Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 1:06 AM
|
Protein structure serves as a critical bridge between sequence and functional annotation, particularly in establishing functional links among distantly homologous proteins with low sequence similarities. However, systematic protein structure-based functional annotations have been lacking in plants, where functions for a significant portion of the proteomes are still elusive. In this study, we leveraged protein structural data from 17 angiosperms to uncover previously unannotated protein functions in plants. After structural clustering, we used the plant clusters to query the UniProtKB/Swiss-Prot database (the expertly curated component of UniProtKB), a repository of expertly curated and reliably annotated proteins, and identified structural matches for thousands of plant clusters that were undetectable by sequence-based BLAST searches. We further selected 120 clusters, which are highly reliable in structural quality and alignment and are well-conserved across plant species, and uncovered various protein functions that are potentially widely important in plants. Finally, we experimentally analyzed one plant cluster structurally resembling the yeast peroxisomal peroxin 8 (PEX8) protein and verified that plant PEX8-like proteins can functionally complement yeast pex8 mutants. Our findings highlight the power of structural comparison in uncovering protein functions in plants.

|
Scooped by
mhryu@live.com
Today, 12:57 AM
|
CRISPR-Cas-based genome editing has revolutionized precise genome manipulation in plants, yet its practical application is still constrained by the inefficient delivery of editing reagents across different genotypes. Plant viruses are promising vehicles for delivering genome-editing components, bypassing plant transformation and/or tissue culture. Virus-induced genome editing (VIGE) has provided powerful tools for achieving heritable edits in model plants such as Arabidopsis thaliana and Nicotiana benthamiana. VIGE has now progressed from proof-of-concept to practical applications in agricultural crops. Notably, a recent breakthrough in VIGE in tiller has successfully achieved heritable genome editing in hexaploid wheat. This review outlines the latest advances in VIGE across diverse plant species, highlights its potential for crop improvement, and discusses future research directions.

|
Scooped by
mhryu@live.com
Today, 12:36 AM
|
Bacterial outer membrane vesicles (OMV), are nano-sized, spherical structures released by Gram-negative bacteria that play diverse roles in bacterial physiology, including communication, nutrient acquisition, and host interactions. These vesicles bud from the bacterial outer membrane and contain lipopolysaccharides, periplasmic proteins, nucleotides, and other biomolecules. The Vesicle Nucleating peptide (VNp) is a short peptide tag that, when fused to the amino terminus of a protein of interest, promotes the formation of bespoke recombinant extracellular vesicles in E. coli, enabling efficient production and simplified purification of recombinant proteins. Here, we characterize VNp-induced vesicles and compare their composition and organization with naturally produced E. coli OMVs. While both vesicle types possess a single outer membrane-derived lipid bilayer, recombinant protein is highly enriched within the VNp vesicles compared to endogenous OMVs. VNp-fusions and periplasm-targeted recombinant proteins localize to distinct vesicle populations, with VNp- fusions showing markedly higher intra-vesicular concentrations and vesicular purity, compared to the OMV targeted protein. OmpX co-expression further enriched the VNp-fusion content of vesicles, further enhancing yield. The VNp-vesicle lumen is an oxidizing environment, thus supports formation of inter- and intra-molecular disulfide bonds within encapsulated proteins. Overall, VNp-induced vesicles represent a distinct class of recombinant extracellular vesicles that offer a simple and efficient route for producing and purifying concentrated, correctly folded recombinant proteins, expanding the utility of bacterial vesicle systems for biotechnological applications.

|
Scooped by
mhryu@live.com
Today, 12:24 AM
|
Phage therapy is being used to combat pathogenic bacterial infections that threaten plant, animal, and human health. However, its application remains limited by high host specificity and the emergence of bacterial resistance. In this study, we addressed the key issues in phage therapy using rice bacterial blight pathogen Xanthomonas oryzae pv. oryzae (Xoo) strain N1 and its lytic phage NP1. Strain N1 acquired resistance to the phage NP1 through mutations and downregulation of lipopolysaccharide (LPS) biosynthesis genes. A directed evolution assay using phage NP1 and the resistant strain N1R resulted in the development of phage E12–2, which overcame bacterial resistance, expanded its host range and improved bacterial suppression by targeting alternative LPS binding sites. Moreover, genome analysis identified two amino acid substitutions (V303L and G317V) in its tail fiber protein. Additionally, phage E12–2 improved disease control efficiency by 51 % compared to the wild-type phage NP1 and induced plant immunity in a plant disease model. These findings enhance our understanding of how bacteria-phage evolution shapes the dynamics of phage therapy in plants.

|
Scooped by
mhryu@live.com
Today, 12:19 AM
|
Microbial methanogenesis is a major contributor to global warming and methane fluxes represent a loss of energy and electrons from industrial ecosystems. The chemical space of methane control strategies is still under-explored. Most known methanogenesis inhibitors target methanogenic archaeal enzymes. However, interference with syntrophic electron exchange in methanogenic systems presents an additional target for methane control. Here we show that hypophosphite, an inorganic formate analog, is a potent and selective inhibitor of syntrophic methanogenesis versus primary fermentation in rice field sediments and cattle rumens. Hypophosphite is also generally recognized as safe and relatively non-toxic to plants and animals. Genetic screens and physiological assays in the model methanogen Methanococcus maripaludis S2 implicate formate metabolism as the target of hypophosphite inhibition. Currently, there is no known biological pathway for anaerobic hypophosphite oxidation and hypophosphite is stable in anoxic sediments for weeks to months. Given its widespread natural occurrence, we propose that hypophosphite may modulate carbon cycling in natural environments. Taken together, our results suggest that hypophosphite could be used as a safe, inexpensive, strategy for methane control in syntrophic methanogenic ecosystems.

|
Scooped by
mhryu@live.com
March 12, 11:33 PM
|
Transformer-based models (TBMs) are state-of-the-art deep learning architectures that predict protein structural and functional features with high accuracy. Despite methodological differences, they all rely on large protein sequence datasets structured by homology, as homologous proteins typically share structure and function. However, 5-30% of eukaryotic proteomes consist of orphan proteins - sequences without detectable similarity to known families. Although they may share structural or functional traits with characterized proteins, their lack of homology makes them ideal for evaluating TBM generalization beyond familiar sequence space. We compared predictions from several widely used TBM architectures on an expert-curated set of orphan proteins from the Meloidogyne genus. None of these proteins has an experimentally determined structure. To assess model performance, we conducted consistency analyses, comparing predicted features with those observed in sets of known homologous proteins and across models. Multiple sequence alignment-based approaches such as AlphaFold2 performed poorly on orphan proteins, as did single-sequence or embedding-based language models including ESMFold, OmegaFold, and ProtT5. This limited performance cannot be fully attributed to intrinsic disorder, as confirmed by independent non-TBM disorder predictors. While accurate tertiary structure prediction remains out of reach, secondary structure is more reliably captured: predictors share about 70% of secondary structure elements on average, regardless of global fold similarity, and these elements are consistently identified by dedicated secondary structure tools.

|
Scooped by
mhryu@live.com
March 12, 11:21 PM
|
The ability to sequence proteins without reliance on a genomic template defines a critical frontier in proteomics. This approach, known as de novo protein sequencing, is essential for applications in antibody sequencing, microbiome proteomics, and antigen discovery, which require accurate reconstruction of target sequences. To advance this field, we here explore two hyperthermoacidic archaeal (HTA) proteases for de novo antibody sequencing, benchmarking them against trypsin and chymotrypsin. Each HTA-protease generated about five times more unique peptide reads than trypsin or chymotrypsin, providing high redundancy across all complementarity-determining regions. Combined with EAciD fragmentation on a ZenoTOF, this methodology enabled complete, unambiguous antibody sequencing. De novo analysis showed much higher alignment scores and reduced the sequence errors by using the HTA-generated data. With short digestion times, minimal sample cleanup, and analysis in just a single liquid chromatography-mass spectrometry (LC-MS/MS) run, this streamlined single-protease approach delivers a scalable and efficient strategy for de novo protein sequencing across diverse applications.

|
Scooped by
mhryu@live.com
March 12, 4:54 PM
|
A device capable of sampling natural gas under aseptic conditions and in complete safety has been deployed along the transmission grid for the first time. Microbial endospores, resilient enough to survive the extreme conditions of gas transmission and storage, have been detected and isolated throughout high-pressure pipelines and underground reservoirs. In four underground gas storage (UGS) facilities, three in deep aquifers and one in a depleted reservoir, endospores of the same hydrogenotrophic bacterial species from the family Peptococcaceae have been identified, sometimes separated by hundreds of kilometers, and at two different points in the pipeline network. Cultural and genomic analyses show these bacteria can perform acetogenesis, biofilm formation, and produce formate. Hidden within pipelines, these microbes survive long journeys and actively participate in biogeochemical cycles in UGS. Several recent studies on dihydrogen injection into deep aquifers have shown the ubiquity of bacteria similar to these, responsible for formate formation through modified acetogenesis. This formate can serve as a carbon source or inhibit sulfate reduction at high concentrations. Understanding their role offers critical insights into microbial life in the deep biosphere and the potential impacts of future dihydrogen injection into natural gas systems. Their ability to thrive in extreme environments makes these microbes key players in the evolving landscape of underground energy storage and transport.

|
Scooped by
mhryu@live.com
March 12, 4:47 PM
|
Microbial exopolysaccharides (EPSs) serve multiple industrial and environmental purposes operating as complex biopolymers produced by bacteria and fungus, as well as by yeast and microalgae. The structural diversity of microbial EPS enables the biofilm formation, the stress resistance and the nutrient storage, comprising homopolysaccharides and heteropolysaccharides. Soil structure receives substantial improvement through EPS because the polymers help aggregate particles, retaining more water and trapping heavy metals, that results in enhanced soil fertility useful in sustainable agricultural practices. Moreover, through the presence of EPS-producing bacteria, plants can establish beneficial connections with microorganisms that improve their tolerance to environmental factors, including salt exposure, drought conditions and extreme temperature changes. Such polymers find applications in the bioremediation and pharmaceutical fields because they present significant pharmacological properties, such as antibacterial, anti-inflammatory activities and antioxidant behavior. Their biodegradable nature and eco-friendly properties make it eligible as a sustainable choice to replace synthetic polymers. This paper broaches the multiple ways how EPS improves plant wellness and enhances soil quality. Potential solutions emerge from microbial EPS research to address global challenges in agricultural sectors, biotechnological fields, and environmental management domains.

|
Scooped by
mhryu@live.com
March 12, 4:35 PM
|
Like many tools in modern biology, fluorescent proteins have developed from broad cross-disciplinary effort. Colleagues push an emerging branch of fluorescent protein development — magnetosensitive fluorescent proteins, in particular, MagLOV into some intriguing new areas, demonstrating the use of MagLOV for often-disconnected fields like quantum property inference using a standard fluorescence microscope and a novel fluorescence-based magnetic resonance imaging (MRI) approach. To develop MagLOV for sensing capacities and standard widefield fluorescence microscopy readouts, the researchers apply directed evolution to the ancestor of the original MagLOV proteins, AsLOV2, toward magnetoresponsive fluorescent proteins (MFPs) with improved magnetic field effect contrast and optically detected magnetic resonance saturation rate, resulting in new variants called MagLOV2 and MagLOV2 fast. The researchers characterized the variation in magnetic responses and tested a theorized quantum mechanism for magnetically regulated fluorescence. To demonstrate the utility of MFP systems that can be fully expressed within cells and have stronger magnetic properties, Steel and colleagues developed a combined MRI–widefield fluorescence imaging system, capturing differences from MFP-expressing cells on the scale of millimeters and testing the utility of MagLOV 2 in sensing its magnetic microenvironment.

|
Scooped by
mhryu@live.com
March 12, 4:17 PM
|
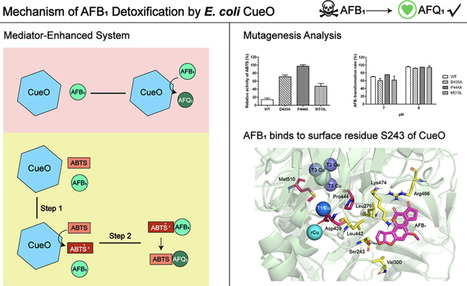
Aflatoxin B1 (AFB1) is a highly toxic mycotoxin that threatens global food and feed safety. While enzymatic detoxification is a promising strategy, robust and efficient enzymes remain scarce. This study identified that the multicopper oxidase CueO from E. coli CG1061 transforms AFB1 into the less toxic aflatoxin Q1, exhibiting optimal activity at pH 8 and 60 °C. The CueO-2,2′-azino-bis (3-ethylbenzothazoline-6-sulfonic acid) (ABTS) mediator system achieved >90% of AFB1 transformation in 10 min and complete transformation in 20 min, significantly outperforming CueO alone (51% in 60 min). Mechanistically, CueO oxidizes ABTS to form the ABTṠ+ radical, which subsequently oxidizes AFB1. Mutants (Met510Leu, Asp439Ala, and Pro444Ala) exhibited enhanced enzymatic activity toward ABTS but showed no improvement in the catalytic efficiency for AFB1 transformation. Molecular docking suggests this is because AFB1 binds to surface-exposed residue Ser243, distal to the active site. These findings highlight E. coli CueO’s a promising candidate for managing AFB1 contamination.

|
Scooped by
mhryu@live.com
March 12, 1:49 AM
|
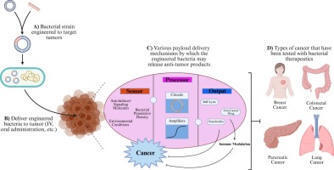
Microorganisms have had an established relationship with the maintenance of human health, and recent advancements in genetic engineering and synthetic biology have allowed the development of engineered microbes as targeted disease treatment. This review evaluates the rising use of microorganisms as therapeutic agents, particularly in the treatment of cancer. These engineered microbes are able to detect disease markers and precisely deliver therapeutic payloads to affected sites. These bacterial-based therapeutics show remarkable promise against cancers like pancreatic, breast, lung, and colorectal cancers by targeting tumor microenvironments and enhancing antitumor immunity. While obstacles still remain, such as interindividual variability, safety concerns, and regulatory barriers, the medicinal potential of microbial therapeutics is promising. Future research should focus on furthering the understanding of microbe-host interaction mechanisms and refining bacterial engineering techniques to continue to develop more precise and effective therapies. Ultimately, harnessing the power of microorganisms has the potential to revolutionize the treatment of complex diseases, such as cancer.

|
Scooped by
mhryu@live.com
March 12, 1:45 AM
|
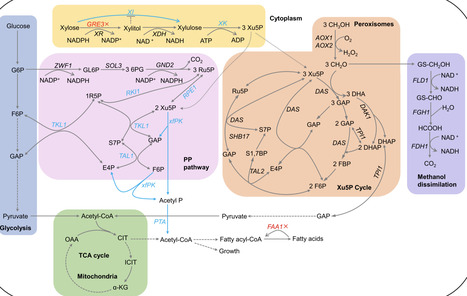
Efficient xylose utilization is crucial for biomass hydrolysate valorization. However, Komagataella phaffii cannot efficiently utilize xylose. Here, we constructed xylose isomerase (XI)-xylulokinase (XK) pathway, nonoxidative pentose phosphate pathway (PPP), and nonoxidative glycolysis (NOG) pathway in K. phaffii to increase cell growth on xylose. Additionally, the bypass pathway of xylose metabolism, the high osmolarity glycerol/mitogen-activated protein kinase (HOG-MAPK) signaling pathway, and possible negative transcription factors were blocked to further promote xylose catabolism. Moreover, adaptive laboratory evolution (ALE) dramatically improved xylose utilization, and six potential targets were identified through multiomics and reverse engineering. The engineered strain exhibited the highest reported specific growth rate μmax of up to 0.042 h−1 and lag time of 25.0 h with a biomass yield of 0.366 g dry cell weight/g from sole xylose in minimal media. This strain also showed faster metabolite turnover, efficient free fatty acid (FFA) production and partial amelioration of glucose repression effect from xylose alone. The engineered metabolic plasticity described here will facilitate the regulation of xylose catabolism in other nonnative xylose-consuming yeasts.
|

|
Scooped by
mhryu@live.com
Today, 1:01 AM
|
Rice is the main source of calories for more than half of the world’s population, so yields must increase to feed an estimated 5 billion people by 2050, despite the effects of resource depletion and climate change. The urgent demand for resilient, higher-yielding varieties cannot be met by conventional breeding, which is too slow, or transgenesis, which is burdened by regulatory issues. In contrast, genome editing enables precise, efficient, and transgene-free improvements for gene knockouts. This review covers major advances in rice genome editing since 2020, including innovations in base and prime editing, multiplex editing, and delivery. We highlight applications that improve yield, abiotic and biotic stress resilience, and nutritional quality, as well as challenges and future directions affecting precision, delivery, and regulatory policy.

|
Scooped by
mhryu@live.com
Today, 12:47 AM
|
Microbial natural products are a foundational source of therapeutic agents, yet a vast majority remain inaccessible due to the limited expression of their biosynthetic gene clusters under standard laboratory conditions. To address this challenge, we developed a simple and broadly applicable screening approach based on the idea of antibiotic hormesis wherein high-dose growth-inhibitory antibiotics serve as low-dose elicitors of secondary metabolites. We generated an in-house library of all available clinical antibiotics and exposed nine phylogenetically diverse bacteria to high-dose growth inhibition and low-dose metabolite stimulation assays. The approach revealed the induction of cryptic metabolites with every strain tested. Four of these were selected for scale-up fermentation and comprehensive metabolomic analysis, leading to the identification of eight known but strain-novel cryptic metabolites and nine structurally unique, previously undescribed natural products. These findings underscore the utility of the antibiotic hormesis approach as a rapid and scalable platform for the discovery of natural products and therapeutic leads. Moreover, the consistent elicitation of cryptic metabolites highlights the generalizability of low-dose antibiotics as key signaling molecules in microbial metabolic regulation. Beyond expanding the repertoire of accessible natural products, this work lays the foundation for systematic studies into the molecular and ecological mechanisms that govern cryptic metabolite biosynthesis.

|
Scooped by
mhryu@live.com
Today, 12:33 AM
|
Transposons are among the most abundant and diverse mobile genetic elements in nature. A unique class termed IStrons encodes a transposase for transposon mobility, an RNA-guided nuclease for maintenance, and a self-splicing group I intron for element removal from host mRNA. However, it is unclear how a single polynucleotide sequence balances these distinct biochemical functions. Here we employed pooled library mutagenesis coupled with high-throughput sequencing to systematically dissect the molecular determinants of IStron transposition, RNA-guided DNA cleavage, and self-splicing. We found that the terminal trinucleotide of the transposon right end is constrained by all three functions, identifying a molecular convergence point. Cross-assay comparisons revealed that most variants maintained or lost activity across multiple assays simultaneously. However, a subset selectively retained one activity while losing another, revealing antagonism between DNA cleavage and splicing governed by guide RNA structural stability. Increased GC content at the base of the guide RNA 3' terminal stem-loop abolished splicing while maintaining DNA cleavage, and the properly folded guide RNA sterically occluded alternative splice sites, ensuring splicing accuracy across variable flanking contexts. Thus, IStron transcripts overcome an inherent trade-off between guide RNA maturation and splicing, with RNA structural stability as the primary determinant of pathway choice.

|
Scooped by
mhryu@live.com
Today, 12:20 AM
|
Droplet sorting technology has the potential to revolutionize the biotechnology sector as it provides massive high-throughput screening capacity, but the technology remains not accessible for a wider audience yet. There is a need for more affordable droplet sorting platforms to design cell factories and screen cell libraries. In here we demonstrate our droplet cytometry/sorter platform for single-cell screening of yeast cells based on their fluorescence signal.

|
Scooped by
mhryu@live.com
March 12, 11:37 PM
|
The availability of large-scale protein structure collections enables structure-based analysis of their function and evolution beyond what is possible from sequence alone. However, applying three-dimensional structure comparison at scale remains computationally demanding and limits practical exploration of large experimental and predicted collections. This creates a need for fast, structure-based search methods that retain biological relevance while enabling large-scale exploration. In this paper, we present AlphaFind v2, an application for finding structurally similar proteins in the AlphaFold Database https://alphafold.ebi.ac.uk/ of predicted structures. AlphaFind v2 uses fast pre-filtering via state-of-the-art protein embeddings that preserve structural information, followed by refinement with US-align. The application presents multiple complementary search modes, including (i) search over full protein chains, (ii) search aware of the AlphaFold pLDDT metric, restricting similarity computation to the most stable and structurally relevant regions, (iii) search over protein domains from the TED database https://ted.cathdb.info/ , and (iv) a multidomain search mode, combining multiple chain-level domain matches within a single score and alignment. The application accepts protein identifiers and returns similar proteins with metrics, rich metadata, and interactive superpositions. AlphaFind v2 additionally allows searching within an organism or CATH label and matches the proteins with experimental structures. AlphaFind v2 is accessible at https://alphafind.ics.muni.cz/

|
Scooped by
mhryu@live.com
March 12, 11:30 PM
|
Large Language Models (LLMs) are increasingly applied to genomic tasks, yet core challenges remain concerning tokenization, evaluation, and data scarcity. This study focuses on promoter classification and systematically evaluates four tokenization methods: non-overlapping 6-mer, overlapping 6-mer, Byte Pair Encoding (BPE), and WordPiece (WPC). We show that the commonly used k-mer approach, specifically the non-overlapping variant, outperforms BPE and WPC across eight organisms, challenging assumptions derived from natural language processing. To ensure robustness, we evaluated performance under two distinct negative data strategies: positive-promoter-shuffled and random-non-promoter-fragments. Using a positional SHAP framework, we demonstrate that the model learns biologically plausible positional patterns rather than exploiting artifacts from these negative data generation processes. Furthermore, evolutionary-informed transfer learning experiments and external validation on an unseen organism reveal that training on phylogenetically related species significantly improves performance, particularly in low-data regimes. These findings underscore the significant impact of tokenization and negative data design, providing practical guidance for refining genomic classifiers.

|
Scooped by
mhryu@live.com
March 12, 5:03 PM
|
Microbial communities are structured through complex interactions that are difficult to observe directly. Co-occurrence networks offer a way to infer community structure, revealing (not exclusively) potential biotic interactions. Such networks have been inferred for diverse biomes and repeatedly found to be modular, yet the ecological significance of this modularity remains underexplored. We tested whether clusters within co-occurrence networks (“cohorts”), are universal and ecologically meaningful units by assessing their ubiquity, stability, and environmental specificity across diverse ecosystems. Our meta-analysis spans 25 previously published 16S rRNA gene amplicon sequencing datasets (14 160 samples) and covers high environmental variability ranging from aquatic, terrestrial to anthropogenic environments. Microbial co-occurrence networks consistently exhibited high modularity across biomes. Inferred cohorts were ubiquitous and represented up to 90% of the community composition. Our findings demonstrate that modularity is a fundamental and generalizable feature of microbial community organization, indicating the existence of stable subcommunities. Highly similar cohorts were inferred even across different, unconnected environments and datasets, and showed consistent responses to environmental gradients, indicating that their composition is to a large degree deterministic and predictable. The overall cohort structure and environmental preferences were independent of the sample size and the inference algorithm, underlining the robustness and applicability of the results. Recognizing these microbial cohorts as a meaningful level of microbial organization will refine microbial community ecology, cultivation strategies, and predictive modelling of microbial dynamics.

|
Scooped by
mhryu@live.com
March 12, 4:50 PM
|
Bacteria of the genus Bacillus are widely used in agriculture due to their ability to synthesise IAA, but the metabolic pathways of IAA synthesis and the regulatory mechanisms of key genes remain unclear. In this study, the biochemical characteristics of Bacillus cereus, Bacillus subtilis and Bacillus safensis were detected. It was found that Bacillus subtilis has a higher indoles synthesis efficiency, Bacillus safensis has stronger salt tolerance and Bacillus cereus has more prominent alkali tolerance. Through whole-genome sequencing and comparative analysis, combined with the retrieval of key genes in the tryptophan metabolic pathway and the metabolic analysis of indole compounds, it is clarified that Bacillus cereus has three IAA synthesis pathways, namely TAM, IAM and IPyA; Bacillus subtilis contains IAM and IPyA pathways; and Bacillus safensis only has the IPyA pathway. This study offers new insights into Bacillus metabolism, high-quality strains for microbial agents, and matters for plant-growth bacteria application and agricultural sustainability.

|
Scooped by
mhryu@live.com
March 12, 4:43 PM
|
Polyethylene terephthalate (PET), an abundant synthetic polyester, is the only plastic that has been enzymatically recycled at an industrial scale. Over the last decades, research efforts have focused on screening and engineering PET-degrading hydrolases (PETases), aiming to identify variants that can operate efficiently in both environmental and industrial settings. The detection of potential PETases from marine and terrestrial ecosystems has primarily been conducted via metagenomics using homology strategies. However, the use of benchmark PETases as references has limited the searches, narrowing the sequence landscape. Currently, there remains a need to identify efficient thermophilic, halotolerant and pH-robust PETases for the industrial biocatalysis of PET. In line with this, in this article, we discuss recent findings related to the following topics: (i) the identification of suitable ecosystems for mining PETases; (ii) the discovery of PETases via the restructuring of microbiomes; (iii) advancements in metagenomics and artificial intelligence (AI)-based approaches for the detection and ranking of PETases and (iv) the future of PET biocatalysis. Overall, we suggest that disrupting microbiomes with polyester-rich substrates, combined with innovative computational and AI-based strategies, can be an effective pathway for the discovery of PETases that can be used as scaffolds for protein engineering and biotechnological applications.

|
Scooped by
mhryu@live.com
March 12, 4:21 PM
|
Limonene is a high-value monoterpenoid used in food, pharmaceutical, and fragrance applications. To overcome limitations of conventional production, this study developed a biosynthesis platform using acetate as the sole carbon source. Acetate is an ideal nonfood substrate that can be sustainably derived from industrial waste streams, syngas, and CO2 reduction. Initially, the heterologous synthetic pathway was constructed to yield 0.13 mg/L limonene from acetate. By introducing the mutated EfMVASA110G to enhance catalytic efficiency, along with promoter engineering and substitution of the synthesis substrate, the production titer was enhanced to 49.24 mg/L. Subsequent host optimization through NADPH regeneration and pathway knockout, combined with fermentation optimization, increased the titer to 167.45 mg/L. Ultimately, whole-cell biocatalysis achieved 450.72 mg/L, establishing the highest reported acetate-based limonene titer. Overall, this study establishes a framework for biosynthesis of high-value terpenoids from nonfood feedstocks and demonstrates acetate as a promising carbon source for advanced bioproduction systems.

|
Scooped by
mhryu@live.com
March 12, 2:42 PM
|
Robust biocontainment is essential for the safe use of engineered microbes, but existing strategies suffer from genetic instability and/or laborious construction. Here, we present ATTACH, a kill switch that associates toxin–antitoxin with CRISPR-Cas to harness engineered microbes. Our approach employs a CRISPR-repressed toxin–antitoxin (CreTA) module to make microbes addicted to the type I-F Cas effector proteins, and places both the Cas3 nuclease and the chromosome-targeting guide RNA under inducible promoters, thereby improving the genetic stability and stringency of the CRISPR-based suicidal program. Additionally, we have developed a single-plasmid, antibiotic-independent ATTACH device, which shows robust, stringent containment of a microbial chassis in murine gut, and negligible impacts on culture growth or lycopene production during batch fermentation. Our data highlight the potential of CreTA to stabilize CRISPR-based kill switches, advancing their development into more portable and reliable biocontainment tools for engineered microbes.

|
Scooped by
mhryu@live.com
March 12, 1:48 AM
|
The identification of post-translational modifications (PTM) of protein residues is vital for understanding protein functions and their subsequent role in cellular processes. Researchers have collected substantial experimental data on PTMs along with their validation and assortment into online databases. However, these databases do not offer integration with protein search and identification tools which then prevent researchers from seamlessly employing them in deciphering PTMs in biological samples. Additionally, there is a lack of tools that evaluate protein residues in light of their neighborhoods for the variety of PTMs found in PTM databases. Here, we propose PERCEPTRON-PTMKB, a web server designed for the analysis and evaluation of PTM sites using empirical datasets. PERCEPTRON-PTMKB provides a quantitative evaluation of per-residue PTM site through its propensity scoring algorithm. The search and propensity calculation pipelines are also served via a secure RESTful API, which enables their seamless integration into protein sequence search engines.
|
 Your new post is loading...
Your new post is loading...