Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 9:26 AM
|
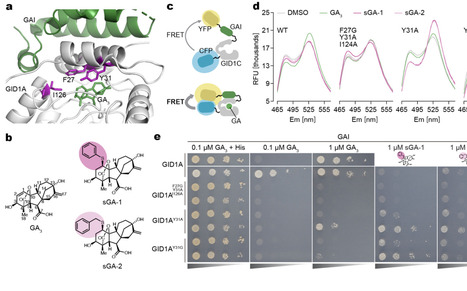
Gibberellins (GAs) influence cell division and elongation, profoundly shaping plant architecture and yield. GA perception occurs when bioactive GAs bind the receptor GID1, promoting DELLA degradation and activating transcriptional programs. While GA signaling in the root endodermis is essential for promoting root elongation, functions of other layers in spatial control of GA responses have not been explored. Here, we developed a synthetic GA (sGA) that does not bind endogenous GID1, together with a modified GID1 (mGID1) engineered to selectively recognize sGA, enabling cell-specific activation of GA signaling in vivo. Using this system in Arabidopsis, we demonstrate that coordinated action of GA signaling in the endodermis, epidermis, and other layers is required for full root elongation. Moreover, cell type-specific expression of GA biosynthetic enzymes indicates the existence of intercellular GA transport. The sGA-mGID1 system provides a versatile platform for spatially precise reprogramming of hormone signaling, enabling synthetic control of developmental processes such as root-shoot growth balance, thereby advancing applications in plant synthetic biology and sustainable crop improvement.

|
Scooped by
mhryu@live.com
Today, 9:18 AM
|
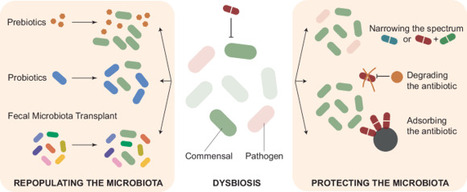
The discovery of antibiotics and their subsequent therapeutic use revolutionized our ability to treat once deadly infectious diseases, and antibiotics have become one of the most commonly prescribed drug classes. Unfortunately, these compounds not only target pathogenic strains, but also non-pathogenic bacteria that fulfill important functions for the human host. As such, antibiotic treatment can cause severe collateral damage, resulting in dysbiosis, for example, in the human gut microbiome. Given the immense importance of the gut microbiome for human health, antibiotic-induced dysbiosis can cause a variety of detrimental health outcomes. In addition, antibiotic (over-)use causes selection of antibiotic-resistant strains, and the human gut microbiome has become a major reservoir for resistance determinants that can transfer to pathogenic isolates and cause hard-to-treat infections. In this review, we describe various adverse effects that antibiotic use has on the human gut microbiome, how we can approach this problem experimentally, and discuss pathways to mitigate antibiotic-induced collateral damage.

|
Scooped by
mhryu@live.com
Today, 12:36 AM
|
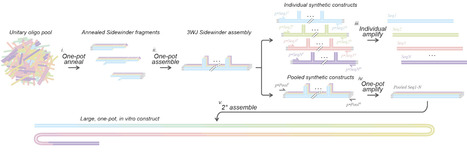
Reliable and cost effective de novo DNA production has become central to studying and engineering biology. Short synthetic single-stranded DNA oligo pools offer substantially reduced costs at the sacrifice of yield and individual oligo isolation. Efficiently constructing longer synthetic double-stranded DNA molecules from oligo pools as the input has remained an engineering challenge with the potential to drastically reduce costs, labor, and experimental turn-around time. Here we show one-pot, parallel assembly of hundreds of DNA fragments simultaneously into dozens of defined constructs with high fidelity using Sidewinder. We designed a novel string-based bespoke barcode design algorithm which rapidly generates Sidewinder barcodes at unprecedented scale. We apply the new algorithm to Sidewinder using oligo pools, demonstrating construct-specific amplification from pooled assemblies with misconnection rates as low as 1 in 10,000,000. Further, we demonstrate universal amplification of pooled assemblies to generate a library of specific target sequences that we combine with in vitro hierarchical assembly to 12.5 kilobases.

|
Scooped by
mhryu@live.com
Today, 12:19 AM
|
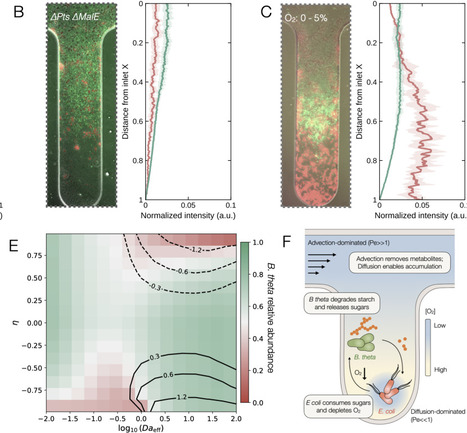
Microbial interactions unfold within environments structured by physical transport and chemical gradients. Yet most mechanistic studies rely on well-mixed systems that mask the reciprocal influences of environmental heterogeneity on metabolism and ecology. Here, we investigate how the physical environment modulates the interaction between the gut commensal Bacteroides thetaiotaomicron and E. coli. In anoxic liquid culture, cell-resolved isotope imaging and genetic perturbations reveal exploitative cross-feeding, where E. coli consumes diffusible sugars released by B. thetaiotaomicron during starch degradation. When exposed to intestinal-like oxygen gradients in microfluidics, the interaction is restructured by spatial organization. The species self-organize into complementary niches: E. coli locally depletes sugars and oxygen, thereby expanding the anoxic niche required by B. thetaiotaomicron. A reactive transport model confirms that this organization arises from coupled feedback between physical transport and metabolic reaction rates. Together, our results reveal how physical structure and chemical gradients convert an exploitative cross-feeding interaction into a dynamic niche-construction process that generates emergent spatial organization and stabilizes coexistence.

|
Scooped by
mhryu@live.com
May 2, 11:51 PM
|
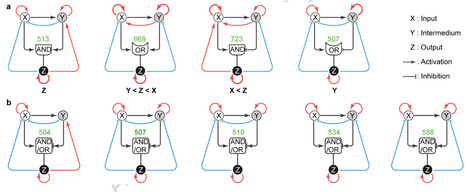
Feedforward loops (FFLs) and feedback loops (FBLs) are ubiquitous network motifs that mediate signal filtering, pulse generation, and state switching; yet, how coupling FBLs to FFLs produces robust multistability—a key mechanism for cellular decision-making—remains unclear. Here, we systematically investigate coupled FFL–FBL architectures by focusing on two prevalent FFL types, each with AND or OR logic, yielding four distinct frameworks. For each framework, we enumerate all 36 = 729 possible circuits, corresponding to three possible states (activation, inhibition, or absence) for each of six feedback edges, formulate each circuit as a system of ordinary differential equations, and quantify robustness as the proportion of 100,000 randomly sampled parameter sets exhibiting multistability. Our results reveal two key principles. First, positive self-activation is a primary driver of multistability, but the identity of the critical node(s) depends on the FFL type and logic. Second, coherent FFLs support multistability more readily than incoherent ones, whereas the choice between AND and OR logic has a comparatively weaker effect. Notably, we identify representative high-performing circuits within each framework and find that a small set of circuit designs remain robustly multistable across all four frameworks. These findings advance the theoretical understanding of motif design and provide practical guidelines for engineering synthetic multistable circuits.

|
Scooped by
mhryu@live.com
May 2, 10:52 PM
|
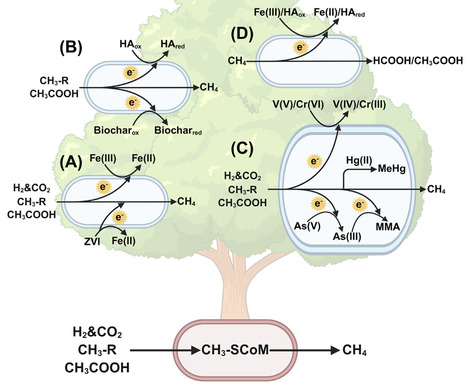
As key players in anoxic environments, methanogenic archaea account for the vast majority of biogenic methane emissions and play important roles during organic matter mineralization. Growing evidence indicates potential for broader ecological roles beyond methanogenesis. This review summarizes the physiology of biogeochemically relevant transformations of iron minerals, humic acid, biochar, and toxic heavy metals/metalloids. We examine how methanogenic archaea couple carbon cycling with other element cycles through methanogenic pathways and electron transfer processes. We also discuss known and potential molecular mechanisms that underlie these interactions. Furthermore, we highlight the discovery of the reversal of methanogenesis driven by extracellular electron transfer and its potential for methane bioconversion technologies. This work underscores the physiological versatility of methanogenic archaea, suggesting their diverse roles in biogeochemical cycles.

|
Scooped by
mhryu@live.com
May 2, 10:46 PM
|
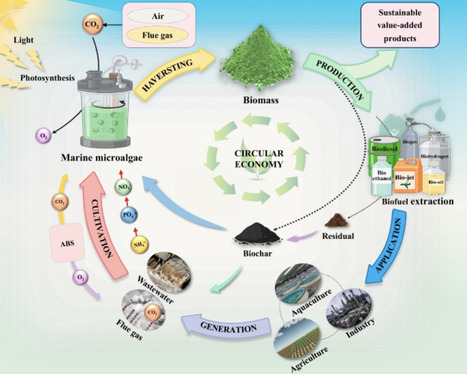
Marine microalgae are frequently promoted as sustainable biofuel feedstocks because of their halotolerance, high photosynthetic efficiency, and limited land requirements, yet commercial deployment remains elusive. This gap is primarily systemic rather than biological, reflecting the fragmented development of strain engineering, harvesting, conversion, and sustainability assessment. This review reframes marine micro-algae as circular biofactories and advances a system-centric paradigm for integrated biorefineries. We synthesise recent advances in metabolic and genetic engineering, low-energy harvesting, and thermochemical and biochemical conversion, highlighting how cross-stage interdependencies dominate overall performance. We further discuss how artificial intelligence, digital twins, nutrient recycling, carbon utilization, and high-value coproducts enable predictive optimization and techno-economic viability. This perspective provides a road map for translating marine microalgal biofuels from laboratory promise to industrial relevance.

|
Scooped by
mhryu@live.com
May 2, 10:40 PM
|
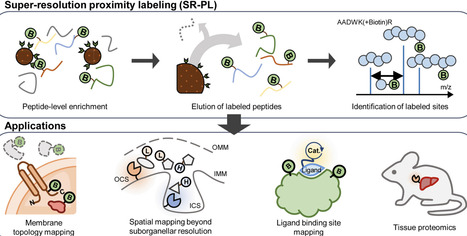
Super-resolution proximity labeling (SR-PL) advances spatial proteomics beyond conventional protein-level enrichment, enabling residue-resolved analysis of subcellular organization in living cells. Conventional proximity labeling relies on streptavidin-based capture and on-bead digestion, producing protein-centric readouts with limited structural insight. In contrast, SR-PL directly recovers biotinylated peptides and identifies labeled amino acid residues by LC–MS/MS. These site-specific labels serve as direct evidence of proximity, allowing for the precise mapping of protein surfaces, solvent accessibility and interaction interfaces. By linking spatial proximity to specific structural features, SR-PL enables mechanistic interpretation of spatial proteomic data and reframes proximity labeling as a structure-informed analytical framework. Recent advances in affinity capture strategies—including engineered probes, reversible affinity matrices, and optimized antibody reagents—have improved selective enrichment and gentle peptide release while reducing background contamination. Together, these developments position SR-PL for broad applications such as membrane topology mapping, organelle contact site analysis, and ligand-dependent interactions.

|
Scooped by
mhryu@live.com
May 2, 10:33 PM
|
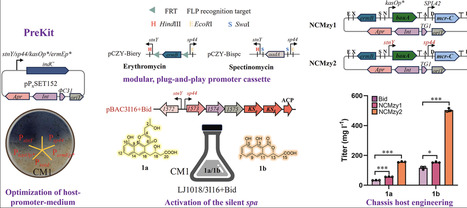
The discovery of novel scaffold compounds or enzymes through genome mining of biosynthetic gene clusters (BGCs) represents cornerstone of drug development. A major obstacle, however, is that most microbial BGCs encoding secondary metabolites remain silent under standard laboratory conditions. While promoter engineering coupled with heterologous expression has become a mainstream strategy to activate these silent BGCs, its efficiency is often limited by inherent incompatibilities among key factors, including host selection, promoter strength, and culture medium. To overcome this issue, we developed a streamlined platform integrating a promoter-library with an indC-based reporter system. This platform enables combinatorial screening across diverse media conditions and different heterologous hosts, systematically resolving the multi-factor incompatibility issue. Building on this strategy, we further developed a plug-and-play promoter toolkit, which was successfully applied to activate a silent type II polyketide synthase (PKS) gene cluster (spa). This led to the production of one new aromatic polyketides compounds strepanthenes A (1a) and streptoketide C (1b). Furthermore, enhancement of precursor supply was achieved by integrating a non-carboxylative malonyl-CoA (NCM) pathway into S. lividans LJ1018 through attPTG1 site-specific integration system, we achieved a 3.6-fold and 3.3-fold increase in the titers of compounds 1a and 1b, respectively. Collectively, this work provides a versatile and efficient strategy for activating silent BGCs and improving metabolite production. Bioactivity evaluation revealed that new compound 1a exhibited antibacterial activity against S. aureus and E. faecalis, with a consistent MIC value of 4 μg mL−1 against both strains.

|
Scooped by
mhryu@live.com
May 2, 4:53 PM
|
‘Structurally known’ and ‘Structurally unknown’ have long served as a binary classification of natural products. However, with the emergence of synthetic biology, the boundaries of this simple classification are being redrawn, because SynBio expands both what we can see and what we can build. To reflect this change, we propose a SynBio-centered framework that maps antimicrobial natural products along two axes, chemical novelty and engineering accessibility, yielding four quadrants: (i) known knowns, well-characterized natural products accessed mainly through pathway refactoring and producer strain optimization; (ii) unknown knowns, engineerable scaffolds with established diversification routes but uncertain chemical and functional outcomes; (iii) known unknowns, architectures with precedent but limited realization due to missing transferable engineering routes, often because the relevant coupling logic has not yet been converted into a practical engineering handle; (iv) unknown unknowns, unanticipated scaffolds that need to be recognized through minimal chemical, genetic, or mechanistic anchors before engineering. Since these quadrants are dynamic rather than fixed, compounds may shift across the framework as discovery, route-building, and outcome predictability improve. This two-axis map highlights complementary SynBio strategies, from optimizing existing antibiotics to accessing first-in-class entities, which may help guide efforts to combat current resistance challenges.

|
Scooped by
mhryu@live.com
May 2, 4:43 PM
|
The systematic exploration of novel bioactive compounds with superior functional properties is critical for driving innovations in agriculture, healthcare, and related fields, thereby becoming essential for advancing sustainable biotechnological solutions. Nonprotein amino acids (NPAAs), functional amino acids not incorporated into proteins, exhibit unique physiological activities and provide distinctive advantages in nutritional enhancement, functional product formulation, and food/feed processing. These attributes challenge the conventional perception of proteins as mere nutritional carriers, positioning NPAAs as promising bioproducts for biosynthesis and functional applications in agriculture, food, and medicine. This review summarizes the classification of the available NPAAs based on their synthetic substrates for the first time and then outlines their diverse functional roles. A comprehensive analysis of recent advances in biosynthetic pathways, engineering strategies, and production level demonstrates their primary research progress in the laboratory phase. The further sustainable biomanufacturing of NPAAs is hampered by several challenges, including poorly elucidated biosynthetic mechanisms, limited robustness and low productivity of microbial strains, and difficulties in scaling up production for industrial applications. Addressing these bottlenecks will require innovative strategies and technologies to facilitate the translation of NPAA production from bench to industry. This review offers valuable insights into the potential of NPAAs in the development of next-generation bioproducts of nutrition, immune regulation, antioxidant defense, and intestinal homeostasis maintenance, suggesting a promising direction for microbial production of high-performance bioactive molecules in agricultural synthetic biomanufacturing.

|
Scooped by
mhryu@live.com
May 2, 4:33 PM
|
CRISPR–Cas effectors typically rely on RNA guides to recognize target sequences. In Cas12a, the protospacer adjacent motif on DNA engages conserved protein residues, triggering target binding and nuclease activation. Here we reprogram Cas12a into a DNA-guided, RNA-targeting effector. Exploiting protospacer-adjacent motif-dependent interaction, we engineer synthetic CRISPR DNA that engages Cas12a to form a functional deoxyribonucleoprotein complex, while repurposing solely RNA as the programmable target. Structural, biophysical and biochemical analyses reveal the molecular basis of this DNA-guided, RNA-targeting configuration and support an activation pathway distinct from that of canonical RNA-guided systems. DNA-guided Cas12a enables direct RNA detection and efficient intracellular RNA knockdown, establishing a modular activation architecture for CRISPR–Cas12a and expanding the design space for programmable RNA manipulation. Synthetic DNA guides (crDNA) reprogram Cas12a nucleases for RNA targeting.

|
Scooped by
mhryu@live.com
May 2, 4:18 PM
|
Metabolic interactions are central to the functioning of microbial communities. In spatially structured environments, such interactions typically require close physical proximity between partner cells. However, cell division drives spatial segregation of interaction partners, as the cells emerging from division remain adjacent and form clonal clusters, weakening these interactions. Here, we hypothesized that active cell motility can reduce this spatial segregation by enabling cells to leave clonal clusters and thereby enhance metabolic interactions. We tested this hypothesis by performing time-lapse single-cell imaging of a synthetic cross-feeding consortium growing in microfluidic chambers. We found that surface motility disrupted clonal clustering, enhanced spatial intermixing, and consequently increased the growth rates of individual cells and community productivity. Individual-based simulations further revealed that this effect is robust across motility modes and a wide range of ecological and physiological parameters. Together, our findings demonstrate that even non-directed, random motility, without chemotactic sensing, is sufficient to enhance metabolic interactions by separating cells from their clonal lineages and repositioning them in proximity to their metabolic partners, thereby acting as a key driver of community functioning in spatially structured microbial systems.
|

|
Scooped by
mhryu@live.com
Today, 9:23 AM
|
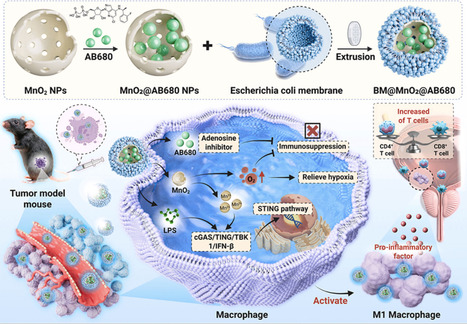
Bladder cancer is a highly malignant tumor with limited treatment options. Although adoptive cell therapy has shown promise in oncology, its efficacy is often constrained by poor intrinsic antitumor activity and a highly immunosuppressive tumor microenvironment. To address these challenges, this study develops a novel living adoptive macrophage therapy armed with nano-bionic bacteria for bladder cancer treatment. Specifically, hollow manganese dioxide (MnO2) nanoparticles were synthesized, loaded with an adenosine inhibitor, and coated with E. coli membranes. Under mild conditions, these nano-bacterial particles were internalized by macrophages, transforming them into living cell drug factories. Upon accumulation into the bladder, these engineered macrophages actively infiltrated tumor tissues. Lipopolysaccharide (LPS) from the bacterial membrane, along with Mn2+ ions, activated the STING pathway in macrophages, promoting and sustaining an M1-like antitumor phenotype. The activated macrophages released pro-inflammatory cytokines, thereby stimulating resident immune cells within the tumor and initiating robust antitumor immunity. Additionally, the macrophages released the adenosine inhibitor while MnO2 generated oxygen, synergistically counteracting adenosine-mediated immunosuppression. In a subcutaneous tumor model, this nano-bionic bacteria-armed macrophage therapy significantly enhanced the therapeutic outcome against bladder cancer. Mac@ABMn triggered reprogramming of the tumor microenvironment, resulting in enhanced anti-tumor immunity, characterized by increased infiltration of activated CD4+ and CD8+ T cells without expanding regulatory T cells (Tregs), thereby shifting the milieu toward a potent immunostimulatory state. The immunostimulatory effect is molecularly defined by the cGAS/STING/TBK1/IFN-β signaling axis, and Mac@ABMn demonstrates significant synergistic efficacy when combined with standard immunotherapies like BCG or anti-PD-L1 checkpoint blockade, leading to superior tumor control. Collectively, this study developed a novel therapy regiment, featuring macrophages engineered with bacterial membrane-coated nanoparticles are activated via the STING pathway to sustain an anti-tumor M1 phenotype while simultaneously alleviating hypoxia and scavenging immunosuppressive adenosine.

|
Scooped by
mhryu@live.com
Today, 1:06 AM
|
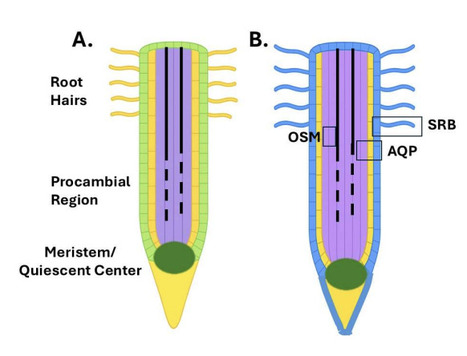
Salinity exerts a major constraint on global crop production while seawater intrusion impacts coastal aquifers and surface waters. Using a blueprint from nature, we produced highly salt tolerant Arabidopsis and rice. Endodermal-like barriers, duplicated to the root epidermis and distally expanded to protect sensitive regions, provide salt tolerance to 600 mM NaCl, levels comparable to seawater. Two additional genetic modules are added to reduce adverse effects on ion uptake and provide osmotic protection. Arabidopsis and rice containing all three genetic modules can survive 600 mM NaCl and set seed. RNA-seq analysis suggests that our rational engineering primes plants for salt tolerance, even without salt exposure, while our ionic analysis provides means for improvement. Our results, duplicating suberin and the Casparian Strip to the epidermis, adding symplastic transport and providing a means to address osmotic stress, provides a new approach to salt tolerance and insight to genes involved in salt responses.

|
Scooped by
mhryu@live.com
Today, 12:33 AM
|
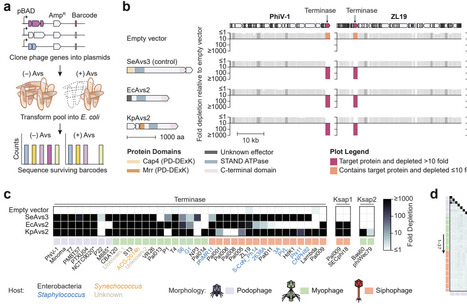
Prokaryotes employ diverse defense strategies to detect and halt the progression of phage infection. Multiple defense systems sense phage proteins through direct binding, including antiviral STAND NTPases (Avs), which oligomerize upon target recognition to induce programmed cell death. The widespread Avs2 family was previously shown to detect the large terminase subunit of tailed phages, but the mechanism of terminase sensing was unknown. Here, we determine the structural basis of terminase recognition by Avs2 from E. coli (EcAvs2). A cryo-EM structure at 2.3 Å resolution reveals that EcAvs2 forms a flat, C4-symmetric tetramer in which each protomer is bound to a single terminase monomer. Terminase recognition is mediated by a large, shape complementary binding pocket in the EcAvs2 sensor domain, including specific contacts with an unexpected ATP molecule at the interface of EcAvs2 and terminase. Furthermore, we demonstrate that the defense protein Upx also recognizes diverse phage terminases, despite lacking sequence and structural homology to Avs. AlphaFold 3 models indicate that Upx binds an unfolded state of the core terminase ATPase domain, mediated by β-augmentation. These findings highlight the distinct modes of terminase recognition across structurally diverse defense proteins.

|
Scooped by
mhryu@live.com
May 2, 11:59 PM
|
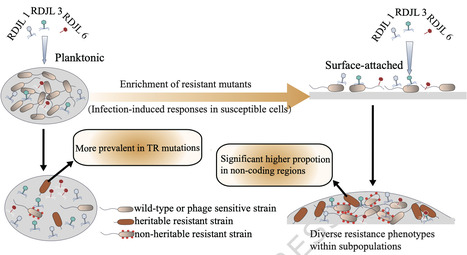
Marine bacteria alternate between planktonic and surface-attached lifestyles, facing continuous phage predation. However, how these lifestyles shape resistance evolution remains poorly understood. Using a Roseobacter model strain, we demonstrate that surface-attached populations exhibit 26-fold higher survivability than planktonic counterparts during lytic phage infection. This advantage emerges through the evolution of heterogeneous subpopulations exhibiting diverse resistance phenotypes, a pattern absent in planktonic populations. Whole-genome sequencing of 139 heritable phage-resistant mutants revealed fundamentally divergent mutational patterns, with planktonic populations predominantly harboring tandem repeat mutations, whereas surface-attached populations favor non-coding mutations. Despite this, both lifestyles independently converged on mutations in the CtrA phosphorelay system, identifying CtrA as a previously unrecognized evolutionary target of phage-driven selection and triggering planktonic-to-surface-attached switch. Further analyses revealed systematic downregulation of motility genes and enhancement of biofilm formation, mechanistically linking phage resistance to lifestyle transitions. The identified CtrA mutations occur in regions highly conserved across ecologically important marine Alphaproteobacteria (Rhodobacterales) that are known to switch between planktonic and surface-attached states, suggesting lifestyle-dependent evolutionary trajectories may broadly shape phage resistance in marine ecosystems. Marine bacteria are often under constant phage predation. Li et al. present that surface-attached populations exhibit 26-fold higher survivability than planktonic counterparts during lytic phage infection. They identify CtrA as an evolutionary target for phage-driven selection towards an attached lifestyle.

|
Scooped by
mhryu@live.com
May 2, 10:56 PM
|
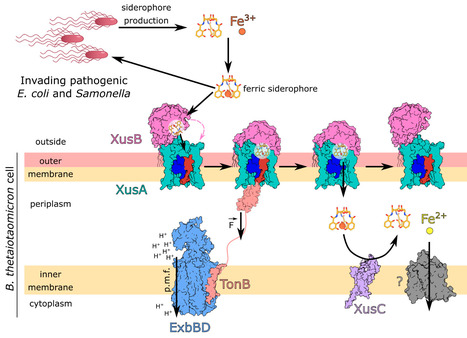
Iron is an essential element that can be growth-limiting in microbial communities, particularly those present within host organisms. To acquire iron, many bacteria secrete siderophores, secondary metabolites that chelate ferric iron. These iron chelates can be transported back into the cell via TonB-dependent transporters in the outer membrane, followed by intracellular liberation of the iron. Pathogenic E. coli and Salmonella produce siderophores during gut infection. In response to iron starvation, the human gut symbiont Bacteroides thetaiotaomicron upregulates an iron piracy system, XusABC, which steals iron-bound siderophores from the invading pathogens. Here, we investigated the molecular details of xenosiderophore uptake across the outer membrane by the XusAB complex. Our crystal and cryogenic electron microscopy structures explain how the XusB lipoprotein recognizes iron-bound xenosiderophores and passes them on to the XusA TonB-dependent transporter. Moreover, we show that Xus homologues can transport a variety of siderophores with different iron-chelating functional groups.

|
Scooped by
mhryu@live.com
May 2, 10:48 PM
|
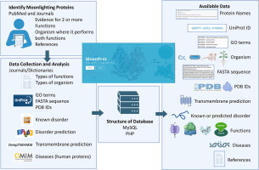
MoonProt 4.0 ( http://moonlightingproteins.org) is an updated open-access database storing manually-curated annotations for moonlighting proteins. Moonlighting proteins exhibit two or more physiologically relevant distinct biochemical or biophysical functions performed by a single polypeptide chain. Here we describe an expansion in the database since our report published in 2021. With the assistance of five undergraduate annotators, we have added approximately 200 protein entries to give a total of over 700 moonlighting proteins. The new entries include more examples from plants, more transmembrane proteins and additional combinations of functions. The MoonProt Database collection of proteins with multiple functions serves as a resource for developing algorithms for predicting protein functions and provides examples of the evolution of new functions on a protein scaffold that can be valuable in developing novel methods for designing proteins with added functions.

|
Scooped by
mhryu@live.com
May 2, 10:45 PM
|
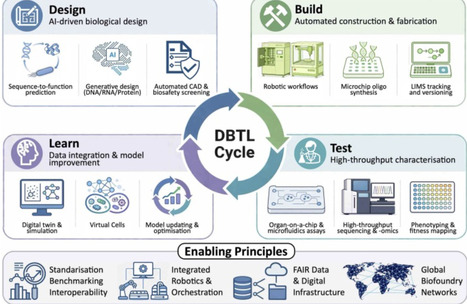
Since the emergence of synthetic biology, biofoundries have developed as enabling infrastructures that scale engineering biology globally. Landmark initiatives, such as Genome Project-Write, JCVI-syn3.0, Sc2.0, SynMoss and the Synthetic Human Genome Project, have significantly advanced the feasibility of constructing chromosome-sized DNA and revealed key principles of genome function and design. Nevertheless, the intrinsic complexity of cellular systems and the resource-intensive nature of experimental design-build-test-learn cycles continue to constrain innovation. Recent advances in artificial intelligence (AI), whole-cell modelling and digital twinning are now creating opportunities for self-improving, AI-driven biofoundries that seamlessly integrate in silico design and validation with miniaturised and automated in vitro testing. This review surveys the technologies shaping AI-driven synthetic biology, highlighting their convergence with automation, digitisation and miniaturisation to enable fully autonomous biofoundries that unify computational design, automated fabrication and data-driven learning within a single adaptive framework.

|
Scooped by
mhryu@live.com
May 2, 10:37 PM
|
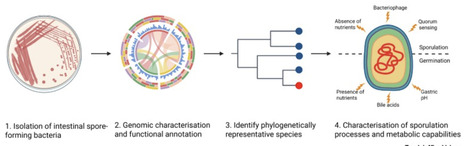
Anaerobic gut bacteria have highly evolved functions that promote transmission between human hosts. These include resilient spores produced by many Bacillota (formerly Firmicutes) bacteria, which are tolerant to extended ambient oxygen exposure. Recent culturing and genomic studies have revealed the taxonomic diversity of gut spore-formers and their prevalence in human populations. However, due to a reliance on spore-forming model organisms that are not representative of commensal gut bacteria, we still have a limited understanding of gut microbiome-specific sporulation processes and their underlying genetics. In this opinion article, we outline a rational path for development of new model organisms derived from commensal spore-forming gut bacteria in order to obtain fundamental insights into sporulation and to provide a discovery platform for novel probiotic or microbiome-based therapeutic development.

|
Scooped by
mhryu@live.com
May 2, 4:54 PM
|
Land plants underpin civilization and planetary health, yet their genomic diversity remains largely uncharted. Current resources are unstandardized and scarce, lacking reference genomes for 95% of genera, 70% of families, and 51% of orders, impeding evolutionary and functional insight. We thus propose the PLANeT initiative, an international effort to generate high-quality, standardized genomes across the plant tree of life. Integrating artificial intelligence (AI) with genomics, we will decode conserved principles to advance fundamental plant biology, biodiversity conservation, crop improvement, and natural product discovery. Engaging around 100 labs to train 1,000 scientists, we will tackle pivotal questions for a sustainable future.

|
Scooped by
mhryu@live.com
May 2, 4:46 PM
|
Integrons are genetic elements that drive bacterial adaptation by capturing and expressing mobile gene cassettes. They play a key role in dissemination of antimicrobial resistance (AMR) genes, particularly in Gram-negative bacteria. In addition to known AMR determinants, integron gene cassettes carry a vast reservoir of novel genes whose functions are largely uncharacterised, making it difficult to assess their full contribution to the resistome. Contributing to this are limitations in current sequence-based prediction methods which often lack the ability to identify unknown AMR or other adaptive genes with novel mechanisms. To address this, we developed a high-throughput gene cassette capture system that utilizes site-specific recombination activity of integrons and a counter selection strategy to capture and express gene cassettes from metagenomes. Coupling this platform with a functional screening approach allowed us to rapidly assay large libraries of environmental gene cassettes. Using this system, we recovered previously unknown AMR determinants while also providing insights into the prevalence of known clinical AMR genes in a range of environmental samples, including food and fertiliser. Here we provide experimental data on multiple novel bleomycin resistance genes and a stress response gene conferring gentamicin and tobramycin resistance. Our sequence analysis of the captured library also highlighted the diversity of the environmental cassette pool, with 656 unique cassettes recovered, the majority of which encoded proteins with unknown functions. The cassette capture system is a powerful tool for accessing hidden elements of the resistome and discovering novel adaptive genes that may go undetected using current sequence-based approaches.

|
Scooped by
mhryu@live.com
May 2, 4:41 PM
|
Carboxysomes are self-assembling proteinaceous microcompartments that encapsulate ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco) and carbonic anhydrase within a semi-permeable shell, thereby elevating local CO2 concentrations around Rubisco to improve carbon fixation. Their inherent design principles, including programmable architecture, cargo encapsulation, semi-permeability, and modular assembly, position carboxysomes as powerful paradigms in synthetic biology and bioengineering, offering unprecedented opportunities to boost carbon assimilation and unlock novel biotechnological functions. This review summarizes current knowledge of the molecular mechanisms of carboxysome structure, assembly, and function, and highlights key recent breakthroughs and key challenges such as achieving precise control of shell permeability and efficient cargo encapsulation, integrating carboxysomes into heterologous hosts. We also outline emerging strategies and future perspectives for engineering carboxysomes as enhanced CO2-fixing engines and repurposing them as versatile nanomaterials in biotechnological applications. Together, these advances underscore the growing potential of carboxysome engineering to transform carbon-fixation pathways across diverse biological systems.

|
Scooped by
mhryu@live.com
May 2, 4:32 PM
|
Plant-based materials provide a renewable and sustainable platform for tackling global challenges in energy, environment and healthcare. Among these, chronic diabetic wounds, currently affecting over 10% of the global population, remain a pressing public health crisis, largely owing to persistent hypoxia and unresolved inflammation. Here we report photosynthetic microspheres derived from the invasive aquatic plant Eichhornia crassipes (water hyacinth) as a green therapeutic strategy for chronic wound repair. Harnessing intact chloroplasts, these biogenic microspheres deliver sustained oxygen evolution for up to 3 weeks, alleviating local hypoxia and restoring redox balance. When integrated with the metabolic regulator metformin, the system couples exogenous photosynthesis and endogenous glucose metabolism, synergistically reprogramming the wound microenvironment. In both mouse and pig diabetic wound models, this approach suppresses inflammation, promotes angiogenesis and expedites re-epithelialization, leading to adaptive tissue regeneration. Our findings reveal that inexpensive, invasive aquatic biomass can be upcycled into advanced therapeutic biomaterials, offering a scalable and sustainable framework for next-generation regenerative medicine. This study showcases the upcycling of invasive water hyacinth into photosynthetic microspheres which provide sustained oxygen evolution and metabolic regulation, significantly accelerating the healing process of chronic diabetic wounds.
|
 Your new post is loading...
Your new post is loading...