 Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 5:18 PM
|
Biological information can be encoded in signaling dynamics, which have been implicated in many physiological processes; yet the diversity of dynamic expression profiles driven by a single gene remains unclear. To explore this, we screen 80 chromatin-associated proteins (CAPs) for their potential to drive diverse dynamic gene expression profiles from the same genome-integrated reporter in yeast. Using locus-specific optogenetic recruitment and live-cell microscopy, we measure dynamic expression profiles within single cells. CAP recruitment elicits a range of responses varying in activation delay, strength, production rate, and noise. We find that promoter activity is characterized by graded, rather than switch-like, transitions. A kinetic model with three promoter states and a positive feedback loop successfully captures the key features of expression driven by each CAP. These results reveal the rich dynamic landscape possible from a single gene, offering insights into native cellular processes and enhancing gene expression control in synthetic biology.

|
Scooped by
mhryu@live.com
Today, 3:43 PM
|
IQ-TREE (https://iqtree.github.io/) is a widely used open-source software tool for efficiently inferring phylogenetic trees under maximum likelihood. Here, we present IQ-TREE version 3, the third major release of the software. IQ-TREE 3 significantly extends version 2 with new features, including mixture models as an alternative to partitioned models, gene and site concordance factors to quantify discordance between genomic regions, integration with phylogenomic divergence time estimation, and a fully-featured sequence simulator. The IQ-TREE 3 source code is available at https://github.com/iqtree/iqtree3.

|
Scooped by
mhryu@live.com
Today, 3:36 PM
|
High-quality datasets that span broad sequence diversity are essential for understanding protein sequence-function relationships beyond local mutational landscapes. Here, we applied Growth-based Quantitative Sequencing (GROQ-seq) to measure function across an 11,722 member protease library, comprised of natural homologs and AI-shrunken variants. This library spans vast sequence diversity, with Levenshtein distances of up to 245 and a mean pairwise sequence identity of 41% to TEV protease S219V. We identified sequence-divergent TEV protease homologs that preserve function against the native TEV protease substrate. These findings reveal the robustness of protease activity across highly diverse sequences. Here, we demonstrate the aptitude of the GROQ-seq assay for screening large, diverse protein libraries for function, enabling efficient data generation at scale for training machine learning models across broad sequence landscapes.

|
Scooped by
mhryu@live.com
Today, 3:03 PM
|
Scalable genetic circuits are essential for implementing complex functions in living cells. Toward this goal, RNA regulators can provide a much-needed parts library with added benefits of low metabolic load, design flexibility, and logic capacity. However, despite the great potential of synthetic RNA circuits, constructing such circuits with wide dynamic ranges and multiplexed regulatory cascades remains a challenge. To address this, we introduce RATEX (Ribosome-Assisted Transcriptional EXpression controller) by integrating a translation-to-transcription converter with synthetic RNA regulators, enabling a compact and scalable RNA-programmed circuit architecture. The RATEX platform repurposes a large library of well-characterized translation regulators with up to 1,492-fold gene regulation, while leveraging natural ribosome-mediated sensing of diverse environmental inputs, such as metabolites. We demonstrated multi-input logic processing with up to a 6-input OR logic gate for RNA inputs and hybrid 3-input logic gates to sense diverse metabolite and small-molecule inputs alongside RNA signals. Signal amplification with multiplexed combinatorial control of RNA outputs was achieved through multiplexed signaling cascades. Finally, the RNA- and metabolite-sensing 3-input AND gates were used to control cellular morphology and intracellular spatial organization. Together, the RATEX platform, with its scalable and modular architecture, offers a broad potential design space for synthetic biology and biotechnology.

|
Scooped by
mhryu@live.com
Today, 2:53 PM
|
In this study, we report an integrated strategy combining directed evolution and semirational design to engineer a high-performance Pyrococcus furiosus Argonaute (PfAgo) variant. The optimized variant exhibited an 8.85-fold increase in catalytic efficiency, substantially enhanced soluble expression, and improved tolerance to elevated salt and metal ion conditions, enabling sensitive nucleic acid detection with a detection limit in the picomolar range. Mechanistic analyses suggested that these improvements may arise from reinforcement of the dimerization interface to stabilize the enzyme’s active state, optimization of the catalytic center, and the introduction of charge-shielding salt bridges. Finally, a biosensing platform based on the engineered PfAgo achieved a detection limit of 10 fmol l−1 for tetracycline, demonstrating its strong potential for sensitive and robust detection. This work provides a generalizable framework for enhancing Argonaute performance and advances their application in field-deployable diagnostics across clinical, environmental, and food safety settings.

|
Scooped by
mhryu@live.com
Today, 2:35 PM
|
Science is losing knowledge it cannot afford to lose. Negative results go unpublished, hard-won expertise walks out the door with departing researchers, and preservation efforts remain fragmented. The consequences are wasted resources, duplicated effort, and missed discoveries. In this perspective, we argue that the research community can act now by embracing alternative dissemination channels, improving documentation best practices, and building sustainable digital infrastructure. We envision moderated platforms for sharing null results and practical know-how, community-driven standards, and AI-powered tools that lower barriers to implementation. With coordinated effort, science can become more open, efficient, and resilient for future generations. Science routinely discards valuable knowledge, from unpublished negative results to the tacit expertise lost when researchers move on. Here, the authors propose a roadmap for preserving this hidden knowledge through community-driven platforms, open standards, and AI-assisted documentation.

|
Scooped by
mhryu@live.com
Today, 2:16 PM
|
Rapid and accurate pathogen detection serves as a core component in infectious disease prevention and control, clinical diagnosis and treatment, and public health surveillance systems. Although traditional detection methods have been widely adopted in clinical practice, they still exhibit significant limitations in terms of detection speed, throughput, automation levels, and adaptability to complex samples. In recent years, artificial intelligence (AI) technology has provided novel technical pathways for pathogen detection by leveraging its strengths in feature learning, pattern recognition, and multidimensional data modeling. The core contribution of this review lies in providing a novel, integrated analytical framework that overcomes the limitations of existing reviews, which often focus on a single modality (such as imaging alone or molecular diagnostics alone). Based on this framework, this paper systematically reviews AI research progress in pathogen detection, focusing on typical applications of machine learning and deep learning algorithms in analyzing imaging data, molecular diagnostic data, sensor signals, microscopic images, and multimodal data. It summarizes AI’s enabling value in enhancing detection sensitivity, specificity, automation, and point-of-care capabilities. Concurrently, this paper delves into key challenges facing AI-assisted pathogen detection, including data standardization, model generalization, interpretability, and clinical translation. It also outlines future trends toward intelligent, integrated, and clinically deployable applications. This paper aims to provide researchers and clinicians in the interdisciplinary field of artificial intelligence, biosensing, and clinical medicine with a comprehensive reference and roadmap for future development.

|
Scooped by
mhryu@live.com
Today, 2:06 PM
|
xBind is an interactive, freely accessible, and fully configurable webserver for large language model (LLM)-enabled cross-molecular protein binding-site prediction. xBind leverages LLM embeddings from the ESM-2 model together with sequence- and structure-derived features to predict protein–protein, protein–DNA, and protein–RNA binding sites using symmetry-aware deep graph neural networks. The input to xBind is either a single-chain protein sequence in FASTA format or a monomer protein structure in PDB or mmCIF format and it outputs predicted residue-level binding sites of the input protein with its pre-selected interaction partner. The customizable xBind web interface provides: (i) choice of interaction partners including protein–protein, protein–DNA, and protein–RNA; (ii) on-the-fly AlphaFold-based protein structure prediction for sequence-only inputs; (iii) on-demand selection of the likelihood threshold for calibrating structure-aware binding site annotations; (iv) interactive and interpretable web-based results, including sequence and structural visualizations and plots of residue-level binding likelihoods with user-adjustable threshold calibration; and (v) extensive help information for usage and results interpretation through a web-based tutorial and guide. xBind is freely available at https://fusion.cs.vt.edu/xBind.

|
Scooped by
mhryu@live.com
Today, 1:40 PM
|
Elucidating gene function in highly redundant genetic programs such as signaling pathways is challenging in model and nonmodel plants with current whole-plant genetic screening tools. Many of these challenges could be overcome if screens were instead carried out using individual cells harboring genetic perturbations. Here we report a single-cell screening platform, PIVOT (protoplast isolation after virus overexpression in planta), to accelerate identification and functional characterization of plant genes. We use Nicotiana benthamiana as a heterologous host to test gene libraries arrayed in a single leaf. PIVOT harnesses viral superinfection exclusion to ensure single multiplicity of infection per cell during pooled library delivery. Additionally, we engineer a cell-surface protein as a phenotypic marker for isolating cells of interest from a heterogeneous population. Using this system, we recover regulators of cytokinin signaling from an Arabidopsis open reading frame library. We anticipate PIVOT will be broadly applicable for high-throughput, single-cell functional genetic screening across the plant kingdom. A pooled, cell-based, genetic screening platform in plants is used for the functional analysis of cytokinin signaling proteins.

|
Scooped by
mhryu@live.com
Today, 1:16 PM
|
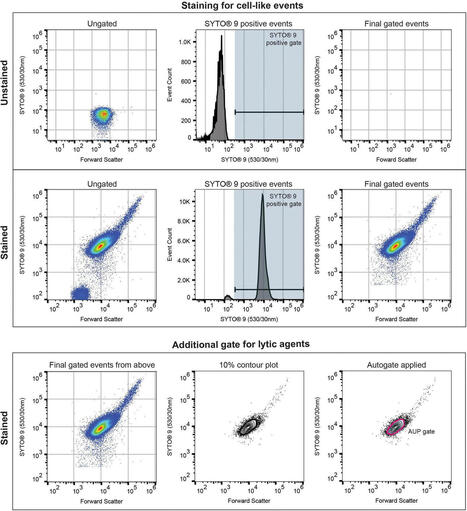
The global rise of multidrug-resistant bacteria necessitates the development of new antimicrobials and faster diagnostic tools. Conventional antimicrobial susceptibility testing is slow, relying on culture-based methods that delay effective treatment, often with fatal consequences in severe infections. In this study, we evaluate flow cytometry as a rapid, culture-minimal method to assess bacterial responses to six antimicrobials: ceftazidime-avibactam, meropenem-vaborbactam, cefiderocol, doxycycline, omadacycline, and lefamulin. Across 165 evaluable antibiotic-isolate combinations, essential agreement between flow cytometry and broth microdilution minimum inhibitory concentrations was 90.71%. Assessable categorical agreement, determined using the European Committee on Antimicrobial Susceptibility Testing and Clinical and Laboratory Standards Institute breakpoints, was 92.59% for doxycycline, 91.67% for omadacycline, and 100% for meropenem-vaborbactam. Cefiderocol exposure was associated with substantial cell elongation, demonstrating cellular-level antimicrobial effects observed using confocal microscopy and imaging flow cytometry. These findings demonstrate the potential of flow cytometry for novel antimicrobial evaluation, offering rapid insights into drug efficacy with potential to improve clinical outcomes in patients.

|
Scooped by
mhryu@live.com
Today, 1:29 AM
|
Does taking probiotics really matter? The idea is enticing. Swallow a capsule, add helpful microbes, support immunity, and strengthen the gut. Yet the microbiome is not a vacant landscape waiting for reinforcements. It is a densely woven ecosystem that behaves like an old-growth rainforest. Every niche is filled, every interaction balanced through biochemical negotiation, and any newcomer must face strong colonization resistance. With such a fortified system, what impact can a probiotic truly make? Most strains pass through the adult gut without becoming permanent residents. Still, they are not biologically inconsequential. During transit, they can influence epithelial barrier integrity, alter short-chain fatty acid and bile acid profiles, modulate immune signaling, and participate in cross-feeding interactions that reshape metabolic activity. These effects are best understood as functional ripples rather than structural reconfiguration. Accordingly, probiotic efficacy often reflects transient biochemical and host–microbe interactions, although the balance between transient activity and durable colonization depends on strain and formulation, dosing duration, host factors, and the baseline microbiome ecosystem, including recent disturbances such as antibiotics. Probiotic efficacy should therefore be evaluated using outcomes aligned with the intended mechanism, prioritizing clinical endpoints and biomarkers, supported by complementary compositional and functional microbiome readouts.

|
Scooped by
mhryu@live.com
Today, 1:22 AM
|
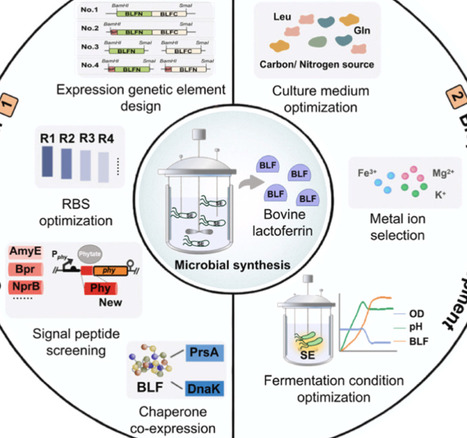
Bovine lactoferrin (BLF), a high-value multifunctional protein renowned for its iron-binding capacity and broad-spectrum antimicrobial properties. Despite its potential as a functional food ingredient, industrial application is restricted by the low abundance and high cost of extraction from natural sources. To address this bottleneck, this study establishes the first successful secretory expression of recombinant BLF in Bacillus amyloliquefaciens, a Generally Recognized as Safe (GRAS) host. Initially, the engineered strain yielded a titer of 18.0 ± 0.3 mg/L. To enhance secretion efficiency, a systems engineering optimization strategy was employed. This involved optimizing the ribosome binding site (RBS) and fusing a novel phytase-derived signal peptide (SPphy) to facilitate translocation. Crucially, the co-expression of the molecular chaperone PrsA was implemented to alleviate folding stress. These modifications culminated in a 6.1-fold yield increase, achieving a final titer of 110.0 ± 0.8 mg/L in a 5-L bioreactor. This research not only demonstrates the feasibility of B. amyloliquefaciens as a robust chassis for BLF production but also provides a strategic framework for the heterologous biomanufacturing of other complex nutrient proteins.

|
Scooped by
mhryu@live.com
Today, 12:58 AM
|
Agriculture is a major source of anthropogenic greenhouse-gas emissions, being the largest source of nitrous oxide (N2O), an extremely potent greenhouse gas and ozone-depleting agent. Soil N2O emissions are largely driven by microbial nitrification, in which ammonia-oxidizing microorganisms catalyze the rate-limiting oxidation of ammonia to nitrite. Nitrification not only mediates N2O fluxes but also reduces fertilization efficiency and contributes to eutrophication through nitrate leaching. Bacteriophage-based control of microbial communities is rapidly garnering interest in a number of fields; however, phages infecting ammonia-oxidizers are largely uncharacterized, with only one lytic phage having been described, limiting the potential for phage-mediated nitrification inhibition. Here, we show the largest set of phages infecting ammonia-oxidizing bacteria (AOB) to date: 45 dsDNA phages identified from urban wastewater, infecting four AOB species, with 16 demonstrating cross-genus host ranges and capable of eliminating nitrification activity in liquid cultures. Phylogenetic and taxonomic analyses revealed six proposed families of Caudoviricetes and numerous monophyletic clades, likely representing higher-level lineages. Structure-guided genome annotation revealed these phages to carry diverse and seldom-seen auxiliary metabolic genes, ranging from a complete ABC transporter cassette to a large antimicrobial resistance gene cluster. These results unveil the previously unrecognized diversity of AOB phages and their potential to alter host physiology. Our data demonstrates a broad taxonomic and functional repertoire of cultured AOB phages, greatly expanding the panel of known AOB phages, suggesting that viruses play a more significant and complex role in nitrification than previously understood. Moreover, we outline an effective methodological framework for isolating AOB phages from environmental samples. These results will help reframe our understanding of environmental nitrification and enable intensified selection and use of phages for its control.
|

|
Scooped by
mhryu@live.com
Today, 5:14 PM
|
Protein language models (pLMs) capture evolutionary sequence constraints but are limited in modeling underrepresented functional classes due to training data imbalance. Metalloproteins constitute a fundamental but sparsely represented class in sequence databases. We therefore assess whether structure-conditioned synthetic sequences can be used to specialize pLMs toward metal-binding functionality. We fine-tuned the generalist model ProtGPT2 on synthetic sequences generated by the inverse-folding model ProteinMPNN, constructing training sets with controlled variation in size and diversity. Fine-tuning increased recovery of canonical metal-binding motifs from 43% in the baseline model to 91% in the fine-tuned models. Generated sequences retained high predicted structural confidence and structural similarity to known folds, despite low sequence identity. Analysis of latent representations from ProtGPT2 indicated that fine-tuned models occupy distinct regions of embedding space relative to both the baseline model and structure-conditioned sequences, consistent with partial incorporation of structural constraints while preserving sequence diversity. A multi-step filtering pipeline applied to sequences lacking canonical motifs identified candidate metal-binding sites in four-helical bundle topologies not detected in a non-redundant subset of Protein Data Bank structures or in AlphaFold-predicted proteomes. https://doi.org/10.5281/zenodo.18672158 https://huggingface.co/gsgueglia

|
Scooped by
mhryu@live.com
Today, 3:38 PM
|
Although several existing protein-protein interaction (PPI) databases provide yeast PPI data, none unify large-scale network topology information with detailed biophysical, proteostasis, and regulatory annotations in a single protein-centric framework. To address this gap, we developed the ANnotated Yeast Interactome (ANYI), an open, integrated resource that combines experimental yeast PPIs with sixteen feature annotation types, including protein abundance, half-life, disorder content, post-translational modifications, conformational stability, chaperone interactions, sequence, and structure. ANYI integrates 3,927 proteins with 155 annotation features, forming a unified matrix that enables systematic cross-layer analyses. Available via GitHub and Docker Hub with an interactive network browser for broad accessibility, ANYI provides both experienced and beginner computational scientists with tools to investigate the yeast interactome. For example, users can directly test whether highly connected hub proteins exhibit distinct stability, disorder, or proteostasis signatures relative to peripheral nodes.

|
Scooped by
mhryu@live.com
Today, 3:25 PM
|
Long-term biodegradation of soil microplastics such as polyethylene terephthalate (PET) in situ remains inadequately addressed due to the limited expression of efficient PET degrading enzymes in engineered bacteria. Here, we developed a quorum-sensing (QS)-based protein expression system (XylS-LuxI/LuxR) that enhanced reporter green fluorescent protein (GFP) expression by 44-fold in E. coli. Using this system, we constructed whole-cell PET biodegraders expressing PET hydrolases (FASTPETase-MHETase) and leaf-branch compost cutinase (LCCICCG) in E. coli and Pseudomonas putida (P. putida). Soil-based assays using crude enzymes and E. coli XylS-QS-LCCICCG cells showed >80% degradation of bis(2-hydroxyethyl) terephthalate (BHET) within 30 min. Furthermore, Agde-LCCICCG was identified as the most effective signal peptide (SP) for protein secretion in E. coli, whereas LCCICCG without a SP performed best in P. putida. Engineered E. coli achieved up to 63% PET nanoparticle degradation over 30 days, while P. putida reached 42.3% within 20 days in nonsterilized soil, substantially outperforming wild-type controls and indicating synergistic interactions with native microbiota. These results demonstrate that XylS-QS-based systems enable efficient, self-regulated whole-cell PET biodegradation in soil environments, providing new insights for the development of efficient biodegradation strategies of environmental plastic waste.

|
Scooped by
mhryu@live.com
Today, 2:55 PM
|
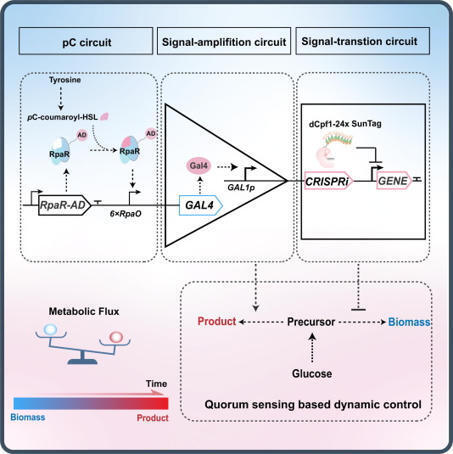
Dynamic regulation of metabolic pathways is critical for optimizing microbial production, yet robust quorum-sensing (QS) systems remain largely unavailable in eukaryotic microorganisms. Here, we establish a bacterial-derived QS platform in Saccharomyces cerevisiae by repurposing the noncanonical RpaI/RpaR system (a LuxI/R-type QS system), which produces p-coumaroyl-homoserine lactone as a signal molecule, bypassing a fundamental metabolic barrier that has prevented functional bacterial QS in eukaryotes. The engineered circuit features low leakage, high sensitivity, and broad dynamic range. By coupling QS with signal amplification and CRISPR interference modules, we create a bifunctional cascade system enabling autonomous transcriptional activation and repression. This QS platform enables growth-production decoupling and improves the production of cordycepin, geraniol, and 3-hydroxypropionic acid in both baseline and high-producing strains. Our work establishes a functional bacterial QS system in yeast and expands the synthetic biology toolkit for eukaryotic hosts.

|
Scooped by
mhryu@live.com
Today, 2:38 PM
|
ARTEM Server is an online platform for comparative analysis of nucleic acid 3D structures, combining two complementary superposition methods based on the ARTEM algorithm. The server provides access to searches for local tertiary motifs using the ARTEM tool, which identifies local isosteric structural arrangements without relying on sequence, interaction annotations, or backbone connectivity. It also offers global structure alignment and search modes via ARTEMIS, a recent extension of ARTEM that performs global sequence alignments based on rigid-body structural superposition. ARTEMIS supports both classical sequentially ordered superpositions and alignments involving sequence permutations, and can enumerate alternative suboptimal matches, enabling structural searches within large molecules or across databases. Benchmarks reported in the original publications demonstrate that ARTEM and ARTEMIS outperform other tools and are particularly effective at detecting 3D motif and 3D fold similarities across diverse backbone contexts, including cases that are challenging for sequence-ordered or annotation-dependent methods. ARTEM Server unifies these capabilities in a web interface, accepting PDB/mmCIF inputs, supporting multiple query and reference structures, and providing interactive 3D visualization and exportable alignment and motif-matching data. ARTEM Server offers a user-friendly web-based environment for exploration of global nucleic acid folds and local tertiary motifs. The web server is available at https://artemserver.genesilico.pl/.

|
Scooped by
mhryu@live.com
Today, 2:30 PM
|
Bioorthogonal chemistry was originally developed for fast and selective ligation of small molecules onto biomolecules in complex biological environments. Over time, the field has grown significantly beyond just clicking molecules together. It now includes reactions that can form or break chemical bonds and control how reactive substances are generated within living organisms. The field is continuing to develop strategies that allow reactions to be turned on when needed, exhibit greater compatibility in biological environments, and enable multi-step transformations to achieve complex operations. In this review, we summarize these new developments, including new click reactions and reagents, inducible and enzyme-activated click systems, click-induced bond dissociation reactions, bioorthogonal chemical operations, and bioorthogonally activated reactive species.

|
Scooped by
mhryu@live.com
Today, 2:08 PM
|
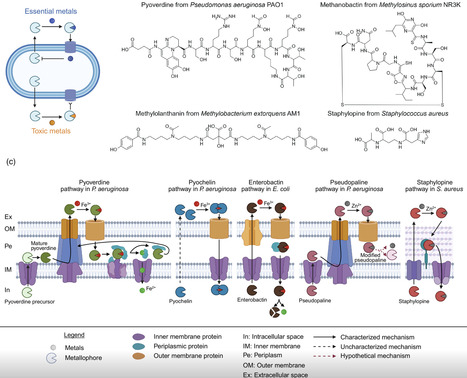
Metals are essential trace elements for almost all organisms including bacteria. Yet, metals are toxic at high concentrations, requiring fine-tuned regulatory mechanisms to steer metal homeostasis inside cells. In this primer, we explain how bacterial metallophores – small secreted secondary metabolites – act as gatekeepers by carefully orchestrating the scavenging and uptake of essential metals whilst preventing intracellular toxicity and keeping toxic metals outside the cell. We further introduce metallophore diversity together with main synthesis, secretion and uptake mechanisms. Finally, we show how secreted metallophores shape ecological interactions between bacteria and with eukaryotic organisms and how fundamental research on metallophores opens promising avenues for therapeutic and biotechnological applications.

|
Scooped by
mhryu@live.com
Today, 1:58 PM
|
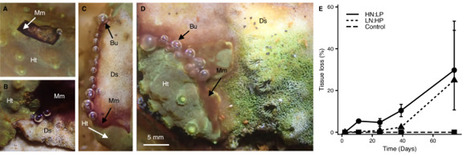
Coral diseases are increasing in prevalence, accelerating the global decline of tropical reefs, which threatens over 25% of marine biodiversity and vital ecosystem services for human societies. While outbreaks are frequently linked to environmental change, including heat stress, sedimentation, and reduced water quality, the mechanisms by which such factors promote disease remain poorly understood. Here we show that nutrient stress, caused by skewed seawater nitrogen-to-phosphorus (N:P) stoichiometry, promotes the onset of Black Band Disease (BBD), a common and easily recognisable syndrome that affects corals around the globe. Using Turbinaria reniformis as a model system, controlled laboratory experiments demonstrate that skewed N:P ratios disrupt the functional integrity of coral-associated microbial networks while favouring opportunists that exploit dysfunctional host–symbiont interactions. Disease lesion-associated microbial mats are dominated by cyanobacteria and include sulphur-metabolising bacteria, hallmarks of natural BBD communities. Strikingly, similar cyanobacterial taxa are also detected in the visually healthy coral tissue ahead of the expanding lesions, suggesting an opportunistic recruitment of disease-associated members from the resident microbiome. Global analyses of BBD outbreaks reveal that over 88% occurred in regions with skewed N:P ratios, compared with only 16% that were linked to prior heat stress. Together, our findings identify nutrient-driven microbiome destabilisation as a key pathway to coral disease, reinforcing nutrient management as a major lever for reef conservation and restoration practice. Coral diseases contribute to the decline of reefs around the globe. This study reveals that disruptions of the nutrient balance in seawater can change coral-associated microbial communities leading to disease.

|
Scooped by
mhryu@live.com
Today, 1:34 PM
|
The biological significance of the transition metal molybdenum (Mo) lies in its function at the catalytic center of several enzymes that drive a wide spectrum of redox reactions underlying global biogeochemical cycles, yet a paradox persists. While modern life ubiquitously relies on Mo, geochemical evidence suggests that its availability in early Earth’s anoxic oceans was extremely limited. Modern organisms can use Mo down to trace levels; however, the rates of Mo-dependent metabolisms slow down when Mo availability decreases, posing fundamental questions about the extent to which changing Mo abundances shaped the evolution of molybdoenzymes, and when early life began harnessing Mo. Here, we confront this evolutionary enigma by reconstructing the temporal and ecological emergence of molybdoenzymes, their transport systems, and biosynthetic pathways. In parallel, we examine biological tungsten (W) usage due to shared chemical properties and cofactor biosynthetic pathways with Mo. We provide molecular dating evidence of Mo/W utilization back to the Eo- to Mesoarchean (~3.7–3.1 Ga). These findings challenge prevailing assumptions about trace metal availability on the early Earth and underscore the profound antiquity and adaptability of Mo-based biochemistry in shaping early microbial evolution. The study shows that life began using molybdenum and tungsten enzymes as early as 3.7–3.1 billion years ago. Study reveals that key metabolic processes arose despite scarce metals and highlighting the early adaptability of microbial life.

|
Scooped by
mhryu@live.com
Today, 1:31 AM
|
Marchantia polymorpha has emerged as a promising model system for investigations in plant synthetic biology. Quantitatively characterizing plant genetic elements is fundamental to achieving predictable and controlled gene expression. However, only a few genetic parts are currently available for Marchantia. Additionally, the characterization of gene expression elements still relies on stable transformation assays. Here, we developed an Agrobacterium-mediated transient expression system to rapidly evaluate genetic parts in Marchantia. The entire experimental workflow can be completed within 8 days. Using this high-throughput system, we systematically benchmarked 21 promoters, 15 terminators, and 7 signal peptides from diverse sources. We identified a truncated CaMV35S promoter variant (P_35S-3), a native terminator (T_MpAct1), and a heterologous signal peptide (SP_SdMir) as top-performing elements. Notably, the P_35S-3 promoter exhibited a 409-fold activity increase over the standard CaMV35S promoter P_35S. Utilizing this potent element, we achieved an eGFP protein yield of 319.8 μg/g fresh weight in stable transgenic lines. The reliability of this transient system was further validated by stable transformation, where the two signal peptides exhibited a relative performance consistent with the transient assays. Our transient expression system provides a rapid and efficient platform for the characterization of gene expression elements, thereby expanding the genetic toolkit for Marchantia and enhancing protein expression levels.

|
Scooped by
mhryu@live.com
Today, 1:24 AM
|
Biomolecular condensates are formed through liquid–liquid phase separation (LLPS). They are highly dynamic, membraneless compartments within cells. The liquid-to-solid transition (LST) of these condensates plays a central role in regulating cellular physiological functions, maintaining tissue structural stability, and driving disease progression. Engineering LST has emerged as a major research frontier, integrating biophysics, synthetic biology, and materials science. This review systematically outlines the molecular grammar governing LST, key engineering strategies for its spatiotemporal control, and emerging applications in designed biological systems. We further discuss current challenges and future directions for harnessing LST as a design principle in systems chemistry and synthetic biology.

|
Scooped by
mhryu@live.com
Today, 1:02 AM
|
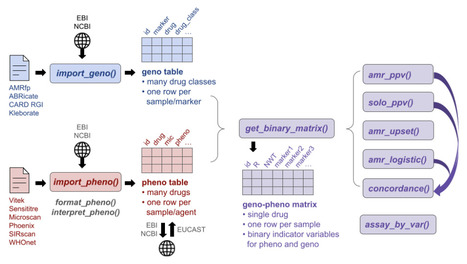
Microbial whole-genome sequence data is now generated at scale, including to support antimicrobial resistance (AMR) surveillance and understand resistance mechanisms, yet analytical infrastructure for systematically linking AMR genotypes to measured phenotypes remains fragmented. Here we present AMRgen, an R package to support systematic AMR genotype-phenotype analysis. AMRgen imports and harmonises genotypic data from common bioinformatics tools, alongside phenotypic data from automated antimicrobial susceptibility testing instruments and public repositories. It supports common analyses linking data to reference distributions, modelling associations, quantifying concordance, and producing publication-ready visualizations including UpSet plots that jointly display genotypic marker combination frequencies and associated phenotypic distributions. We demonstrate AMRgen's utility using publicly available surveillance data for World Health Organization priority AMR pathogens, Neisseria gonorrhoeae, Klebsiella pneumoniae, Escherichia coli and Salmonella enterica. AMRgen, available free and open-source at https://AMRgen.org provides a reproducible end-to-end foundation for genotype-phenotype research in AMR genomics, clinical microbiology, and public health surveillance.
|
 Your new post is loading...
Your new post is loading...




























Kortemme t,