISPR systems, suggesting that multiple CARF-containing proteins may contribute to a coordinated cOA-dependent defense. Biochemical and structural studies reveal the intrinsic dynamics of CAAD hexamer, and demonstrate that cA4/cA6 binding stabilizes CAAD hexamers, triggering metal-ion-dependent conversion of ATP into inosine triphosphate. Concurrently, the downstream Csx1 is exclusively activated by cA4 to cleave single-stranded RNA. Strikingly, we found that both effectors are capable of degrading cA4, suggesting that this CAAD–Csx1 pair may be cross-regulated and achieve immunity through a dual-targeting mechanism: in response to infection, Csx1 degrades viral RNA while CAAD disrupts nucleotide metabolism via ATP deamination, which can be relieved via cA4 degradation when infection has been eliminated. This study proposes an enhanced defense mechanism through coordinated activation and regulation of multiple CRISPR effectors by a single signaling molecule, unveiling unprecedented complexity in CRISPR immunoregulation.

|
Scooped by
mhryu@live.com
March 17, 1:29 PM
|
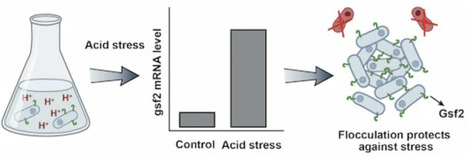
The social behaviors of microbes provide unique opportunities for testing social evolution theories. How can altruistic behaviors arise by natural selection is a central challenge in biology. Green-beard effect has been proposed as a basic mechanism for the evolution of altruistic behaviors. Yet, green-beard genes are generally thought to be rare. Here, we find that the Schizosaccharomyces pombe gsf2 gene mediates flocculation-like aggregation, and flocculation is triggered by acid stresses. gsf2-expressing cells preferentially adhere to each other. The expression of gsf2 is costly, but gsf2-expressing cells preferentially adhere to each other and protect each other from external stress. Gsf2 is highly variable in natural populations, likely contributing to different flocculation intensity. These findings suggest that gsf2 is a gradient green-beard gene that drives the altruism among gsf2 carriers. Moreover, we find that gsf2 is a new gene that originated very recently. Our results provide insights into the origin and evolution of green-beard genes.

|
Scooped by
mhryu@live.com
March 17, 1:07 PM
|
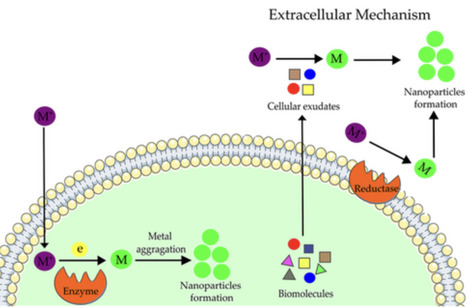
Biogenic nanoparticles are distinguished by their unique physical and chemical attributes, notably their potent antimicrobial activity against bacterial and fungal pathogens, as well as their cytotoxic effects on cancer cells. These nanoparticles are characterized by their biocompatibility, indicating their potential as effective antimicrobial agents and in oncological therapies. This article examines the existing literature on the antimicrobial and cytotoxic properties of nanoparticles derived from cyanobacteria, with particular emphasis on their implications for human health.

|
Scooped by
mhryu@live.com
March 17, 11:03 AM
|
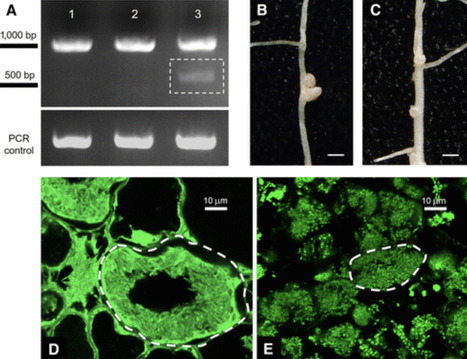
The model legume Medicago truncatula delivers nodule-specific cysteine-rich peptides to the intracellular bacteria within nodules to coerce the microbe into terminal differentiation, which coincides with nitrogen fixation in this species. Inside the host cell, the anterograde protein trafficking pathway is repurposed toward a new compartment, the symbiosome. Precise protein delivery within the nodule is critical to the success of the symbiosis in M. truncatula; without it, nodules form but do not fix nitrogen. For example, when the plant lacks DNF1, the nodule-specific 22-kDa subunit of the signal peptidase complex (SPC), the intracellular bacteria fail to fully differentiate, leading to defective nitrogen fixation. The present study shows that DNF1 became specialized in symbiosis through its nodule-specific expression, and we identified nodule-specific cis-elements that are crucial for that transcriptional control. Furthermore, we identified the nodule-specific SPC catalytic subunit and demonstrated that CRISPR/Cas9-induced mutation of this gene causes a symbiosis defect, which phenocopies the dnf1 mutant. These results suggest that a dedicated SPC in the nodule is co-opted for symbiosis through transcriptional regulation.
|

|
Scooped by
mhryu@live.com
Today, 12:52 AM
|
Synthetic microbial consortia (SMCs) represent a paradigm shift from monocultures to multi-strain systems that leverage ecological interactions for enhanced environmental adaptation and bioproduction. This review systematically sorts out engineering strategies for constructing stable SMCs, focusing on three core principles regarding host selection based on obligate mutualism (e.g., auxotrophs), pathway modularization to resolve metabolic conflicts, and dynamic regulation using tools like quorum sensing and optogenetics. We demonstrate the efficacy of SMCs in diverse applications including high-value compound synthesis and lignocellulosic biomass conversion through consolidated bioprocessing and inhibitor mitigation. SMCs enabling advanced functions in engineered living materials, environmental remediation, and biomedical innovation via division of labor are also described. Despite such progress, challenges in scalability and real-time control of SMCs under industrial conditions remain. We conclude that SMCs serve to bridge evolutionary ecology and biotechnology, offering robust solutions for sustainable biomanufacturing and beyond.

|
Scooped by
mhryu@live.com
Today, 12:42 AM
|
While environmental gradients are known to result in heterogeneous distributions of bacterial species along the gastrointestinal tract, the spatial distribution of genetic diversity within these species remains poorly understood. Because bacterial genetic variants influence host traits like inflammation and metabolism, understanding their distribution is critical. Here, we analyze ~30 common gut commensals in germ-free mice colonized with the same healthy human stool. Unexpectedly, we find that while species composition varied significantly across gut regions, genetic diversity within species remained remarkably uniform. This uniformity is driven by similar strain frequencies along the gut lumen, indicating that genetically divergent strains can coexist without spatial segregation. Furthermore, ~60 evolutionary adaptations arising within the mice tend to sweep globally throughout the gut, showing little region-specificity. We observe similar dynamics in conventional mice and humans, suggesting that uniform bacterial genetic diversity is a conserved, robust feature of mammalian gut ecosystems. Here, the authors show that while species composition varies significantly across gut regions, genetic diversity within species is remarkably uniform and driven by similar strain frequencies along the gut lumen and global selective sweeps throughout the gut.

|
Scooped by
mhryu@live.com
March 17, 11:56 PM
|
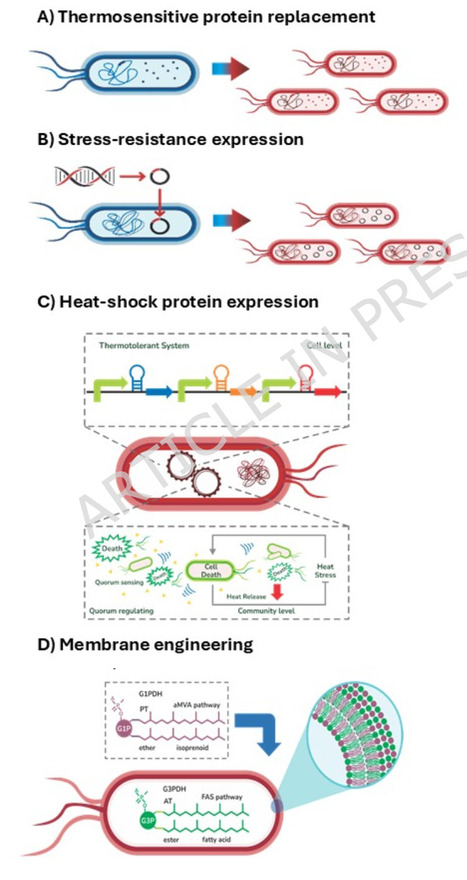
E. coli is one of the leading microbial chassis for bioproduct manufacturing in biotechnology. However, its use in high-temperature bioprocesses is restricted to its mesophilic nature. Rational engineering for improving thermotolerance in E. coli is challenging due to the limited understanding of the molecular functions and interactions involved in supraoptimal thermal adaptation. In recent decades, various approaches have been applied to increase the thermotolerance of E. coli. In this review, we examine the effect of temperature on cellular growth and thermal adaptation at supraoptimal temperatures and discuss how this knowledge can be applied to increase thermotolerance in E. coli. We particularly emphasize systems, synthetic, and evolutionary biology approaches that translate into systems metabolic engineering strategies to improve E. coli thermotolerance. We expect that systems-level insights into heat-stress physiology will enable data-driven strategies for the development of thermotolerant E. coli strains that can be used in high-temperature bioprocesses.

|
Scooped by
mhryu@live.com
March 17, 11:43 PM
|
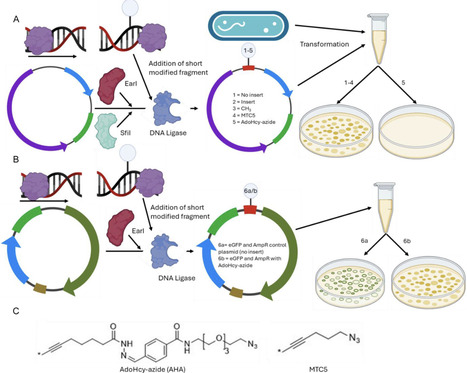
Artificial control of gene expression in bacteria offers interesting prospects for influencing bacterial pathogenicity and antibiotic resistance. We show that the methyl-transferase cofactor, AdoHcy azide, can silence gene expression in modified plasmids in some strains of E. coli, where ampicillin and kanamycin resistance as well as eGFP genes were selectively and independently disabled. The disabling of transcription is likely due to steric inhibition during transcription initiation, which is supported by Sanger and nanopore sequencing results. Both sequencing methods showed that 3–6 nucleotides were absent from around the modification site. Postgrowth, extracted AmpR/eGFP plasmid shows evidence of restriction, with sections of the plasmid, including the modification site, missing for the AdoHcy azide modified plasmids. Notably, the AdoHcy azide modification on the DNA appears to be resistant against demethylation in the BL21 strain of E. coli.

|
Scooped by
mhryu@live.com
March 17, 10:11 PM
|
The biological activity of many proteins is influenced by glycosylation, underscoring its essential role. However, no established recombinant technique currently enables the controlled shedading of glycosylated extracellular loops from transmembrane proteins. Here, we describe enzymatically controlled release of displayed proteins and peptides (ENCOREP), a strategy that enablesin situ expression at the cell surface and protease-mediated release, followed by collection from the cell culture medium. Using ENCOREP, we achieved the production of glycosylated peptides. These glycopeptides, derived from the large extracellular loop of the highly glycosylated CD63, illustrate the type of targets that are otherwise inaccessible with current methods but can be readily obtained using ENCOREP. Overall, ENCOREP provides a rapid and reliable approach to obtain glycosylated proteins or peptides while bypassing the conventional signal peptide–dependent secretory pathway.

|
Scooped by
mhryu@live.com
March 17, 6:43 PM
|
Gene regulation through promoter engineering is a cornerstone of synthetic biology, enabling precise control over transcriptional networks. However, experimental approaches remain labor-intensive. While artificial neural networks (ANNs) have improved regulatory element prediction, tools for promoter–transcription factor binding site (TFBS) recombination are still lacking. We present an ANN framework for context-aware design of synthetic promoters in Saccharomyces cerevisiae. The model predicts optimal TFBS insertion sites and the extent of promoter rewriting needed for successful integration. Applying this, we screened 6,011 native yeast promoters for compatibility with the TetR TFBS, generating a ranked list of high-confidence promoter–TFBS pairs. Experimental validation showed that model-designed promoters achieved repression rates up to 98.4%, without prior experimental characterization or tuning. We further rewired the yeast transcriptional network by introducing glucose-dependent regulation of an essential gene via Mig1 TFBS insertion. These results establish a scalable, predictive method for engineering regulatory sequences and reprogramming transcriptional logic.

|
Scooped by
mhryu@live.com
March 17, 2:33 PM
|
In E. coli, chromosome replication is regulated through ATP/ADP state of the DnaA initiator. The DDAH system inactivates DnaA in the post-initiation stage by promoting ATP hydrolysis through timely binding of the DNA-bending protein IHF to the datA locus, while the DARS2 locus reactivates DnaA in the pre-initiation stage via binding of IHF and another nucleoid protein Fis. The iron-sulfur cluster [(Fe-S)] assembly factor YgfZ is known to sustain replication initiation, central carbon metabolism, redox state and modification of tRNA A37 residues by MiaB, but the link between initiation and the others remains unclear. This study shows that YgfZ regulates initiation primarily by downregulating the DDAH system by repressing datA-IHF binding in a manner independent of MiaB. Also, the [Fe-S]-binding protein MnmA moderately downregulates datA-IHF binding. Furthermore, YgfZ globally downregulates basal IHF binding across the genome, while preserving IHF's timely binding at key loci including oriC and datA during the cell cycle, highlighting a novel strategy: YgfZ modulates both the cellular metabolic states and global genome dynamics to control replication initiation under various growth conditions.

|
Scooped by
mhryu@live.com
March 17, 1:53 PM
|
Xyloglucan oligosaccharides (XyGOs) are emerging prebiotics with limited sustainable production strategies. Here, two endoxyloglucanases, XyGH5 and XyGH74, from Paenibacillus sp. XP01 were identified. Genetic knockout revealed their synergistic role in bacterial growth on xyloglucan. Biochemical characterization showed that XyGH74 exhibited superior activity (Vmax = 76.25 U/mg) and thermal stability (optimum at 70 °C) compared to XyGH5, with its carbohydrate-binding modules (CBMs) crucially enhancing catalytic efficiency and substrate affinity. We further employed the highly efficient XyGH74 to produce tamarind xyloglucan oligosaccharides (TXOS). In vitro fermentation with human gut microbiota demonstrated that TXOS significantly modulated microbial composition, enriching beneficial genera such as Bacteroides and Parabacteroides, and notably enhanced the production of short-chain fatty acids, particularly butyrate, compared to the native polysaccharide. This study provides direct genetic evidence for synergistic glycoside hydrolase function in vivo and establishes XyGH74 as a robust enzymatic tool for scalable, high-value prebiotic production with enhanced health benefits.

|
Scooped by
mhryu@live.com
March 17, 1:47 PM
|
A miniaturized variant of the artificial luciferase (ALuc), named picALuc, has been generated through the deletion of N- and C-terminal residues in ALuc. Although picALuc is small and active, questions remain regarding its the structural organization and inter-residue interactions in the protein. Here, combining computational analysis and mutational studies, we show that the E50A mutation in picALuc results in an increased bioluminescence activity of the protein. Specifically, we generated a structural model of picALuc using the available structure of the Gaussia luciferase (GLuc) that revealed a ‘hole’ in the structure due to the deletion of N-terminal α-helices. Gaussian-accelerated molecular dynamics (GaMD) simulation revealed a rapid ‘compaction’ of the picALuc structure during the initial phase of the simulation and a number of residues such as E10, E50, and D94 showed salt bridge interactions. Mutation of the residues E10, E50, and D94 individually to an A revealed increased bioluminescence activity of the E50A mutant, while E10A and D94A mutants showed activities similar to the WT protein in living cells. In vitro assays revealed an increase in the Vmax of the E50A mutant, while Khalf and thermal stability of the mutant remained unchanged. Further, dynamic cross-correlation and principal component analyses of the GaMD simulation trajectories of the WT and the E50A mutant picALuc revealed altered collective dynamics in the protein. Finally, we developed a protein fragment complementation assay using picALuc that allows for the monitoring protein–protein interactions (PPIs) in live cells. We envisage that the brighter picALuc reported here will find broad applicability in developing bioluminescence-based assays.

|
Scooped by
mhryu@live.com
March 17, 1:32 PM
|
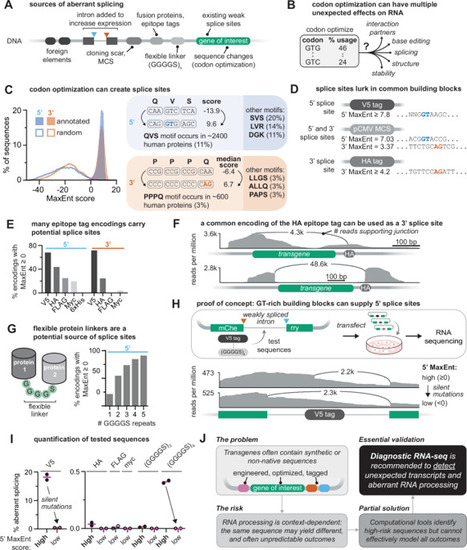
Plasmids are the workhorses of molecular biology: fast, flexible, and often taken for granted. We clone, overexpress, tag, and mutate freely, assuming they will faithfully produce RNA transcripts that match the intended DNA sequence. This assumption is rarely tested and often invalidated. Sequences in plasmid backbones, epitope tags, and codon-optimized regions may inadvertently harbor cryptic promoters or splice sites. The resulting unexpected transcripts and proteins, while often undetected, can distort results and propagate false conclusions through papers, grants, and even clinical trials. In this perspective, we highlight published cases where plasmids have distorted results and misled interpretation. We examine the mechanisms and consequences of plasmid-associated expression artifacts and offer practical strategies to minimize them. Finally, we call for a revision of community standards for experiments using transgenes: deposit complete plasmid sequences and verify the resulting transcripts using RNA-seq.

|
Scooped by
mhryu@live.com
March 17, 1:23 PM
|
Metagenome-assembled genomes (MAGs) provide crucial insights into the genomic diversity of uncultured microbes. However, MAG datasets deposited in public repositories such as INSDC are often difficult to reuse due to heterogeneous quality, inconsistent taxonomic and functional annotations, and insufficiently curated environmental metadata. While secondary MAG databases such as MGnify, IMG/M, and SPIRE provide standardized resources, they reconstruct MAGs de novo from public metagenomic reads and therefore do not represent the original MAGs reported in publications. To address this gap, we developed Microbiome Datahub, an open-access platform that systematically aggregates and re-annotates original MAGs from INSDC. We collected 214,427 MAGs, predicted genes by DFAST, performed quality assessment with CheckM, standardized taxonomic assignments with GTDB-Tk, inferred 27 phenotypic traits using Bac2Feature, assigned proteins to MBGD ortholog clusters and KEGG Orthology IDs using PZLAST, and annotated environmental metadata with the Metagenome and Microbes Environmental Ontology. Across these MAGs, the average completeness was 80.5% and contamination 1.8%; notably, the most frequent values were >95% completeness and <1% contamination, indicating that the majority of MAGs are of high quality. Comparative analyses showed that Microbiome Datahub provides phylogenetically and environmentally diverse MAGs: while the majority originated from vertebrate gut environments, a substantial number were also recovered from other habitats such as groundwater, including nearly 10,000 MAGs from the Patescibacteria. Inference of 27 phenotypic traits, including optimum growth temperature, further revealed ecological differentiation across phyla. Protein clustering revealed 56 million identity 40% clusters, with the majority unique compared with MGnify and GlobDB, and ~19% of proteins unassigned to MBGD ortholog clusters, underscoring their novelty. Microbiome Datahub integrates MAG genome sequences, gene and protein predictions, quality metrics, environmental and taxonomic annotations, ortholog cluster assignments, and phenotype predictions, all accessible via a web interface, API, and bulk downloads. By combining original MAGs with curated metadata and functional annotations, Microbiome Datahub constitutes a comprehensive and reusable resource that will accelerate microbiome and microbial genomics research. Video Abstract

|
Scooped by
mhryu@live.com
March 17, 1:06 PM
|
Microbial resources are crucial for biotechnology development and fundamental scientific research. Traditional microbial techniques fail to isolate and cultivate the vast majority of microorganisms in nature, severely limiting the discovery of novel microbial resources. The rise in artificial intelligence (AI) technologies provides new computational tools to overcome bottlenecks in microbial resource discovery and utilization. This review comprehensively examines the development of AI technologies in microbial isolation and cultivation over the past three decades from the perspective of microbial resource discovery. We propose a five-stage framework: the germination period (1997–2008), the early exploration period (2008–2015), the rapid development period (2015–2019), the deep learning (DL) explosion period (2020–2022), and the AI integration period (2023–present). We focus on how AI technologies at each stage address core challenges in microbiology—including insufficient knowledge reserves, dynamic phenotypic changes, and complex cultivation conditions—through applications at the genome, individual, and community levels. Our analysis demonstrates that, as AI technologies advance iteratively, microbial isolation and cultivation methods are transitioning from experience-driven to data-driven approaches, from single-objective to systematic integration, and from passive screening to active design. This methodological transition is expanding the scope of microbial resource discovery.
|
 Your new post is loading...
Your new post is loading...

























species changes while strains remain?