Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 12:10 PM
|
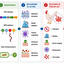
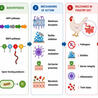
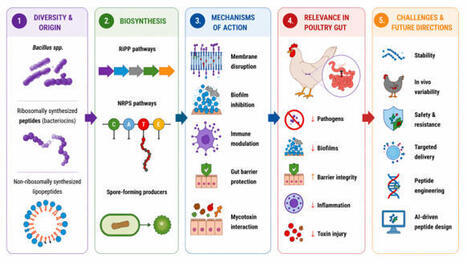
Bacterial enteric pathogens, antimicrobial resistance, and mycotoxin-associated intestinal injury remain important challenges in poultry-associated systems. In this context, Bacillus-derived antimicrobial peptides (AMP) have attracted attention as potential alternatives to conventional antibiotics due to their structural diversity and multifunctional properties. These peptides include ribosomally synthesized bacteriocins and non-ribosomally synthesized lipopeptides, such as surfactin, iturin, and fengycin. Their amphipathic structures enable interaction with microbial membranes, leading to permeabilization and disruption of cellular homeostasis. In addition to direct antimicrobial activity, these AMPs may interfere with biofilm-associated processes, modulate host immune responses, and help protect against toxin-induced epithelial injury. This review summarizes current knowledge on the diversity, structural characteristics, biosynthesis, mechanisms of action, and microbiological relevance of Bacillus AMPs in poultry-associated environments. Emphasis is placed on membrane targeting, biofilm regulation, immunomodulation, and mycotoxin-related gut protection, as well as limitations associated with antimicrobial resistance. Available evidence indicates that these peptides have diverse mechanisms of action; however, their activity is influenced by peptide class, formulation, microbial ecology, and host physiological factors. In addition, the potential for adaptive or genetically encoded resistance should be considered. Key translational challenges include peptide instability, variability in in vivo efficacy, strain-specific differences, safety considerations, and the lack of standardized comparative models. Future progress will depend on improved delivery systems, microbiome-resolved in vivo studies, and the integration of genomic mining, synthetic biology, and computational peptide design. These approaches may support the development of AMPs with improved stability, specificity, and functional performance in poultry-associated microbial systems.

|
Scooped by
mhryu@live.com
Today, 12:04 PM
|
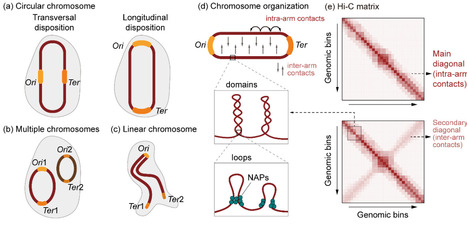
Bacterial chromosomes are compacted into hierarchical three-dimensional (3D) architectures that underpin essential processes such as replication, transcription, and segregation. However, detailed principles governing genome architecture and its regulation remain incompletely understood. Here we review current knowledge on structural organization of bacterial 3D genomes and their regulatory factors, integrating insights from chromosome conformation capture and imaging studies. We highlight how genome architecture dynamically responds to environmental cues. We also discuss potential applications of bacterial 3D genomics, particularly in synthetic biology. These findings provide a framework for understanding bacterial genomic homeostasis and adaptation, offering opportunities for engineering microbial systems and novel therapies.

|
Scooped by
mhryu@live.com
Today, 11:54 AM
|
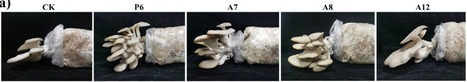
Some microorganisms present in the cultivation environment serve as biocontrol agents and contribute to enhanced mushroom production. However, the native bacteria naturally associated with commercial cultivation bags of Pleurotus ostreatus, as well as their growth-promoting roles and underlying mechanisms, remain poorly understood. This study aimed to identify native bacteria that promote Pleurotus ostreatus development and to investigate the underlying mechanisms. Four native bacteria, including Brevibacterium epidermidis (P6), Acinetobacter soli (A7), Pseudomonas parafulva (A8), and Pseudomonas hunanensis (A12), were isolated based on their ability to promote mycelial growth of P. ostreatus. B. epidermidis P6 shortened complete mycelial colonization time from ~30 d to 14 d in dual cultivation bags. All four strains increased fresh mushroom yield, with B. epidermidis P6, A. soli A7, and P. parafulva A8 increasing the number of basidiomata, while P. hunanensis A12 enhanced their size. These strains produced exopolysaccharides that enhanced mycelial growth. Additionally, B. epidermidis P6, A. soli A7, and P. parafulva A8 also secreted extracellular crude proteins that also promoted mycelial growth. Bi-plates and further gas chromatography–mass spectrometry analysis demonstrated that volatile organic compounds from P. hunanensis A12, including acetone, 2-butanone, benzaldehyde, and 1-undecene, enhanced fungal mycelial growth. The mycelial growth rates of Ganoderma lucidum and Pleurotus pulmonarius were also enhanced by these four strains. These results reveal that four native bacterial strains promote mushroom development through complex mechanisms.

|
Scooped by
mhryu@live.com
May 23, 11:01 PM
|
Microorganisms can use different enzymes to perform nitrite ammonification, the reduction of nitrite to ammonium, an important process to retain nitrogen in soils. Yet, the organisms mediating this process and their distribution in terrestrial ecosystems remain poorly resolved. Here, we determined the phylogenetic diversity of bacteria performing fermentative nitrite ammonification via the NAD(P)H-dependent nitrite reductase NirB, assessed their distribution across terrestrial ecosystems, and identified their environmental preferences. We found that these organisms are broadly distributed, spanning 29 phyla including Bacillota, Pseudomonadota and Actinomycetota. Screening 1,587 globally distributed soil metagenomes using a phylogeny-based approach revealed that fermentative nitrite ammonifiers are ubiquitous across biomes and particularly abundant in Mediterranean forests and desert soils. In these ecosystems, they outnumbered NrfA-dependent ammonifiers, the best characterized ammonifier group to date, suggesting distinct ecological niches for the two groups. Consistent with this, random forest modelling revealed a negative relationship between fermentative nitrite ammonifiers and the carbon-to-nitrate ratio, which contrasts with a preference for high carbon-to-nitrate conditions in NrfA-dependent ammonifiers. However, moisture and salinity emerged as the strongest predictors of the abundance of fermentative nitrite ammonifiers, indicating a high tolerance to osmotic stress in this group. Overall, our results demonstrate that fermentative nitrite ammonifiers are both phylogenetically diverse and environmentally widespread, calling for future efforts to determine the conditions under which they contribute to nitrogen retention in soils.

|
Scooped by
mhryu@live.com
May 23, 5:18 PM
|
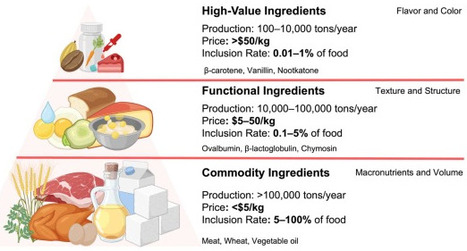
Precision fermentation has emerged as a promising platform for producing food ingredients. However, only a limited number of precision fermentation-derived ingredients have reached commercial markets. The concept of the minimum viable product (MVP) provides a useful framework for accelerating the practical adoption. MVP in precision fermentation is a food ingredient that provides a clear value proposition for a defined target consumer and can be safely and economically produced by an engineered microorganism. We introduce the Precision Fermentation Ingredient Pyramid, which can categorize targets into three tiers based on production volume, market value, and functional roles. A pipeline integrates market needs, biological and regulatory feasibility, economic competitiveness, and scalability for MVP selection. Applying this framework, we highlight opportunities across ingredient classes, including single-cell protein, ovalbumin, and β-carotene. Finally, we highlight the importance of techno-economic analysis for precision fermentation, particularly in linking strain performance and downstream processing to process economics.

|
Scooped by
mhryu@live.com
May 23, 5:05 PM
|
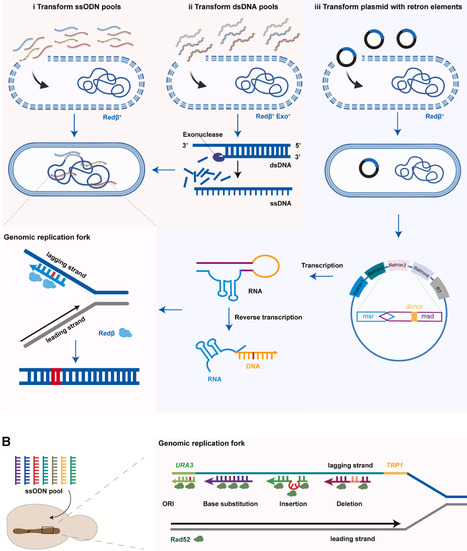
To facilitate the study of applied genetics and enable a rapid translation of genetic insights, it is highly desirable to concurrently modify many genomic loci in an organism of interest. While single-locus editing is well-established and technically straightforward, multiplex genome engineering (MGE) poses significant technological barriers. With the convergence of low-cost DNA synthesis, advanced genome editing techniques, and laboratory automation, a plethora of MGE methodologies were recently developed and applied in fields ranging from basic research to applied sectors. This review analyzes one-step and iterative MGE methodologies, with an emphasis on recombineering and CRISPR-Cas systems, and showcases emerging paradigm-shifting applications in biomanufacturing, agriculture, and therapeutics. We conclude by analyzing the limitations of existing technologies and discussing future directions for further optimizing MGE to solve system-level problems.

|
Scooped by
mhryu@live.com
May 23, 4:38 PM
|
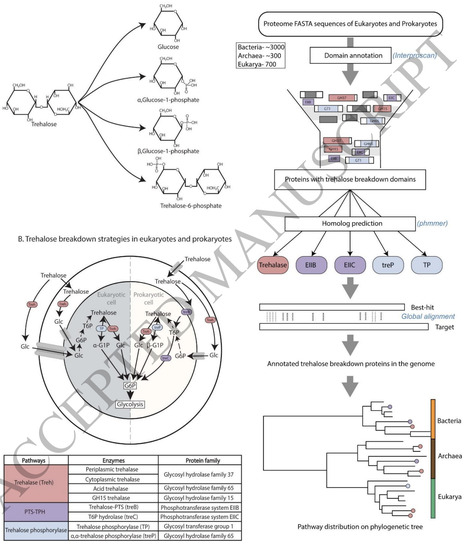
Trehalose is a widely prevalent disaccharide that acts as a cellular stress protectant, and functions as an energy source that enters central carbon metabolism when broken down. The evolution and distribution of trehalose breakdown pathways across kingdoms of life is not well studied, and therefore the ability of different organisms to consume trehalose as a carbon source is unknown. In this study, we build a comprehensive evolutionary analysis of the four known trehalose breakdown pathways - trehalase (acid, neutral, glycosyl hydrolase 15), trehalose phosphorylases (TP, treP), and trehalose specific phosphotransferases (PTS), by studying their distributions across ∼3800 prokaryotic and eukaryotic genomes. Our study suggests the presence of trehalase in the Last Eukaryotic Common Ancestor (LECA), and reveals near-universal presence of trehalase in eukaryotes, except in all birds where trehalase was lost in the first bird ancestor. Fungi alone retain additional trehalose phosphorylases (TP) in addition to trehalase. In contrast, trehalose breakdown in prokaryotes is highly sporadic but can occur via multiple, independently evolved pathways, including trehalase, the trehalose-specific PTS and trehalose phosphorylase. Finally, we observe that a subset of fast-growing Gammaproteobacteria retain the trehalose specific PTS, the loss of which reduces growth in E. coli. Overall, our findings uncover the evolutionary landscape of trehalose breakdown, and use of this versatile disaccharide as an energy reserve in different kingdoms of life.

|
Scooped by
mhryu@live.com
May 23, 4:28 PM
|
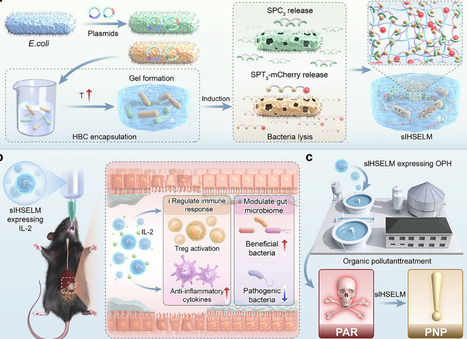
The practical application of engineered living materials (ELMs) is currently hindered by some critical challenges, such as streamlining fabrication processes and achieving long-term stability. Here, a semi-interpenetrating ELM was developed relying on thermosensitive self-assembly of hydroxybutyl chitosan (HBC) and spontaneous covalent protein interactions. This semi-interpenetrating network provided superior mechanical properties over HBC hydrogels. Furthermore, this material can be adapted for diverse scenarios based on engineered bacteria encapsulated, and its applications in biotherapy treatment and environmental remediation were validated. Compared to planktonic bacteria or enzymes, this ELM presented enhanced tolerance to harsh environments, including high temperatures, extreme pH values, high salinity, and digestive fluids, resulting in improved therapeutic efficacy with excellent biosafety in ulcerative colitis treatment and long-term degradation of the pollutant paraoxon. In summary, our material offers advantages including simple preparation, excellent mechanical properties, high stability, customizability, and biosafety, laying a foundation for the application of ELMs.

|
Scooped by
mhryu@live.com
May 23, 4:21 PM
|
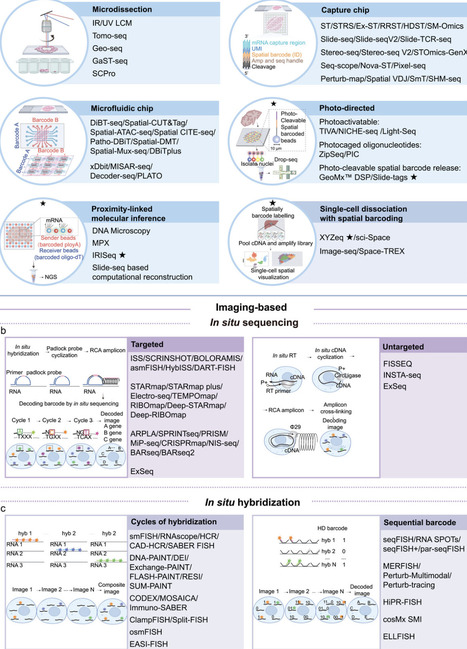
Biological macromolecules assemble into sophisticated spatial architectures to orchestrate fundamental cellular processes. Understanding this architecture is therefore essential for deciphering the mechanisms of life. Driven by advances in high-throughput sequencing and single-molecule imaging, spatial omics technologies have emerged as powerful tools that are revolutionizing biomedical research. This review systematically evaluates current spatial omics methodologies by comparing their key performance parameters. We critically assess their optimal applications and discuss strategies to overcome prevailing challenges in spatial resolution, capture efficiency, robustness, data analysis, and clinical translation. Furthermore, we highlight how these technologies provide unique insights into tissue heterogeneity, cell-cell interactions, developmental dynamics, microenvironmental composition, and neuroanatomy. Our analysis offers guidance for selecting appropriate spatial omics approaches and outlines promising directions for future technological innovation and expanded biomedical applications.

|
Scooped by
mhryu@live.com
May 23, 4:03 PM
|
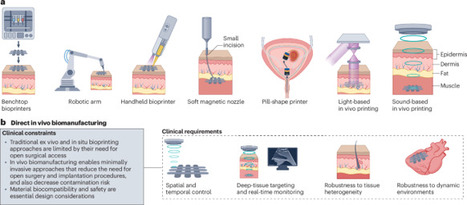
In situ biomanufacturing is redefining personalized and regenerative medicine by enabling therapeutic materials to be formed directly at target sites within the body. This emerging paradigm integrates advances in biomaterials, minimally invasive surgical tools and robotic systems, and ultrasound-guided activation platforms to achieve precise, on-demand material deposition or crosslinking without the need for open surgical access. This Review outlines the clinical challenges and requirements motivating these technologies — minimally invasive access, deep-tissue targeting, spatiotemporal precision and real-time monitoring — and describes how next-generation biomaterials, including shear-thinning, photoreactive, thermoresponsive and acoustically responsive hydrogels, are being engineered to meet these demands. Progress in fabrication strategies, ranging from hand-held and robotic bioprinters to near-infrared light-activated and ultrasound-activated systems capable of noninvasive or deep-tissue biofabrication, is summarized. Key applications in tissue regeneration, wound repair, localized drug delivery and in vivo bioelectronics are highlighted to illustrate the translational potential of these approaches. Finally, the major steps required for clinical adoption are described, including the development of materials optimized for in vivo activation, standardized evaluation frameworks, regulatory considerations for energy-activated platforms, and integration into clinical workflows. Together, these advances point towards a future in which patient-specific therapeutic structures can be fabricated directly within living tissues. Advances in material science have greatly extended the capabilities of 3D bioprinting. This Review discusses material design principles for in vivo biomanufacturing technologies and describes challenges and opportunities for the clinical translation of applications in tissue regeneration, wound repair, therapy and bioelectronics.

|
Scooped by
mhryu@live.com
May 23, 2:52 PM
|
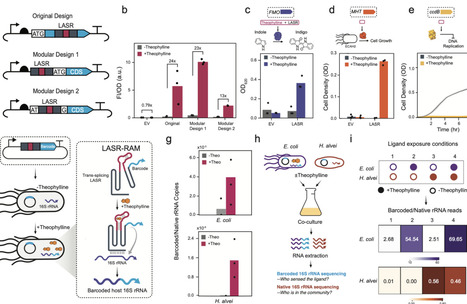
Domain insertion is an established method to engineer ligand-mediated control of activity in protein scaffolds. Whether this strategy can be systematically applied to large, structured RNAs remains unclear. In this study, we investigated the feasibility of engineering ligand-activated splicing ribozymes (LASRs) from group I catalytic introns. Using domain-insertion profiling coupled with high-throughput screening, we mapped the nucleotide-resolution landscape of aptamer insertion across the ribozyme and identified sites that support robust ligand-dependent control. We showed LASRs function across multiple kingdoms of life, including diverse species of bacteria and even fungi, and can be used to regulate various genetic outputs. Finally, we integrated LASRs with a genetic recorder that writes information into ribosomal RNA, enabling sequencing-based recovery of intracellular chemical signals from microbial consortia. This work establishes LASRs as an RNA-based inducible control platform for sensing diverse chemical inputs, regulating the expression of diverse genes of interest, and recording intracellular information.

|
Scooped by
mhryu@live.com
May 23, 1:45 PM
|
The fast-growing cyanobacterium Synechococcus sp. PCC 11901 is emerging as a promising chassis for photosynthetic biomanufacturing. Here we report recombinant protein production in PCC 11901 via signal peptide-mediated secretion, enabling direct recovery of target proteins from the culture medium without cell disruption. Seven signal peptides spanning both Sec and Tat pathways are screened using eYFP as a reporter, with secretion quantified daily over seven days by fluorescence measurements. FutA, belonging to the Tat pathway from Synechocystis sp. PCC 6803, achieves 92.2% extracellular export by day 7, substantially outperforming all Sec candidates, including the best Sec signal peptide thermitase from Cyanobacterium aponinum PCC 10605 (55.7%). Signal peptide-bearing strains exhibit growth reductions of up to 26% relative to the wild-type, with FutA most affected, indicating a general metabolic cost correlated with secretion efficiency. The best-performing signal peptides from both pathways, FutA and thermitase, are validated with secretion of lichenase. Notably, the rank order of signal peptide performance is reversed for lichenase: thermitase demonstrates 2.6-fold higher extracellular activity than FutA, indicating that optimal signal peptide selection is cargo-dependent. These results establish PCC 11901 as a secretion-competent chassis and provide a rational framework for matching signal peptide pathways to target protein properties.

|
Scooped by
mhryu@live.com
May 23, 1:37 PM
|
Genes for CO2 fixation occur in soil microorganisms, but little is known about the pathways that are most common across ecosystem types, the organisms with these genes, where different CO2 fixation pathways are most prevalent, and the energy sources that support autotrophy across ecosystems. Here, we investigated microbial capacity for autotrophy in soils using 853 metagenomes and 201 metatranscriptomes from a wide range of terrestrial ecosystems (agricultural soils, wetlands, weathering rock). Autotrophy-associated RuBisCO (Form I and II) is widely encoded across all soils and occurs in bacteria from numerous lineages (38 phyla). RuBisCO Form IE is consistently more phylogenetically diverse in soils than in marine ecosystems, suggesting that it may have evolved to function in soil-like environments. A newly discovered deeply branching Form I RuBisCO, Form I triple prime, supports the hypothesis that Form I RuBisCO originated in anaerobic environments. Further, saturated soils harbor more, and more distinct, autotrophic microbes, many of which may use the Calvin-Benson-Bassham cycle or Wood-Ljungdahl pathway for CO2 fixation. Overall, the results indicate that autotrophy is a particularly important metabolism in deep, saturated soils and weathering rock.
|

|
Scooped by
mhryu@live.com
Today, 12:08 PM
|
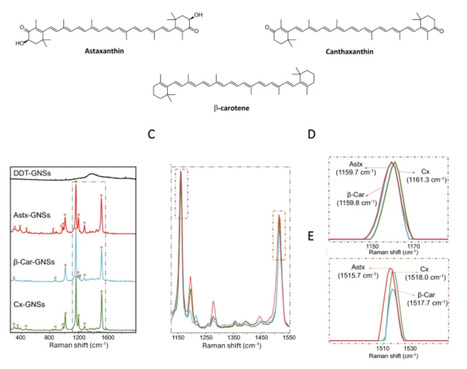
Carotenoids are natural tetraterpenoid pigments with important nutritional properties and broad industrial applications. Enhancing their production in plant-based biofactories offers a sustainable alternative to current manufacturing processes. In this work, we developed a label-free, single-cell analytical platform combining Raman imaging with a multi-layer perceptron neural network to classify tobacco BY-2 cells based on their carotenoid content. Carotenoid standards analysis, including astaxanthin, canthaxanthin, and β-carotene, was performed by surface-enhanced Raman scattering using hydrophobic gold nanostars due to the low concentration available. This analysis allowed the assignment of characteristic Raman peaks, specifically at 1160 cm−1 and 1520 cm−1, of key carotenoids and their identification inside of the cells by Raman imaging. The Raman fingerprints were correlated with carotenoid profiles obtained by HPLC, enabling accurate differentiation between wild-type and transgenic cell lines. In the analyzed transgenic lines, carotenoids accumulated in vesicle-like structures near the nucleus and along the cytoplasmic membrane. This method provides a non-destructive, label-free approach with high classification accuracy and sorting potential based on carotenoid composition, and may be a useful tool for plant synthetic biology and metabolic engineering.

|
Scooped by
mhryu@live.com
Today, 12:02 PM
|
Bacillus methanolicus MGA3 is a thermotolerant methylotroph that utilizes methanol, a renewable C₁ substrate, as its sole carbon and energy source. The strain naturally overproduces and secretes L-glutamate, making it a promising platform for engineering pathways toward L-glutamate-derived amino acids such as L-proline, which has applications in nutrition, stress protection, and industry. Heterologous expression of an osmotic stress–responsive L-proline biosynthetic operon from the mesophile Bacillus licheniformis in the thermotolerant B. methanolicus strain MGA3 did not increase L-proline levels but instead led to accumulation of L-citrulline. This was likely due to heat sensitivity of pyrroline-5-carboxylate reductase (ProH), the last enzyme of the osmoregulatory L-proline biosynthetic route, and metabolic crosstalk between L-proline and L-arginine pathways operating in Bacilli. To overcome these limitations, a synthetic operon containing the native anabolic proBA and proI L-proline biosynthetic genes from B. methanolicus MGA3 was engineered to remove transcriptional T-box regulation and biochemical feedback inhibition of ProB enzyme activity. Expression of this engineered operon enhanced L-proline synthesis and triggered its secretion during methanol-based growth of B. methanolicus MGA3 at 50° C. In fed-batch fermentation with methanol as carbon and energy source, extracellular L-proline levels reached 262 ± 20 mg L⁻1 after 40 h. During the fermentation process, a stepwise increase in medium osmolarity was observed, likely due to large-scale L-glutamate excretion, which impaired cellular growth. This study links osmolarity dynamics to methanol-based fermentation in B. methanolicus MGA3 and demonstrates its potential as a cell factory for L-proline and L-citrulline production. These findings support further strain optimization for producing value-added amino acids and highlight the relevance of methylotrophic thermophiles in sustainable biotechnology.

|
Scooped by
mhryu@live.com
May 23, 11:35 PM
|
The three-dimensional organization of the bacterial chromosome is critical for gene regulation. Chromosome conformation capture (Hi-C) has enabled genome-wide mapping of chromosomal folding, yet ensemble- averaged contact maps entangle biologically specific-folding interactions (SFIs) with nonspecific polymer compaction and mask single-cell heterogeneity. Here, we developed a polymer-based simulation framework in E. coli to address these limitations. We found that a null model of 300,000 random polymer configurations recapitulated the global chromosomal organization observed in Hi-C data, establishing that most Hi-C signals reflect generic polymer behavior in a confined volume. Contrasting null-model predictions with experimental Hi-C data isolated a small subset of SFIs (< 7%) and generated a specific-fold ensemble of 20,000 single-cell conformations that reproduced chromosomal interaction domains and single-cell heterogeneity. SFIs were enriched in the ter region, reduced nucleoid accessibility, colocalized with cryptic prophages, and depleted in positively supercoiled regions. H-NS and MatP emerged as major chromosome-wide and local determinants of SFIs, respectively. Furthermore, high-SFI regions correlated with stress-adaptive genes, whereas low-SFI regions harbored housekeeping genes. Together, our results established that a small number of biologically encoded SFIs superimposed on a polymer background shape the E. coli chromosome and gene expression, providing a quantitative framework for dissecting chromosome architecture and function.

|
Scooped by
mhryu@live.com
May 23, 5:22 PM
|
Rapid, specific quantification of erythritol in complex matrices remains challenging due to its structural similarity to sugars like sucrose. To overcome this, we developed a survival-fluorescence cascade screening circuit (uFAST) to engineer a selective, σ54-dependent BmoR-based biosensor. This circuit integrated SacB-mediated negative and mCherry-mediated positive selection to rapidly isolate erythritol-specific mutants. Guided by structural analysis, we optimized BmoR, achieving a 6.03-fold increase in GFP output and a 2.09-fold improvement in apparent affinity (Km). Coupled with a VioABCE-based visual module, the biosensor enabled accurate erythritol quantification in complex matrices, matching HPLC results. Integrated with an erythritol production module and fluorescence-activated droplet sorting (FADS), this platform screened an ARTP-mutagenized library, identifying a strain with 2.88-fold higher titer, yielding 3.51 g/L erythritol in fermentation. The uFAST circuit provides a generalizable framework for developing highly specific biosensors, offering an efficient tool for erythritol detection and strain improvement.

|
Scooped by
mhryu@live.com
May 23, 5:08 PM
|
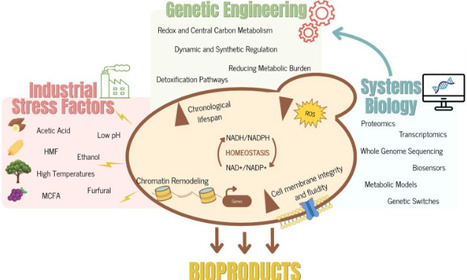
Yeast industrial bioproduction is often constrained by stress, substrate variability, and metabolic imbalance. This review provides a critical overview of recent advances in engineering yeast metabolism for robust performance under industrial conditions, increasingly defined by the use of renewable and heterogeneous feedstocks, which impose additional physiological and metabolic challenges. We highlight strategies that have gained particular relevance in the current literature, including the engineering of redox homeostasis and central carbon metabolism, detoxification and stress mitigation pathways, and the reduction of metabolic burden. The increasing contribution of systems biology and multi-omics, alongside dynamic and synthetic regulatory strategies, is also addressed. Finally, we explore the potential of naturally robust yeast chassis and outline future directions for resilient, sustainable, and scalable bioproduction systems.

|
Scooped by
mhryu@live.com
May 23, 4:56 PM
|
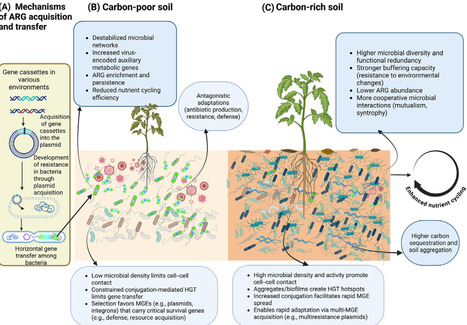
Antibiotic resistance genes (ARGs) are widespread in soils, yet their persistence is often viewed only through the lens of chemical selection. Here, we propose soil carbon as an integrative ecological driver structuring ARG dynamics in terrestrial systems. By shaping microbial growth, community assembly, colonization resistance, and horizontal gene transfer, soil carbon can either constrain ARG persistence or, under certain conditions, facilitate ARG spread.

|
Scooped by
mhryu@live.com
May 23, 4:33 PM
|
Cancer immunotherapy is increasingly moving toward personalized, precision-based strategies, with cancer vaccines emerging as a promising approach to reshape treatment. However, despite their potential, current tumor vaccines often yield limited clinical responses and subpar immunogenicity, underscoring the urgent need for innovative delivery systems to enhance immune activation. Bacterial outer membrane vesicles (OMVs), which possess natural immunomodulatory properties and impressive engineering flexibility, have attracted attention as versatile platforms for vaccine development and bioengineering applications. This review thoroughly summarizes recent advances in using OMVs to enhance the effectiveness of cancer vaccines. First, we explain the key biological features of OMVs that support their immunotherapeutic potential. Next, we carefully analyze the primary mechanisms by which OMVs enhance immune responses, as well as cutting-edge engineering strategies to improve their safety, immunogenicity, and specificity. Additionally, we discuss the significant challenges that hinder the clinical use of OMV-based cancer vaccines and provide a comprehensive review of current progress and future outlooks. Looking forward, combining artificial intelligence, tumor microenvironment profiling, and neoantigen discovery is expected to drive the development of next-generation, personalized OMV-based immunotherapies. Overall, OMVs stand out as a transformative platform capable of overcoming major obstacles in cancer vaccine development and pushing forward future cancer immunotherapy.

|
Scooped by
mhryu@live.com
May 23, 4:24 PM
|
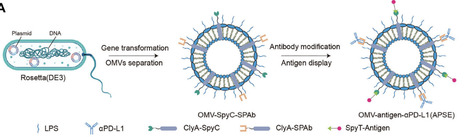
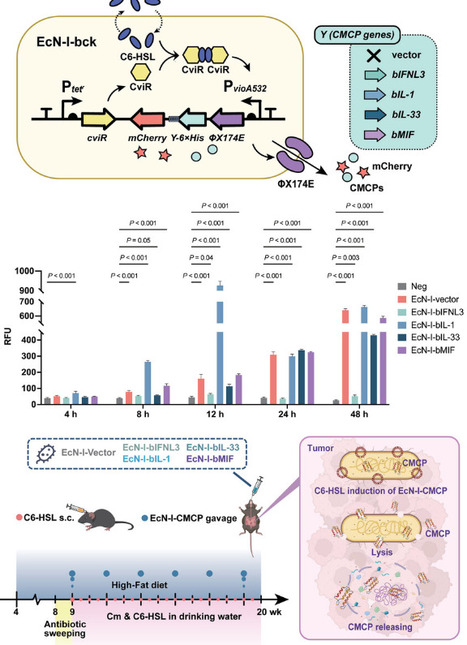
Accumulating evidence emphasizes the importance of microbiota–immune interactions in health and disease development, and identified bacteria-derived small-molecule metabolites as well as macromolecules such as peptides and proteins as promising therapeutic approaches. Here, we identify cytokine motif-containing, immunomodulatory bacterial proteins (CMCPs) as a special category of bacterial proteins in both bacterial genomes and gut metagenomes using Hidden Markov Models (HMMs). We further find eight colorectal cancer‑associated CMCPs differentially enriched in patients or healthy controls. Engineered E. coli Nissle 1917 (EcN) expressing selected CMCPs administered to Apcmin/+ mice selectively colonize intestinal tumors, deliver functional CMCPs in situ, and elicit significant antitumor immune responses while reducing tumor burden. In vitro, purified CMCPs modulate mouse splenic T cells, bone marrow‑derived macrophages and dendritic cells. Our findings indicate that bacterially encoded CMCPs can directly modulate tumor immunity and serve as microbiota‑derived proteins as candidate immunomodulators, which can further be applied in microbiome-mediated immune therapies for CRC.

|
Scooped by
mhryu@live.com
May 23, 4:09 PM
|
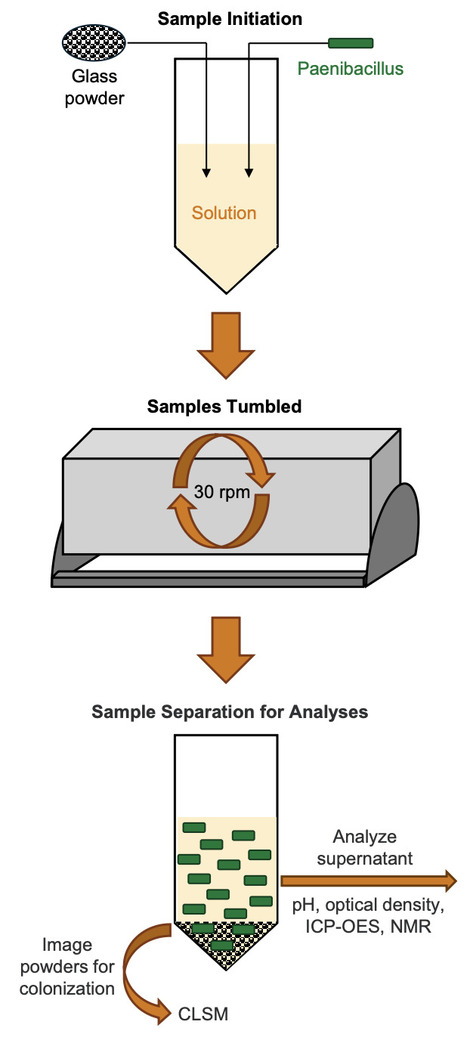
Silicate glasses are an accepted option for immobilizing nuclear waste and waste glass can be disposed in near-surface environments. It is important to understand glass alteration mechanisms under site-relevant conditions to predict glass corrosion rates upon disposal. Microbial activity near the glass surface may influence glass alteration. However, waste glass chemical durability is currently evaluated without consideration of microbial alteration. Here, four glass compositions were tested in three solutions, with and without a subsurface Paenibacillus bacterium, to compare the extent of glass leaching. Results indicate that the Paenibacillus cells increased glass alteration, resulting in higher concentrations of boron, iron, sodium, and silicon released into solution. The combination of microbially mediated organic acid production, which decreased pH, and glass dissolution, which increased pH, resulted in a net neutral or slightly acidic solution that could promote further glass alteration. The amount of each element released depended on glass composition and solution chemistry. This study revealed the dynamic relationship between microbial metabolism, elemental release, and corresponding changes to solution pH, showing that microbial processes can indirectly accelerate glass alteration. This work supports a greater understanding of microbially-influenced glass alteration and informs development of standardized durability tests to assess microbial influence at disposal facilities.

|
Scooped by
mhryu@live.com
May 23, 2:57 PM
|
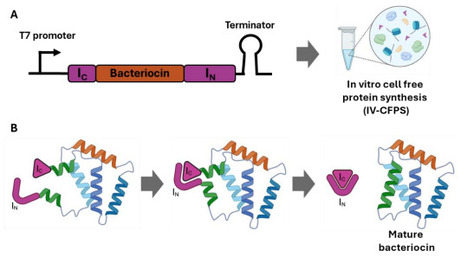
Bacteriocins are ribosomally synthesized antimicrobial peptides with promising applications in biotechnology, particularly in food preservation and animal and human health. Circular bacteriocins are especially attractive due to their head-to-tail cyclized structure, which confers enhanced stability and antimicrobial potency relative to linear peptides. Here, we report an in vitro cell-free protein synthesis system coupled with an enhanced Split Intein-Mediated Ligation platform (IV-CFPS/SIML) for the efficient synthesis of circular bacteriocins through systematic evaluation of cyclization sites and alternative split inteins. Using enterocin AS-48 as a model, we systematically evaluated multiple serine-based cyclization sites in combination with three split inteins, NpuDnaE, Gp41-1, and SspGyrB, to identify configurations supporting efficient splicing and high antimicrobial activity. Gp41-1 emerged as the most effective intein and was subsequently applied to the production of garvicin ML, amylocyclicin, and 27 naturally occurring sequence variants, demonstrating that cyclization site selection, intein identity, and minor sequence variations strongly influence antimicrobial potency and target range. Finally, SIML expression cassettes encoded in pUC-derived vectors enabled in vivo production and functional expression of selected circular bacteriocins in recombinant E. coli. Collectively, these results establish SIML as a versatile platform for in vitro and in vivo production, screening, and functional characterization of known and putative circular bacteriocins.

|
Scooped by
mhryu@live.com
May 23, 2:01 PM
|
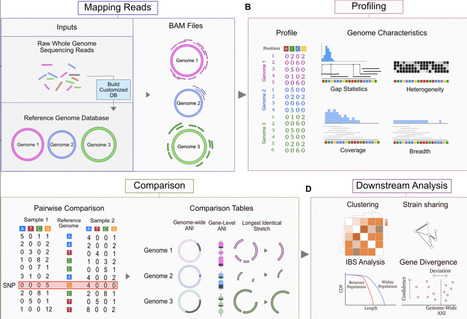
Strain-resolved metagenomics characterizes microbial communities at nucleotide-level resolution, enabling researchers to differentiate identical from closely related organisms and characterize population structure and gene content variation. Here we introduce ZipStrain, a program that performs highly accurate strain-resolved metagenomics over 500 times faster than available methods while offering superior RAM management. Applied to a dataset of 2,754 samples spanning human populations, we identify a strain-sharing gradient across social relationships, reveal striking variation in clonal structure across bacteria and bacteriophage, and pinpoint genes whose nucleotide identity deviates from genome-wide expectations. ZipStrain is distributed as an open-source Python package and accompanying Nextflow pipeline at https://github.com/OlmLab/ZipStrain.

|
Scooped by
mhryu@live.com
May 23, 1:43 PM
|
Flagella are large transenvelope nanomachines but how they transit the peptidoglycan in Gram positive bacteria is poorly understood. A recent model suggested that flagellar basal bodies diffuse in the membrane and become captured at locations in the peptidoglycan with a pore diameter that could accommodate the axle-like flagellar rod. Mutation of penicillin binding protein 1 (PBP1/PonA), a cell wall repair protein thought to decrease peptidoglycan pore frequency and/or size, resulted in a severe growth defect and cell lysis in the ancestral strain of Bacillus subtilis that was dependent on flagellar synthesis. Genetic analysis indicated that toxicity was due to completion of the flagellar hook, which activated the flagellar sigma factor SigD. SigD, in turn, activated a suite of peptidoglycan hydrolases that caused cellular lysis when PBP1 was absent. In addition, mutations that resulted in high levels of the stress response factor Spx could lessen the toxicity, while PBPX, a putative teichoic acid D-alanylase, was required for autolysis. In sum our results indicate that flagellar synthesis, not normally associated with cell viability, causes cell wall stress and under some conditions, cell death. Moreover, our work indicates that cost of envelope integrity by flagellar synthesis may be underappreciated due to strain domestication, and suggests that specialized systems may compensate for the cost of assembly of transenvelope machines in general.
|
 Your new post is loading...
Your new post is loading...