 Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 2:08 PM
|
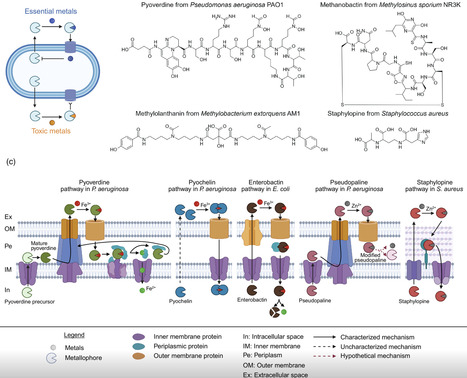
Metals are essential trace elements for almost all organisms including bacteria. Yet, metals are toxic at high concentrations, requiring fine-tuned regulatory mechanisms to steer metal homeostasis inside cells. In this primer, we explain how bacterial metallophores – small secreted secondary metabolites – act as gatekeepers by carefully orchestrating the scavenging and uptake of essential metals whilst preventing intracellular toxicity and keeping toxic metals outside the cell. We further introduce metallophore diversity together with main synthesis, secretion and uptake mechanisms. Finally, we show how secreted metallophores shape ecological interactions between bacteria and with eukaryotic organisms and how fundamental research on metallophores opens promising avenues for therapeutic and biotechnological applications.

|
Scooped by
mhryu@live.com
Today, 1:58 PM
|
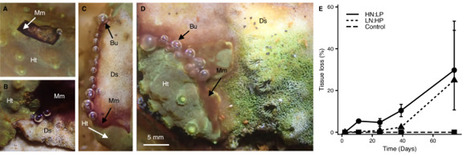
Coral diseases are increasing in prevalence, accelerating the global decline of tropical reefs, which threatens over 25% of marine biodiversity and vital ecosystem services for human societies. While outbreaks are frequently linked to environmental change, including heat stress, sedimentation, and reduced water quality, the mechanisms by which such factors promote disease remain poorly understood. Here we show that nutrient stress, caused by skewed seawater nitrogen-to-phosphorus (N:P) stoichiometry, promotes the onset of Black Band Disease (BBD), a common and easily recognisable syndrome that affects corals around the globe. Using Turbinaria reniformis as a model system, controlled laboratory experiments demonstrate that skewed N:P ratios disrupt the functional integrity of coral-associated microbial networks while favouring opportunists that exploit dysfunctional host–symbiont interactions. Disease lesion-associated microbial mats are dominated by cyanobacteria and include sulphur-metabolising bacteria, hallmarks of natural BBD communities. Strikingly, similar cyanobacterial taxa are also detected in the visually healthy coral tissue ahead of the expanding lesions, suggesting an opportunistic recruitment of disease-associated members from the resident microbiome. Global analyses of BBD outbreaks reveal that over 88% occurred in regions with skewed N:P ratios, compared with only 16% that were linked to prior heat stress. Together, our findings identify nutrient-driven microbiome destabilisation as a key pathway to coral disease, reinforcing nutrient management as a major lever for reef conservation and restoration practice. Coral diseases contribute to the decline of reefs around the globe. This study reveals that disruptions of the nutrient balance in seawater can change coral-associated microbial communities leading to disease.

|
Scooped by
mhryu@live.com
Today, 1:34 PM
|
The biological significance of the transition metal molybdenum (Mo) lies in its function at the catalytic center of several enzymes that drive a wide spectrum of redox reactions underlying global biogeochemical cycles, yet a paradox persists. While modern life ubiquitously relies on Mo, geochemical evidence suggests that its availability in early Earth’s anoxic oceans was extremely limited. Modern organisms can use Mo down to trace levels; however, the rates of Mo-dependent metabolisms slow down when Mo availability decreases, posing fundamental questions about the extent to which changing Mo abundances shaped the evolution of molybdoenzymes, and when early life began harnessing Mo. Here, we confront this evolutionary enigma by reconstructing the temporal and ecological emergence of molybdoenzymes, their transport systems, and biosynthetic pathways. In parallel, we examine biological tungsten (W) usage due to shared chemical properties and cofactor biosynthetic pathways with Mo. We provide molecular dating evidence of Mo/W utilization back to the Eo- to Mesoarchean (~3.7–3.1 Ga). These findings challenge prevailing assumptions about trace metal availability on the early Earth and underscore the profound antiquity and adaptability of Mo-based biochemistry in shaping early microbial evolution. The study shows that life began using molybdenum and tungsten enzymes as early as 3.7–3.1 billion years ago. Study reveals that key metabolic processes arose despite scarce metals and highlighting the early adaptability of microbial life.

|
Scooped by
mhryu@live.com
Today, 1:31 AM
|
Marchantia polymorpha has emerged as a promising model system for investigations in plant synthetic biology. Quantitatively characterizing plant genetic elements is fundamental to achieving predictable and controlled gene expression. However, only a few genetic parts are currently available for Marchantia. Additionally, the characterization of gene expression elements still relies on stable transformation assays. Here, we developed an Agrobacterium-mediated transient expression system to rapidly evaluate genetic parts in Marchantia. The entire experimental workflow can be completed within 8 days. Using this high-throughput system, we systematically benchmarked 21 promoters, 15 terminators, and 7 signal peptides from diverse sources. We identified a truncated CaMV35S promoter variant (P_35S-3), a native terminator (T_MpAct1), and a heterologous signal peptide (SP_SdMir) as top-performing elements. Notably, the P_35S-3 promoter exhibited a 409-fold activity increase over the standard CaMV35S promoter P_35S. Utilizing this potent element, we achieved an eGFP protein yield of 319.8 μg/g fresh weight in stable transgenic lines. The reliability of this transient system was further validated by stable transformation, where the two signal peptides exhibited a relative performance consistent with the transient assays. Our transient expression system provides a rapid and efficient platform for the characterization of gene expression elements, thereby expanding the genetic toolkit for Marchantia and enhancing protein expression levels.

|
Scooped by
mhryu@live.com
Today, 1:24 AM
|
Biomolecular condensates are formed through liquid–liquid phase separation (LLPS). They are highly dynamic, membraneless compartments within cells. The liquid-to-solid transition (LST) of these condensates plays a central role in regulating cellular physiological functions, maintaining tissue structural stability, and driving disease progression. Engineering LST has emerged as a major research frontier, integrating biophysics, synthetic biology, and materials science. This review systematically outlines the molecular grammar governing LST, key engineering strategies for its spatiotemporal control, and emerging applications in designed biological systems. We further discuss current challenges and future directions for harnessing LST as a design principle in systems chemistry and synthetic biology.

|
Scooped by
mhryu@live.com
Today, 1:02 AM
|
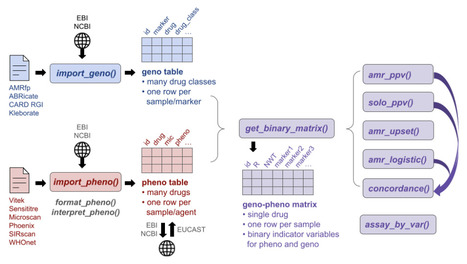
Microbial whole-genome sequence data is now generated at scale, including to support antimicrobial resistance (AMR) surveillance and understand resistance mechanisms, yet analytical infrastructure for systematically linking AMR genotypes to measured phenotypes remains fragmented. Here we present AMRgen, an R package to support systematic AMR genotype-phenotype analysis. AMRgen imports and harmonises genotypic data from common bioinformatics tools, alongside phenotypic data from automated antimicrobial susceptibility testing instruments and public repositories. It supports common analyses linking data to reference distributions, modelling associations, quantifying concordance, and producing publication-ready visualizations including UpSet plots that jointly display genotypic marker combination frequencies and associated phenotypic distributions. We demonstrate AMRgen's utility using publicly available surveillance data for World Health Organization priority AMR pathogens, Neisseria gonorrhoeae, Klebsiella pneumoniae, Escherichia coli and Salmonella enterica. AMRgen, available free and open-source at https://AMRgen.org provides a reproducible end-to-end foundation for genotype-phenotype research in AMR genomics, clinical microbiology, and public health surveillance.

|
Scooped by
mhryu@live.com
Today, 12:55 AM
|
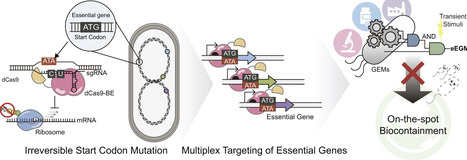
The environmental and therapeutic application of genetically engineered microorganisms necessitates the development of robust, irreversible biocontainment systems. In this study, we present an eEGM (editing-driven essential gene multiplex inactivation) module that utilizes CRISPR-mediated cytidine base editing to induce permanent self-killing via a single transient induction. By targeting the start codons of essential genes, we achieved an irreversible translational blockade that avoids the fitness costs associated with basal toxicity in nuclease-based systems. Multiplexed targeting of non-redundant essential loci (holA, ftsB, and dfp) yielded escape frequencies at or below the NIH guideline criterion (10−8) within 1 h of pulse induction. Furthermore, the eEGM system exhibited robust functional orthogonality and portability across laboratory, industrial, and therapeutic E. coli strains, including MG1655, W3110, and Nissle 1917, without detectable interference with heterologous protein expression. This work establishes base editing as a cleavage-free CRISPR effector for pulse-activated, irreversible biocontainment and provides a practical framework for safer deployment of engineered microbes.

|
Scooped by
mhryu@live.com
Today, 12:46 AM
|
Hexamethylenediamine (HMD), adipic acid, and ε-caprolactam (ε-CL) are essential C6 monomers used in the production of nylon 6,6 and nylon 6. Developing sustainable, bio-based routes to these compounds remains challenging due to pathway complexity. Here, we report a modular E. coli platform for the de novo biosynthesis of all three monomers directly from glycerol. We divided the overall pathway into upstream and downstream modules, with the upstream module converting glycerol to adipic acid. To construct downstream module, two distinct strains were engineered to individually convert adipic acid into HMD or ε-CL. Both strains employed carboxylic acid reductases Macar from Mycobacteroides abscessus and Mmocar from Mycolicibacterium moriokaense, with the latter identified and validated in this work. Specifically, HMD biosynthesis incorporated aminotransferases PatA from E. coli, GabT from Streptomyces avermitilis, and the introduced Bcta from Burkholderia cenocepacia. ε-CL biosynthesis utilized a similar upstream pathway but relied critically on a lactamization step catalyzed by an HLadh–Smnox fusion enzyme containing a flexible linker for efficient NAD+ regeneration. The common precursor, adipic acid, was produced by an upstream strain optimized through reverse β-oxidation pathway reconstruction, PaaJ engineering, and metabolic flux balancing, achieving a titer of 6.1 g/L. In fed-batch fermentation, cocultivation of the engineered strains with delayed inoculation enabled temporally coordinated conversion of glycerol to HMD (230.9 mg/L) and ε-CL (808.0 µg/L), representing low yet the highest titers reported to date. This work opens up the possibility of a unified, modular microbial platform for the sustainable production of nylon monomers from a renewable carbon source.

|
Scooped by
mhryu@live.com
Today, 12:28 AM
|
Despite major advances in biomedical research, dissecting disease-relevant molecular pathways remains challenging due to pathway redundancy, transient protein interactions, and the limited spatiotemporal precision of existing tools. Genetic code expansion (GCE) addresses these limitations by enabling site-specific incorporation of noncanonical amino acids that endow proteins with novel chemical, photophysical, or regulatory properties directly in living systems. This capability provides unique access to dynamic protein interactions, post-translational modifications, and signaling events in cellular environments. Here, we highlight recent advances in GCE that are particularly adapted to studying disease biology in increasingly physiologically relevant contexts, discuss key challenges limiting broader implementation, and outline emerging methodologies that position this technology as a transformative synthetic biology platform for mechanistic dissection of disease processes.

|
Scooped by
mhryu@live.com
Today, 12:06 AM
|
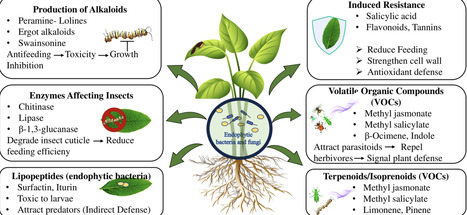
Plant families generate distinct repertoires of specialized metabolites that govern their biotic interactions. Endophytes strengthen host plant defence mechanisms and tolerance to biotic challenges by upregulating metabolite biosynthesis, modifying precursor compounds into more potent forms, or by directly synthesising analogous defence metabolites themselves. This review examines the complex relationships between endophytes and their host plants, with a focus on their colonization strategies, interactions, and contribution to plant immunity. It explores the diverse mechanisms through which endophytes augment plant defences, including the production of specialized metabolites, induction of systemic resistance, and direct antagonism against pathogens and herbivores. Accordingly, this study provides a comprehensive account of role of endophytes in protecting plants against multiple biotic stressors, rather than isolated threats. It further addresses a critical knowledge gap by highlighting how synergistic interactions between endophytic microbes and plant family-specific specialised metabolites shape plant immunity. By examining these unique plant-endophyte interactions across different plant families, the review offers deeper insight into the ecological and functional significance of endophytes in plant defence. Thus, it paves the way for targeted applications of endophytes in sustainable agriculture, where specific microbial strains can be harnessed to naturally improve plant protection and productivity.

|
Scooped by
mhryu@live.com
May 4, 11:19 PM
|
CRISPR-based nucleic acid diagnostics have shown broad potential, yet reliable single-nucleotide variant (SNV) discrimination remains limited by flanking sequence requirements that constrain targetability, and an inherent specificity-sensitivity trade-off where mismatch designs used to suppress wild type recognition often penalize enzymatic activity. Here we develop a scenario-guided Cas13d framework that supports pre-defined operating modes tailored to distinct analytical goals. Leveraging the minimal protospacer flanking site constraints of Cas13d, we first map mismatch-sensitive windows to derive rule-based crRNA designs that improve allelic discrimination. We then restore assay performance through structure-guided engineering of a miniaturized Cas13d scaffold by internally inserting auxiliary RNA binding domains (RBDs). Systematic benchmarking across representative oncology hotspots delineates two practical regimes comprising an ultra-sensitive, amplification-free mode in which a dual-RBD variant paired with optimized mismatched crRNAs achieves ∼0.6% variant allele fraction (VAF) detection, and a robust amplified mode incorporating optional loop-mediated isothermal amplification coupling that favors simpler architectures to balance performance and background across broader low-VAF ranges. In an evaluation of 45 clinical tumor RNA specimens spanning pancreatic, cholangiocarcinoma, and colorectal cancers, the assay correctly classified mutation status with full concordance for KRAS G12D, IDH1 R132C and BRAF V600E, with a subset of positive cases corroborated by orthogonal RT-ddPCR. A prospective IDH1 R132C clinical-matrix spike-in further supported sub-1% detection without pre-amplification. Collectively, this work establishes a configurable Cas13d toolkit and a rule-guided strategy for deploying CRISPR-based RNA SNV diagnostics with application-specific performance objectives.

|
Scooped by
mhryu@live.com
May 4, 7:44 PM
|
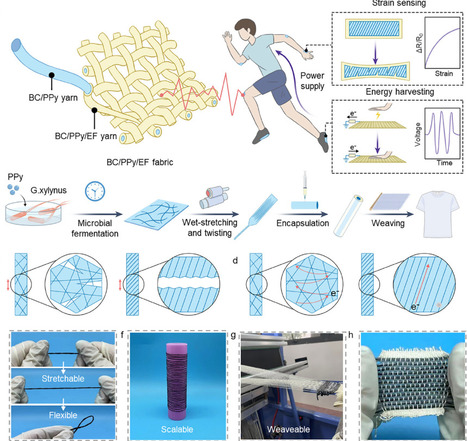
Invoking high-performance bio-based fibres (e.g., Bacterial cellulose) contributes to the sustainability and functionality of wearable electronic devices at the material level. However, the fabrication of self-powered and high-mechanosensitive stretchable BC-based sensors is challenging due to the difficulty in adaptable soft-rigid triboelectrical interfaces and obtaining ordered conductive bacterial cellulose fibres. Here, inspired by the spiral construction from biological systems, we develop an innovative bio-fabrication strategy to develop a core-sheath yarn that features the ordered network and mechanosensitive twisting structures. The yarn sensor integrates the complementary advantages of triboelectric and resistive responses for the integration of strain sensing and energy self-sufficiency. Converging factors of core-sheath structure, modulus-mismatch-governed elongation, and network cracks give the yarn sensor a sensitive mechanosensitive response (8.246), a wide strain range (up to 100%), and high voltage signals (over 50 V). The scalable self-powered fabrics based on yarns are also used as wearable power generation and energy storage for charging the yarn sensing system, achieving continuous health monitoring. The design of the unique structure assists the BC-based sensors to effectively energy charging and driving healthy monitoring system. These empirical insights from bio-manufacturing techniques to structural design of ordered yarns pave the way to obtaining multi-functional high-performance bio-based sensors.

|
Scooped by
mhryu@live.com
May 4, 7:10 PM
|
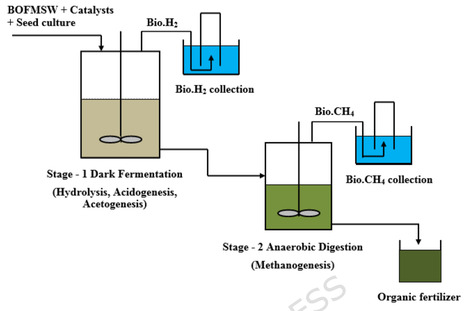
This study presents an integrated waste-to-energy strategy for the sustainable conversion of the biodegradable organic fraction of municipal solid waste (BOFMSW) into biohydrogen (Bio.H2) and biomethane (Bio.CH4) through a two-stage continuous stirred dark fermentation (DF) process. The first-stage bioreactor was inoculated with Clostridium Thermocellum selectively enriched in a 2-bromoethanesulfonic acid (BESA) medium, and the influence of bimetallic ion catalysts NiCl2 + FeCl2 and NiCl2 + FeSO4 was evaluated at various concentrations (25, 50, 75, and 100 mg/L). The catalyst combination NiCl2 + FeCl2 at 75 mg/L produced the maximum Bio.H2 yield of 3162 L, representing a 69% enhancement compared with the catalyst-free substrate. At this optimal catalytic concentration, the percentage of H2 in the gas composition was 69.26%. The second-stage bioreactor utilized the effluent from the first stage for Bio.CH4 generation, achieving the highest cumulative yield of 729 L at 50 mg/L NiCl2 + FeCl2, which was 59% higher than that of the catalyst-free substrate. The two-stage process achieved an overall COD removal efficiency of 93.18%, demonstrating the system’s effective capacity for energy recovery. Fourier Transform Infrared (FTIR) and Field Emission Scanning Electron Microscopy (FESEM) analyses confirmed the biochemical and morphological degradation of complex organics into volatile fatty acids, illustrating efficient substrate conversion and microbial proliferation. The final digested slurry, rich in nitrogen, phosphorus, and potassium, was found suitable for use as a bio-fertilizer, supporting nutrient recycling and soil enrichment. This integrated process not only improved energy recovery efficiency but also achieved near-zero waste discharge, combining waste-to-energy and waste-to-resource approaches. The developed system demonstrates strong potential for pilot-scale implementation of Bio.H2, and Bio.CH4 for co-production from municipal solid waste (MSW), offering a circular bioeconomy pathway toward low-carbon, sustainable urban waste management.
|

|
Scooped by
mhryu@live.com
Today, 2:06 PM
|
xBind is an interactive, freely accessible, and fully configurable webserver for large language model (LLM)-enabled cross-molecular protein binding-site prediction. xBind leverages LLM embeddings from the ESM-2 model together with sequence- and structure-derived features to predict protein–protein, protein–DNA, and protein–RNA binding sites using symmetry-aware deep graph neural networks. The input to xBind is either a single-chain protein sequence in FASTA format or a monomer protein structure in PDB or mmCIF format and it outputs predicted residue-level binding sites of the input protein with its pre-selected interaction partner. The customizable xBind web interface provides: (i) choice of interaction partners including protein–protein, protein–DNA, and protein–RNA; (ii) on-the-fly AlphaFold-based protein structure prediction for sequence-only inputs; (iii) on-demand selection of the likelihood threshold for calibrating structure-aware binding site annotations; (iv) interactive and interpretable web-based results, including sequence and structural visualizations and plots of residue-level binding likelihoods with user-adjustable threshold calibration; and (v) extensive help information for usage and results interpretation through a web-based tutorial and guide. xBind is freely available at https://fusion.cs.vt.edu/xBind.

|
Scooped by
mhryu@live.com
Today, 1:40 PM
|
Elucidating gene function in highly redundant genetic programs such as signaling pathways is challenging in model and nonmodel plants with current whole-plant genetic screening tools. Many of these challenges could be overcome if screens were instead carried out using individual cells harboring genetic perturbations. Here we report a single-cell screening platform, PIVOT (protoplast isolation after virus overexpression in planta), to accelerate identification and functional characterization of plant genes. We use Nicotiana benthamiana as a heterologous host to test gene libraries arrayed in a single leaf. PIVOT harnesses viral superinfection exclusion to ensure single multiplicity of infection per cell during pooled library delivery. Additionally, we engineer a cell-surface protein as a phenotypic marker for isolating cells of interest from a heterogeneous population. Using this system, we recover regulators of cytokinin signaling from an Arabidopsis open reading frame library. We anticipate PIVOT will be broadly applicable for high-throughput, single-cell functional genetic screening across the plant kingdom. A pooled, cell-based, genetic screening platform in plants is used for the functional analysis of cytokinin signaling proteins.

|
Scooped by
mhryu@live.com
Today, 1:16 PM
|
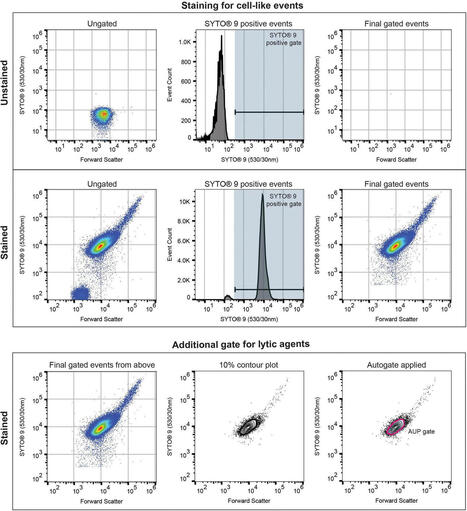
The global rise of multidrug-resistant bacteria necessitates the development of new antimicrobials and faster diagnostic tools. Conventional antimicrobial susceptibility testing is slow, relying on culture-based methods that delay effective treatment, often with fatal consequences in severe infections. In this study, we evaluate flow cytometry as a rapid, culture-minimal method to assess bacterial responses to six antimicrobials: ceftazidime-avibactam, meropenem-vaborbactam, cefiderocol, doxycycline, omadacycline, and lefamulin. Across 165 evaluable antibiotic-isolate combinations, essential agreement between flow cytometry and broth microdilution minimum inhibitory concentrations was 90.71%. Assessable categorical agreement, determined using the European Committee on Antimicrobial Susceptibility Testing and Clinical and Laboratory Standards Institute breakpoints, was 92.59% for doxycycline, 91.67% for omadacycline, and 100% for meropenem-vaborbactam. Cefiderocol exposure was associated with substantial cell elongation, demonstrating cellular-level antimicrobial effects observed using confocal microscopy and imaging flow cytometry. These findings demonstrate the potential of flow cytometry for novel antimicrobial evaluation, offering rapid insights into drug efficacy with potential to improve clinical outcomes in patients.

|
Scooped by
mhryu@live.com
Today, 1:29 AM
|
Does taking probiotics really matter? The idea is enticing. Swallow a capsule, add helpful microbes, support immunity, and strengthen the gut. Yet the microbiome is not a vacant landscape waiting for reinforcements. It is a densely woven ecosystem that behaves like an old-growth rainforest. Every niche is filled, every interaction balanced through biochemical negotiation, and any newcomer must face strong colonization resistance. With such a fortified system, what impact can a probiotic truly make? Most strains pass through the adult gut without becoming permanent residents. Still, they are not biologically inconsequential. During transit, they can influence epithelial barrier integrity, alter short-chain fatty acid and bile acid profiles, modulate immune signaling, and participate in cross-feeding interactions that reshape metabolic activity. These effects are best understood as functional ripples rather than structural reconfiguration. Accordingly, probiotic efficacy often reflects transient biochemical and host–microbe interactions, although the balance between transient activity and durable colonization depends on strain and formulation, dosing duration, host factors, and the baseline microbiome ecosystem, including recent disturbances such as antibiotics. Probiotic efficacy should therefore be evaluated using outcomes aligned with the intended mechanism, prioritizing clinical endpoints and biomarkers, supported by complementary compositional and functional microbiome readouts.

|
Scooped by
mhryu@live.com
Today, 1:22 AM
|
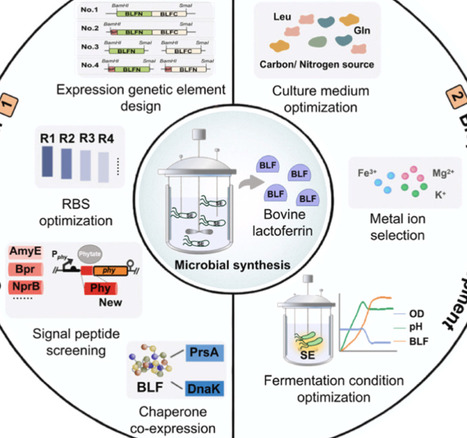
Bovine lactoferrin (BLF), a high-value multifunctional protein renowned for its iron-binding capacity and broad-spectrum antimicrobial properties. Despite its potential as a functional food ingredient, industrial application is restricted by the low abundance and high cost of extraction from natural sources. To address this bottleneck, this study establishes the first successful secretory expression of recombinant BLF in Bacillus amyloliquefaciens, a Generally Recognized as Safe (GRAS) host. Initially, the engineered strain yielded a titer of 18.0 ± 0.3 mg/L. To enhance secretion efficiency, a systems engineering optimization strategy was employed. This involved optimizing the ribosome binding site (RBS) and fusing a novel phytase-derived signal peptide (SPphy) to facilitate translocation. Crucially, the co-expression of the molecular chaperone PrsA was implemented to alleviate folding stress. These modifications culminated in a 6.1-fold yield increase, achieving a final titer of 110.0 ± 0.8 mg/L in a 5-L bioreactor. This research not only demonstrates the feasibility of B. amyloliquefaciens as a robust chassis for BLF production but also provides a strategic framework for the heterologous biomanufacturing of other complex nutrient proteins.

|
Scooped by
mhryu@live.com
Today, 12:58 AM
|
Agriculture is a major source of anthropogenic greenhouse-gas emissions, being the largest source of nitrous oxide (N2O), an extremely potent greenhouse gas and ozone-depleting agent. Soil N2O emissions are largely driven by microbial nitrification, in which ammonia-oxidizing microorganisms catalyze the rate-limiting oxidation of ammonia to nitrite. Nitrification not only mediates N2O fluxes but also reduces fertilization efficiency and contributes to eutrophication through nitrate leaching. Bacteriophage-based control of microbial communities is rapidly garnering interest in a number of fields; however, phages infecting ammonia-oxidizers are largely uncharacterized, with only one lytic phage having been described, limiting the potential for phage-mediated nitrification inhibition. Here, we show the largest set of phages infecting ammonia-oxidizing bacteria (AOB) to date: 45 dsDNA phages identified from urban wastewater, infecting four AOB species, with 16 demonstrating cross-genus host ranges and capable of eliminating nitrification activity in liquid cultures. Phylogenetic and taxonomic analyses revealed six proposed families of Caudoviricetes and numerous monophyletic clades, likely representing higher-level lineages. Structure-guided genome annotation revealed these phages to carry diverse and seldom-seen auxiliary metabolic genes, ranging from a complete ABC transporter cassette to a large antimicrobial resistance gene cluster. These results unveil the previously unrecognized diversity of AOB phages and their potential to alter host physiology. Our data demonstrates a broad taxonomic and functional repertoire of cultured AOB phages, greatly expanding the panel of known AOB phages, suggesting that viruses play a more significant and complex role in nitrification than previously understood. Moreover, we outline an effective methodological framework for isolating AOB phages from environmental samples. These results will help reframe our understanding of environmental nitrification and enable intensified selection and use of phages for its control.

|
Scooped by
mhryu@live.com
Today, 12:52 AM
|
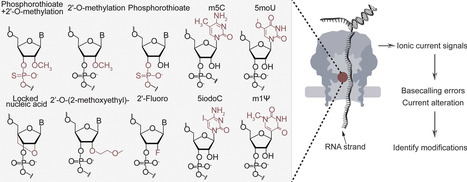
While nanopore direct RNA sequencing has substantially advanced transcriptomics, its detection of RNA modifications remains primarily focused on abundant biological base modifications. However, therapeutic RNAs employ a diverse catalog of modifications, including base, sugar, and backbone modifications, to enhance stability and pharmacological properties. To address this gap, we systematically evaluated a set of therapeutically relevant modifications [phosphorothioate (PS)], sugar [2′-O-methylation (2′OMe), 2′-Fluoro (2′F), locked nucleic acid (LNA), 2′-O-(2′-methoxyethyl) (2′MOE)], and base [N1-methylpseudouridine (m1Ψ), 5-methylcytidine (m5C), 5-methoxyuridine (5moU), and 5-iodocytidine (5iodoC)] using direct RNA nanopore sequencing. Modifications were systematically analyzed using basecall errors, raw current signals, and modification-aware basecalling models. Ribose modifications, m1Ψ, and 5moU induced significant error rate increases and noticeable current alterations, whereas 2′OMe and 2′MOE affected dwell time adjacent to the pore. In contrast, PS linkages produced only slight current alterations without increasing basecalling errors. We further evaluated modification-aware basecallers for 2′OMe and m5C. While these tools can distinguish modification types, they are limited by poor quantification accuracy and high local error rates, especially for 2′OMe. This study establishes a critical performance baseline, clarifying the current capability and limitations of nanopore technology for the analysis of therapeutically relevant RNA modifications.

|
Scooped by
mhryu@live.com
Today, 12:43 AM
|
A bacterial colony rarely exists in isolation – in natural habitats, colonies interact to form spatially structured communities across length and time scales. Eco-evolutionary feedbacks link these scales, such that structure at one level can influence another, yet the interplay between single- and multi-colony organization remains poorly understood. As a step toward addressing this, we develop a high-throughput platform to track population dynamics across spatially extended networks of colonies. A common structural feature observed at the multi-colony scale is the formation of a stable gap region between colonies, even when they are isogenic. Numerous studies observe similar patterns of behavior across species, with few resolving the underlying mechanism. Here, we ask: what are the minimal ingredients shaping this multi-colony structure? We focus on colonies of the opportunistic pathogen Enterococcus faecalis, a model organism for which this behavior has yet to be reported. By combining modeling and experiments, we show that both nutrient competition and direct growth inhibition control colony morphology and expansion of interacting colonies. We identify distinct regimes of gap formation, relating intra- and inter-colony spatial patterns to ecological interactions mediated at the cellular scale. Together, our results suggest that antagonism, even between isogenic populations through self-inhibition, is likely a common behavior of bacterial species in general.

|
Scooped by
mhryu@live.com
Today, 12:26 AM
|
Bacteria require rapid adaptation under fluctuating environmental conditions. Commonly recognized global regulators enable bacteria to respond promptly to external changes, though they are either restricted to specific bacterial taxonomies or physiological statuses, suggesting that additional regulators are required for adaptation. DNA methylation is a reversible modification affecting bacterial gene regulation. However, conventional methods can only detect one DNA methylation form each round, leaving the understanding of DNA methylation in bacterial adaptation mostly unknown. This study aimed to identify genome-wide DNA methylation variation (N6-methyladenine, N4-methylcytosine, and 5-methylcytosine) in E. coli under different culture conditions using Oxford Nanopore sequencing. DNA samples from six conditions (normal, low oxygen, low pH, high temperature, high salt, and recovery after low pH exposure) during the exponential and stationary phases were extracted. When culture conditions were compared to the normal condition, E. coli exhibited more differentially methylated sites during the exponential phase than in the stationary phase. During the exponential phase, the genes differentially methylated in all conditions were involved in cellular activities, such as cellular and metabolic processes. During the stationary phase, universally differentially methylated genes were associated with oxidation responses. Subsequent analysis found that although DNA methylation analysis was affected by batch effects, some genes (e.g. rpoS) showed consistently differential methylation across datasets. Our findings suggest that the E. coli DNA methylation profile was affected by growth phases and conditions, and DNA methylation profiling by Oxford Nanopore sequencing could be a potential approach for gene activity estimation in environmental samples.

|
Scooped by
mhryu@live.com
Today, 12:02 AM
|
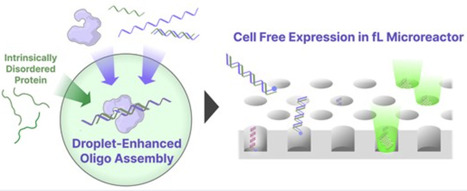
Recent advances in in silico protein design and bioinformatics have enabled the rapid generation of candidate sequences for functional proteins. However, experimental validation remains a bottleneck, largely due to time-consuming DNA assembly and cell-based cloning processes. Technologies that reduce the time required to convert synthetic oligonucleotides (oligos) into expressed proteins are therefore of considerable interest. Here, we demonstrate that phase-separated droplets formed by the intrinsically disordered protein (Ddx4N1) concentrate both oligos and ligation enzymes, enabling efficient oligo assembly at nanomolar to sub-nanomolar concentrations that are typically inaccessible to conventional ligation-based methods. The assembled products can be directly introduced into femtoliter-scale microreactors for digital cell-free gene expression, allowing protein expression from single assembled DNA molecules without polymerase chain reaction amplification or cellular cloning. Simultaneous expression of two distinct proteins from separately assembled DNA templates in a one-pot reaction was also demonstrated. The complete workflow—from oligo assembly to detectable protein expression—can be performed within half a day. While further development will be required to enhance reaction parallelization and enable systematic retrieval of sequence information from expressed products, this amplification-free, low-input system establishes a technical foundation for integrating oligo-pool-based gene assembly with digital protein prototyping platforms.

|
Scooped by
mhryu@live.com
May 4, 11:15 PM
|
Selenoproteins, a unique class of proteins critical for cellular antioxidant defense, are characterized by the incorporation of selenocysteine (Sec) in their active sites. Sec is co-translationally inserted into proteins via a specialized mechanism that reprograms the UGA codon to encode Sec, involving a specific RNA structure designated the Sec insertion sequence (SECIS) element and several essential enzymes. Although numerous selenoproteins have been identified in prokaryotes (primarily bacteria), the detection of selenoprotein genes in these organisms remains challenging, largely due to difficulties in distinguishing the Sec-encoding UGA codon from standard termination signals. In recent years, computational approaches for predicting selenoprotein genes, along with comparative genomic analyses of Sec-encoding machinery and selenoproteomes, have emerged as a promising and rapidly evolving field, offering new insights into Sec utilization in bacteria and archaea. This review provides a comprehensive overview of the latest advancements in the study of selenoproteins in prokaryotes. We summarize the molecular mechanisms underlying Sec biosynthesis and incorporation, and the structural diversity of SECIS elements in bacteria and archaea. We then describe current computational strategies for the identification of prokaryotic selenoprotein genes and present an updated, extensive catalog of prokaryotic selenoproteins documented to date, emphasizing those with well-established functions. Finally, we discuss recent progress in understanding the evolutionary dynamics of the Sec-encoding system and selenoproteins across prokaryotes, with a focus on the archaea-to-eukaryote transition of Sec machinery and selenoproteins. Overall, this review offers a unified perspective on the identification, functions, and evolution of selenoproteins in prokaryotes.

|
Scooped by
mhryu@live.com
May 4, 7:28 PM
|
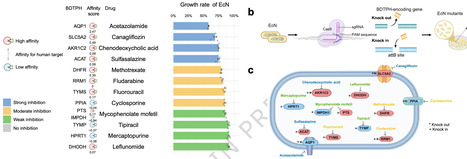
While the impact of non-antibiotic drugs on gut bacteria is well-known, their mechanisms of action remain poorly characterized, and effective mitigation strategies for drug-induced dysbiosis are still limited. Here, we screened bacteria-derived drug-target protein homologs (BDTPHs) mapped to 63 target proteins and 107 associated drugs to quantify “drug–BDTPH–bacterium” interactions. These interactions were validated by co-culture experiments using 10 drugs and 25 strains, enzyme assays, and genetic perturbations in Escherichia coli. Ex vivo and in vivo testing with six drugs showed that over 50% of affected genera exhibited high affinity, indicating microbiota alterations through the “drug–BDTPH–bacterium” axis. Leveraging this quantitative interaction framework, we identified a strain of Bifidobacterium animalis that can competitively bind methotrexate through high-affinity BDTPH, thereby effectively alleviating gut microbiota dysbiosis in vivo. Our findings elucidate a mechanism by which non-antibiotic drug effects on bacterial growth, and suggest a universal homology-based competition strategy to restore drug-disrupted microbiota. Here, the authors uncover a mechanism by which drug-target homologs mediate non-antibiotic drug effects on bacterial growth, and suggest a universal homology-based competition strategy to reverse MTX-induced microbiota dysbiosis.
|
 Your new post is loading...
Your new post is loading...



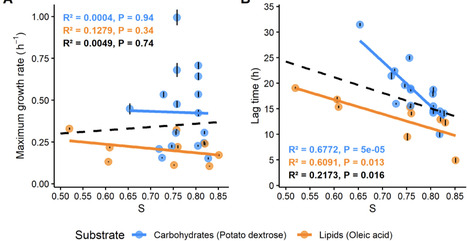


















To infer the translational adaptation for these two lifestyles, we predicted the codon optimization index (S) for lipid and carbohydrate transport and metabolism pathways. The parameter S and respective statistical significance were estimated based on the tRNA gene pool, codon usage bias, and effective number of coding codons among all genes in the respective metabolic pathway. Therefore, S was used as a proxy for the measure of selection on the overall translational efficiency within these pathways. To account for interspecific variation in codon usage bias, S values were normalized by comparing the mean S values across genes associated with lipid and carbohydrate metabolic pathways (S ratio) or by normalizing pathway-specific S values to the genome-wide mean S (S normalized)