 Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 12:25 AM
|
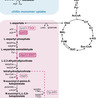
Despite its industrial importance, microbial L-lysine production has largely been confined to classical producer strains, leaving the fast-growing, non-pathogenic marine microorganism V. natriegens largely untapped as an unconventional biosynthetic platform. In this work, we established an L-lysine-overproducing V. natriegens DSM759 strain through a step-wise, systematic rational engineering strategy targeting the native biosynthetic pathway. Guided by our prior systems-level analysis of the strain′s genetic and regulatory architecture, we identified key metabolic bottlenecks and implemented knowledge-driven interventions to relieve pathway constraints. Central to production was alleviation of feedback inhibition in the native key regulatory enzymes, aspartate kinase (AK, lysC) and dihydrodipicolinate synthase (DHDPS, dapA). Site-directed amino-acid substitutions, replicating established E. coli feedback-resistance mechanisms, were introduced into conserved regions of the V. natriegens DSM759 enzymes, producing L-lysine-insensitive variants with kinetic parameters comparable to that of corresponding wild type enzymes. Among the tested configurations, the strain co-expressing Vn.lysC2 and Vn.dapA1:E84T reached the highest L-lysine titer (9.0±0.6 mM) and yield (0.11±0.01 molLys molGlc-1), whereas overexpression of additional L-lysine pathway genes provided no further benefit. Leveraging the host′s metabolic versatility, L-lysine synthesis was also demonstrated from the chitin-derived amino-sugar N-acetylglucosamine (0.09±0.00 molLys molGlcNAc-1), highlighting the potential to valorize chitin-rich waste streams from the seafood industry. This work establishes a minimal, rational strategy for L-lysine biosynthesis in V. natriegens DSM759 and positions it as a promising platform for sustainable amino acid production.

|
Scooped by
mhryu@live.com
April 11, 11:38 PM
|
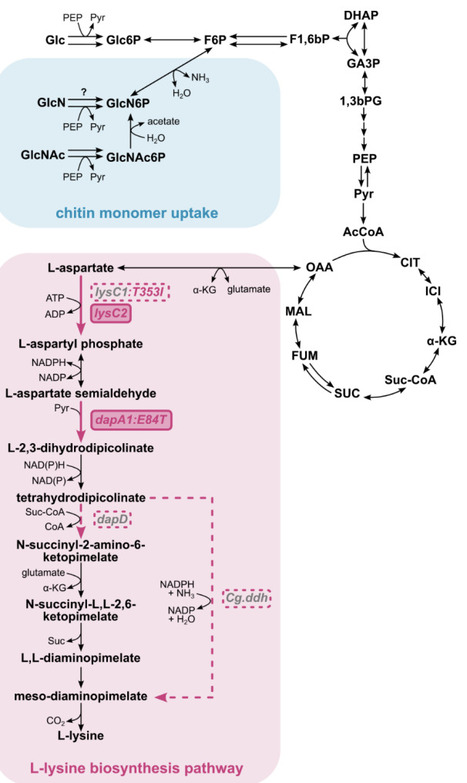
The capacity to adapt to complex environments (adaptability) is a defining property of cells. Yet, its relationship across single-cell physiology, population-level responses, and evolutionary timescales remains unclear. We performed single-cell transcriptional profiling of budding yeast across 20 complex environments before and after long-term selection for increased osmotolerance. In ancestral populations, transcriptional responses organized into a reproducible hierarchy where adaptation to certain cues took precedence over others. This hierarchy mechanistically originated within individual cells through contingent regulation of translation initiation. Evolution for >3,000 generations under osmotic stress increased osmotolerance while reordering this hierarchy, selectively deprioritizing osmotic stress as an organizing axis of adaptation. The derived strain exhibited impaired integration of stress responses, defective translational coordination, and reduced fitness outside the selected condition. Together, these findings illustrate that cellular adaptability is an evolved architecture whose form is set by the history of selection.

|
Scooped by
mhryu@live.com
April 11, 11:18 PM
|
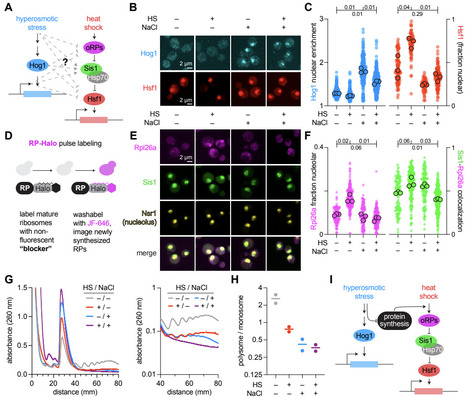
Upon infecting a bacterial cell, temperate phages make a decision between lysis and lysogeny. While research has previously explored how phages sense environmental information to make this choice, most studies have focused on modelling known mechanisms that impact the decision. These mechanisms tell us what environmental information the phage does respond to, but not what it should respond to, as the signals sensed by the phage may serve as proxies for other sources of information. Here, using a mechanism-agnostic population dynamics model, we find that irreversible phage binding to lysogens protects sensitive host cells from infection. This results in lysogens being an additional environmental factor that the phage should sense while making its decision to undergo lysis or lysogeny. Using this model, we derive a responsive lysogeny probability for phages that respond to both cell and lysogen densities optimized towards invading phage-occupied systems, and show that it is more capable of invading and resisting invasion than phage with fixed lysogeny probabilities across different environmental conditions.

|
Scooped by
mhryu@live.com
April 11, 5:10 PM
|
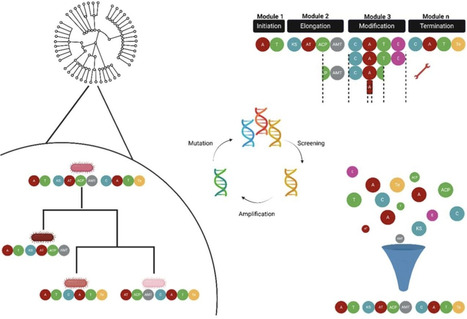
Engineering of non-ribosomal peptide synthetase (NRPS) and/or polyketide synthase (PKS) assembly lines to generate modified products has long offered promise to produce novel antibiotics and other bioactive molecules. However, it is only in recent years that this promise has been realised with any consistency. Key to this has been a shift away from engineering approaches informed solely by structural data, and towards strategies that incorporate insights from evolutionary principles and datasets. Such analyses have not only guided the selection of optimal recombination boundaries for substitution of key subdomains, domains or modules, but also methods for increasing engineering throughput, often trading accuracy for volume. Diverse approaches have proven successful in NRPS systems, but a consistent theme has been that recombinant assembly lines are generally impaired in terms of product yield, and a meta-analysis of published results to date indicates that no one engineering strategy is significantly best for minimising yield losses. Evolution-inspired strategies have advanced the engineering of, and product yields for, PKS systems, and further breakthroughs appear imminent. Although no ‘one size fits all’ solution is apparent for either NRPS or PKS engineering, this review highlights important advances in synthetic biology that will support both discovery and production of next-generation antibiotics.

|
Scooped by
mhryu@live.com
April 11, 5:07 PM
|
Anaerobic digestion (AD) of food waste (FW) is a key wate-to-energy strategy, yet daily biogas yield is often challenging to sustain, partly due to a limited understanding of the internal methanogens and their functional divergence. Here, we investigated seven full-scale mesophilic FW-AD systems distributed across China along a broad latitudinal gradient (>2,800 km), linking methane production variations (0.38–2.11 m3/m3•d-1) with the phylogenetic distributions of methanogens and their methanogenic genes. We found that hydrogenotrophic and aceticlastic pathways were ubiquitous, whereas methylotrophic methanogenesis showed regional enrichment in warmer regions, reflecting persistent influences of climate-associated upstream conditions on downstream methanogenic communities. Gene-level phylogeny of methanogenesis-related alleles, rather than species-level phylogeny, closely tracked biogas yield variation (Mantel's P < 0.05) and showed consistently stronger associations than gene-level compositions (mean standardized total effect: 0.491 vs. 0.298, P < 0.01). Higher methane yields (1.61 vs. 0.61 m3/m3•d-1 in high- vs. low-performing systems, P < 0.01) were significantly associated with reduced Faith's phylogenetic diversity (1.82 vs. 2.30, P < 0.01) and tighter clustering (mean pairwise phylogenetic distance, MPD: 0.25 vs. 0.30, P < 0.01) of methanogenic gene variants, suggesting that phylogenetic coherence may reflect ecological filtering favoring efficient methanogenesis, albeit at the expense of functional redundancy. These findings highlight gene-level trait phylogeny as a potential proxy for functional robustness, offering a framework for ecological design of AD microbiomes.

|
Scooped by
mhryu@live.com
April 11, 4:46 PM
|
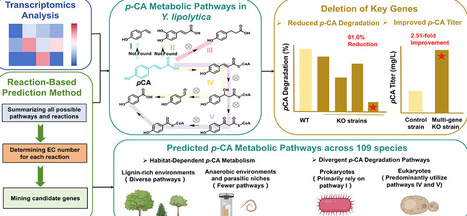
p-Coumaric acid (p-CA) is a universal precursor of plant lignin and polyphenols. A comprehensive understanding of p-CA metabolism is essential for elucidating biological evolution, advancing polyphenol biomanufacturing, and developing lignocellulosic biorefineries. However, p-CA metabolism remains poorly characterized due to the limited number of studies and the absence of universal prediction methods. Here, we systematically elucidate p-CA metabolism in the industrial yeast Yarrowia lipolytica. Through transcriptomic analysis and a reaction-based prediction method, we identify key degradation genes in Y. lipolytica, whose collective deletion reduced p-CA degradation by 81.6% and increased production by 2.51-fold. Comparative analysis of 109 species revealed that the metabolic potential for p-CA is widespread but exhibits significant habitat-dependent divergence. This study establishes a systematic framework for understanding and engineering p-CA metabolism, providing insights for optimizing microbial cell factories and a broader exploration of metabolic pathways.

|
Scooped by
mhryu@live.com
April 11, 4:31 PM
|
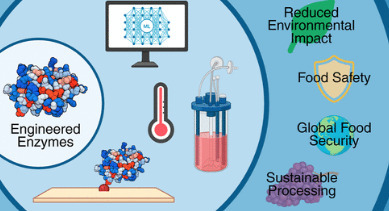
Global food insecurity remains a challenge, with 2.3 billion people worldwide experiencing food insecurity. Applications of synthetic biology offer a promising way to address this crisis through innovative and sustainable enzyme-mediated solutions. This review explores enzymology with food and agriculture systems and how recent advances are aided by synthetic biology. Focusing on how enzymes can be engineered for the greatest good to promote food safety, improved production, and coproduct valorization, we survey state-of-the-art advances in enzyme engineering to achieve these goals, providing a critical review on how technology from other industries could be adapted to food and agriculture. Key areas discussed include biocatalysis of food ingredients, synthetic biology for yield improvements, and computation design of enzymatic pathways for more resource-efficient food processing. This review concludes with a discussion of current limitations, regulations, and future directions for integrating synthetic biology into global food systems.

|
Scooped by
mhryu@live.com
April 11, 4:11 PM
|
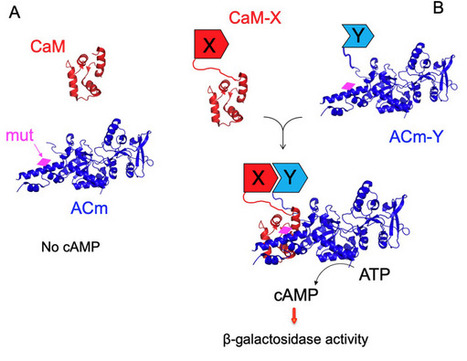
Protein–protein interactions are central in all biological processes. Methods capable of detecting interactions within living, intact cells have been particularly useful to identify and characterize protein interaction networks. We describe here an exquisitely sensitive regulatory circuit that can detect in bacteria, protein–protein interaction with single-molecule sensitivity. This approach involves the interaction-mediated reconstitution of a cyclic AMP signaling cascade in E. coli taking advantage of the high catalytic activity of the adenylate cyclase (AC) from Bordetella pertussis upon activation by its natural activator, calmodulin (CaM). We show that less than one complex of interacting hybrid proteins per cell on average, is enough to confer a selectable trait to the host. This exquisitely sensitive adenylate cyclase hybrid (ESACH) system allows for direct selection, in living bacteria, of ligands exhibiting high affinity for given targets or for studying interactions involving toxic proteins. The extreme sensitivity of the AC/CaM/cAMP signaling cascade may thus be harnessed to interrogate biological processes with single-molecule resolution in live bacteria and could be exploited to design novel synthetic regulatory networks operating at, or even below, the theoretical threshold limit of one molecule per cell.

|
Scooped by
mhryu@live.com
April 11, 3:47 PM
|
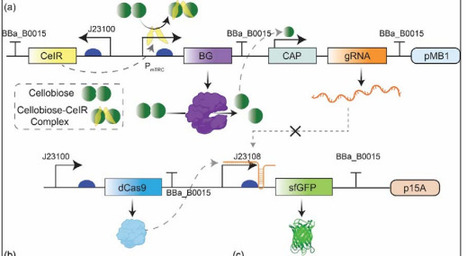
Precise monitoring of intracellular glucose dynamics is essential for understanding carbon flux, optimizing microbial bioprocesses, and enabling responsive control of engineered metabolic pathways. Here, we develop a modular whole-cell biosensor in Escherichia coli that converts the native glucose repression phenotype of a CAP-sensitive promoter into a tunable, glucose-inducible output using CRISPR interference (CRISPRi). By placing a guide RNA (gRNA) under the control of the CAP promoter and positioning dCas9 to target the -10 region of a constitutive promoter driving sfGFP, we created an inversion circuit in which glucose suppresses gRNA expression, thereby relieving dCas9-mediated repression and activating fluorescence. Systematic evaluation of gRNA strand orientation and target site selection revealed that template-strand targeting yielded strong repression (~90 %) but reduced sensing range, whereas moderately repressive non-template gRNAs (~27-35 % repression) enabled optimal signal inversion. The resulting biosensor demonstrated a robust, linear fluorescence response across 200 μM -50 mM glucose (R2 > 0.97), with high specificity against other sugars and a strong correlation between glucose consumption and fluorescence accumulation (R2 ≈ 0.996). To extend the functionality of the platform, we integrated the sensor with a secreted β-glucosidase module that hydrolyzes cellobiose to glucose. The biosensor accurately quantified glucose released during cellobiose degradation, with engineered strains producing up to 33 mM glucose from 50 mM cellobiose in a two-plasmid system. This coupling of enzymatic conversion with intracellular sensing enabled real-time, non-destructive monitoring of metabolic transitions. Together, this work establishes a programmable CRISPRi-based strategy for inverting native promoter logic and provides a sensitive, specific, and modular platform for metabolite sensing in bacteria. The approach is broadly applicable for dynamic pathway regulation, monitoring carbon fluxes, and building responsive genetic circuits in metabolic engineering and synthetic microbial ecosystems.

|
Scooped by
mhryu@live.com
April 11, 3:31 PM
|
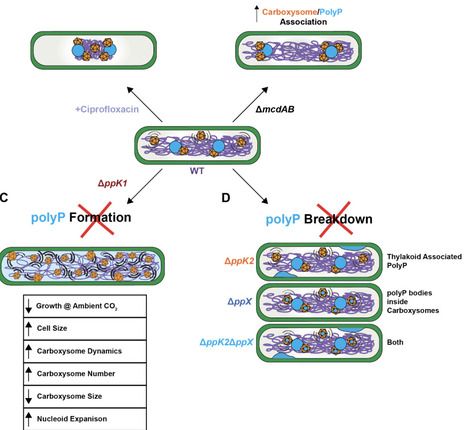
Polyphosphate (polyP) is a conserved inorganic polymer traditionally viewed as a stress-induced phosphate and energy reserve. In cyanobacteria, however, polyP granules are constitutively present and frequently observed in proximity to carboxysomes, the bacterial microcompartments that mediate CO2 fixation. Here we show that polyP functions as a spatially organized regulator of the photosynthetic cytoplasm in Synechococcus elongatus. PolyP granules localize to the nucleoid and are periodically arranged along the cell axis, independently of the McdAB carboxysome positioning system. Despite this independence, polyP and carboxysomes associate non-randomly, and this association is enhanced when active carboxysome positioning by the McdAB system is disrupted. Loss of polyP synthesis leads to nucleoid expansion, an increased number of smaller carboxysomes with high mobility, and severe defects in growth under ambient CO2. Perturbation of polyP turnover further reveals structural connections to both carboxysomes and thylakoid membranes. Together, these findings identify polyP as an architectural integrator that couples chromosome organization, metabolic compartmentalization, and photosynthetic fitness.

|
Scooped by
mhryu@live.com
April 11, 3:16 PM
|
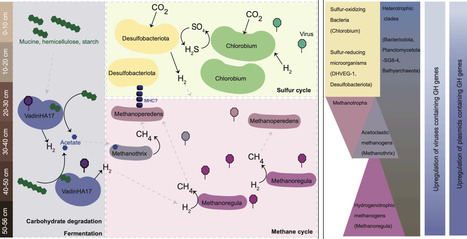
Anaerobic carbon transformation in freshwater sediments drives substantial methane emissions globally, yet the microbial taxa linking complex carbon degradation to methane production remain poorly characterized. Here, we combined metagenomics with the first metatranscriptomic dataset from the anoxic sediments of meromictic Lake Cadagno (Swiss Alps) to identify the active microbial clades, metabolic pathways, and extrachromosomal elements (ecDNA) across a depth gradient within the upper 56 cm of sediment. We recovered 802 species-level metagenome-assembled genomes (MAGs) spanning 66 phyla and identified a Bacteroidota clade (VadinHA17) as one of the most abundant and transcriptionally active populations in the sediment. This clade encodes and transcribes a broad range of diverse glycoside hydrolases (GH), indicating a central role in complex carbohydrate degradation. Transcriptional profiles suggest that this clade ferments organic substrates to acetate and hydrogen, which are key substrates for methanogenesis. In line with this, the acetoclastic methanogen Methanothrix and hydrogenotrophic Methanoregula were among the most abundant and transcriptionally active archaea in the same depth layers. Beyond microbial genomes, we detected 86,905 viral OTUs (vOTUs) and 2,136 plasmid OTUs (pOTUs), with free viruses and plasmids accounting for 5-10% and 0.2% of all sequencing reads, respectively. Notably, plasmids and viruses associated with Bacteroidota VadinHA17 encode and transcribe GHs that could augment host carbohydrate-degrading capacity. Together, these findings reveal new details on how methane production in anoxic lake sediments emerges from a network spanning primary fermentation, methanogenesis and ecDNA-mediated metabolisms.

|
Scooped by
mhryu@live.com
April 11, 2:54 PM
|
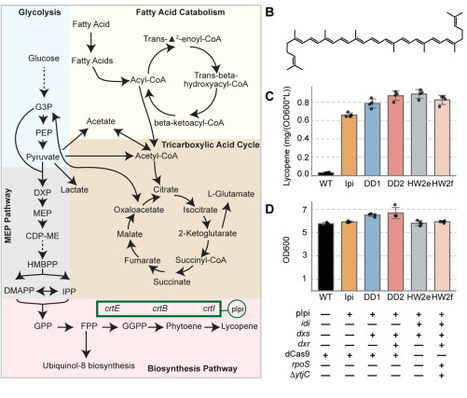
Isoprenoid biosynthesis in E. coli is a promising source of high-value natural products. However, isoprenoid production imposes a substantial metabolic burden on cellular carbon and cofactor metabolism. Optimizing yields thus requires rebalancing gene expression throughout metabolism. To systematically identify the genes in E. coli central metabolism shaping isoprenoid yield, we established a high-throughput CRISPR interference (CRISPRi) assay to examine the impact of gene repression on production of the bright red carotenoid, lycopene. Our approach enables repression of both essential and non-essential genes and links gene expression perturbations to pathway yield and biomass. We screened a CRISPRi library targeting 180 E. coli genes and found 31 genes for which repression significantly modified lycopene yield. Genes whose repression increased lycopene yield were found across fatty acid, amino acid, central carbon, and isoprenoid metabolism as well as stress response pathways. This set included four genes in amino acid biosynthesis, one in phospholipid biosynthesis, and one in the stringent response that, to our knowledge, have not previously been implicated in lycopene production. In contrast, genes for which repression decreased yield were limited to isoprenoid biosynthesis, central carbon metabolism, and stress response pathways. Together, our work reveals genetic targets for increasing lycopene yield throughout metabolism, and defines a tractable, generalizable approach to mapping genetic factors that modulate biosynthetic pathway yield.

|
Scooped by
mhryu@live.com
April 10, 5:01 PM
|
Biological nitrogen fixation in symbiotic diazotrophs is subject to oxygen regulation by an oxygen-sensing FixLJ two-component system under micro-oxic conditions. However, it remains unclear whether this mechanism is conserved in free-living diazotrophs. In this study, we discovered for the first time that FixLJ strongly inhibits the expression of nifHDK genes that encode molybdenum nitrogenase in response to oxygen. The deletion of fixLJ genes, whose expression was stimulated by oxygen, allowed a free-living photosynthetic diazotroph Rhodopseudomonas palustris to express active nitrogenase and grow diazotrophically even under oxic conditions. The unphosphorylated FixJ protein showed high-affinity binding to the promoter of nitrogenase gene cluster (PnifH) and strongly repressed the nitrogenase expression in response to oxygen. The transcriptional repression of nifHDK by FixJ reveals a new regulatory role for the FixLJ system. In addition, transcriptome analysis suggested that the FixLJ regulatory system also plays a role in the energy metabolism of R. palustris, probably through FixK regulation. This newly identified mechanism is speculated to allow R. palustris to rapidly shut down the synthesis of nitrogenase when exposed to oxygen, avoiding the build-up of nitrogenase with impaired activity due to the lack of protection from oxygen damage.
|

|
Scooped by
mhryu@live.com
Today, 12:17 AM
|
Intrinsically disordered protein regions are ubiquitous across all kingdoms of life. These structurally heterogeneous regions play central roles in cellular processes such as transcriptional regulation, cellular signaling, and subcellular organization, yet they have remained largely inaccessible to rational design. Structure-based generative methods are not applicable to proteins that lack a stable fold, and existing sequence-based approaches for disordered regions rely on sampling methods that do not capture the evolutionary statistics of natural disordered regions. Here, we introduce IDiom, an autoregressive protein language model trained on 37 million intrinsically disordered region sequences curated from the AlphaFold Database. Trained using a fill-in-the-middle data augmentation, IDiom generates disordered region sequences conditioned on their surrounding structured context, as well as fully disordered proteins without any context. The model generates diverse sequences that recapitulate biologically relevant sequence features of natural disordered regions, and we demonstrate that post-training via reinforcement learning with a subcellular localization reward model produces sequences with features which are consistent with known sequence determinants of compartment-specific localization. These results establish IDiom as a general platform for the generative design of intrinsically disordered proteins and regions.

|
Scooped by
mhryu@live.com
April 11, 11:31 PM
|
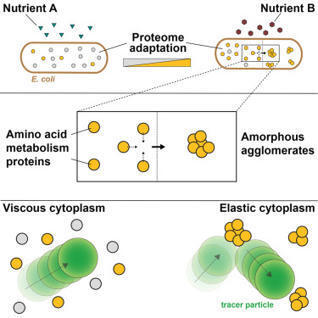
Molecular crowding in the bacterial cytoplasm restricts diffusion of large molecules, impacting cellular processes. To monitor cytoplasmic diffusion and rheology, we used single-particle tracking in E. coli, finding a 3-fold variation in the diffusion of a 40-nm particle across exponential growth conditions. Known determinants of rheology did not account for this variation. Instead, we found a strong anti-correlation between the diffusion coefficient and the abundance of amino acid metabolism proteins (clusters of orthologous groups [COG] category “E”), persisting upon genetic perturbations, and that lower diffusion is associated with increased elasticity. Photoactivated light microscopy revealed that some amino acid metabolism proteins form clusters. Electron microscopy showed that these proteins can form amorphous agglomerates at physiological concentrations in vitro due to their high hydropathy, which also confers low disorder and compactness. These findings show that diffusion is controlled by the formation of protein agglomerates and thus reveal how condition-induced proteome changes affect cytoplasmic rheology.

|
Scooped by
mhryu@live.com
April 11, 11:11 PM
|
The global rise of antimicrobial resistance (AMR) demands urgent attention. While genetic drivers are well studied, epigenetic mechanisms, particularly DNA methylation, are emerging as key contributors to bacterial adaptation under antibiotic pressure. This review examines the roles of N6-methyladenine (m6A), N4-methylcytosine (m4C), and 5-methylcytosine (m5C), each catalyzed by distinct DNA methyltransferases (MTases), in regulating resistance-related processes such as efflux pump expression, β-lactamase activity, and stress responses. Advances in long-read sequencing technologies, including SMRT and ONT, now enable single-base resolution detection of methylation and support strain-specific methylome mapping. These efforts reveal methylation patterns that are dynamic, strain-dependent, and environmentally responsive, complicating resistance profiling. Emerging applications for tackling methylation-linked AMR include methylation-aware diagnostics and CRISPR-based epigenetic editing. Tools like CRISPR-dCas9 fused to DNA methyltransferases enable targeted, reversible suppression of resistance genes regulated by methylation. Current findings position DNA methylation as both a regulator of AMR and a promising target for next-generation diagnostics and therapeutics. However, challenges remain, including the lack of validated biomarkers, inconsistent protocols, and difficulty interpreting mixed-species data. Integrating methylation profiles with transcriptomic and phenotypic data will be essential to fully understand and target resistance mechanisms.

|
Scooped by
mhryu@live.com
April 11, 5:09 PM
|
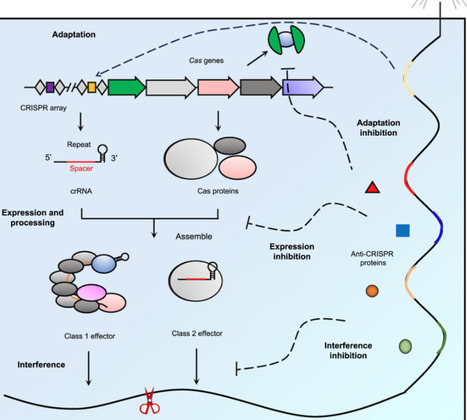
The CRISPR-Cas system constitutes an adaptive immune mechanism in prokaryotes that defends against mobile genetic elements. Within the perpetual co-evolutionary arms race between bacteria and their viral predators, bacteriophages encode anti-CRISPR (Acr) proteins that use sophisticated molecular strategies to sabotage CRISPR-Cas function. While canonical Acr proteins rely on steric blockade of Cas effectors, recent discoveries reveal unprecedented noncanonical mechanisms spanning CRISPR immunity stages. This review synthesizes recent mechanistic advances in this field since 2023, highlighting the expansion of noncanonical inhibition mechanisms beyond type I to include types II, V, and VI, as well as novel Acr interventions targeting multiple functional stages, such as spacer acquisition, translation-coupled inhibition, complex assembly/disassembly, and R-loop DNA binding. Structural insights demonstrate how Acr proteins achieve substoichiometric inhibition via conformational hijacking, catalytic repurposing, and molecular mimicry. Forged by the intense selective pressure of the phage–host conflict, these molecular innovations represent both remarkable evolutionary adaptations and versatile precision tools. They enable spatiotemporal control of CRISPR technologies, from engineered off-switches to diagnostic reset mechanisms, while posing critical challenges for therapeutic safety and microbiome management.

|
Scooped by
mhryu@live.com
April 11, 5:00 PM
|
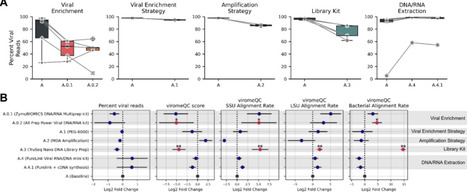
Human virome research is gaining increasing attention as viruses are recognized as critical modulators of microbial communities and human health. Viral metagenomics, however, faces unique challenges, including the low abundance and diversity of viruses in biological samples, the absence of universal marker genes, and biases introduced by experimental protocols. While various virome protocols have been benchmarked using viral particles or nucleic acids from mock communities, these approaches often fail to capture the complexity and heterogeneity of natural viromes. In this study, we systematically evaluated modifications to key methodological steps in the metagenomic analysis of human fecal samples, including viral enrichment, nucleic acid extraction, genome amplification, and library preparation. Using gold-standard bioinformatic approaches on sequencing datasets generated after amplification, we assessed the impact of these modifications on relative viral taxonomic assignment, contig quality, richness, diversity, and inferred genome structure. Our findings reveal striking trade-offs between recovery of viral genomes and retention of non-viral sequences, demonstrating how methodological choices can shape the inferred virome composition. Based on these observations, we propose an optimized protocol that enhances viral genome recovery while reducing contamination from non-viral sequences. This refined workflow provides a more robust and reliable framework for gut virome studies, paving the way for a deeper exploration of the role of viruses in human health and microbial ecosystems.

|
Scooped by
mhryu@live.com
April 11, 4:41 PM
|
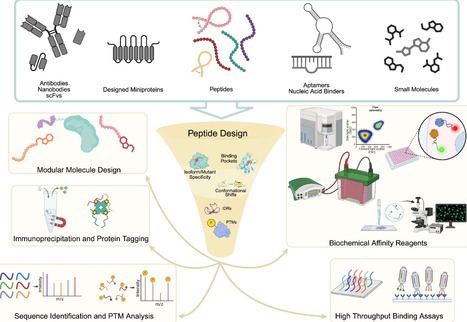
Peptides occupy a unique niche as biochemical tools: they are small, modular reagents capable of perturbing protein function with a precision that is often inaccessible to small molecules or antibodies. Historically, their broader use in biochemical research has been constrained by slow discovery workflows, limited control over specificity, and poor physicochemical properties. Recent advances in artificial intelligence have begun to change this landscape by enabling the rational, data-driven design of peptides tailored for specific experimental tasks. In this review, we focus on AI-designed peptides as practical tools for biochemistry. We survey sequence-based and structure-based design paradigms, highlighting how each supports distinct classes of peptide tools, including isoform- and motif-specific binders, multi-objective assay-ready reagents, and functional peptides that enable degradation, stabilization, or biophysical interrogation of proteins. By emphasizing experimental utility, design constraints, and appropriate use cases, we aim to provide a framework for selecting and deploying AI-designed peptides as on-demand reagents in modern biochemical research.

|
Scooped by
mhryu@live.com
April 11, 4:28 PM
|
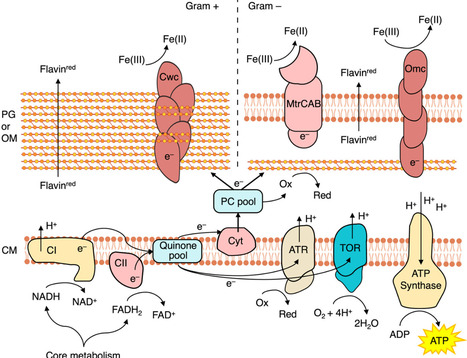
Microorganisms have historically been classified as obligate aerobes or anaerobes, facultative anaerobes, or microaerophiles, reflecting differences in respiratory strategies dictated by the use of oxygen and alternative electron acceptors. Recent discoveries provide evidence that a deviant strategy, the concurrent reduction of oxygen and other electron acceptors, is more widespread than previously thought. Such co-respiring bacteria employ hybrid metabolic strategies that extend models of electron acceptor use. In this review, we investigate mechanisms of co-respiration, summarize the biochemical components enabling parallel electron flow, and discuss the regulation of aerobic and anaerobic pathways under changing redox conditions. We also examine the evolutionary context of these strategies during the rise of oxygen on early Earth and outline experimental approaches needed to resolve co-respiration in individual cells.

|
Scooped by
mhryu@live.com
April 11, 3:53 PM
|
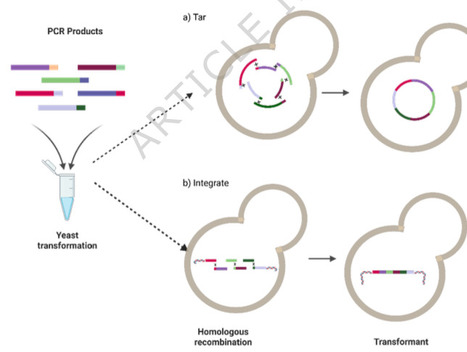
Saccharomyces cerevisiae is a widely used biotechnological workhorse in both academic and industrial settings. One reason for its continued popularity is the extensive legacy of genetic tools, developed over its long history of use, that enable precise manipulation of the S. cerevisiae genome. These tools have enabled extensive genetic characterization and dramatic re-programming efforts for applications ranging from fundamental research to industrial chemical production. However, existing software is often designed for automated pipelines and can be cumbersome to integrate into existing laboratories. Here we present a digital toolkit called PYEAST (Python Enabled Automated Strain Transformation) that encodes some of the most widely used methods for working with S. cerevisiae, modernizes them to leverage advances in DNA synthesis, and facilitates digital sequence management.

|
Scooped by
mhryu@live.com
April 11, 3:34 PM
|
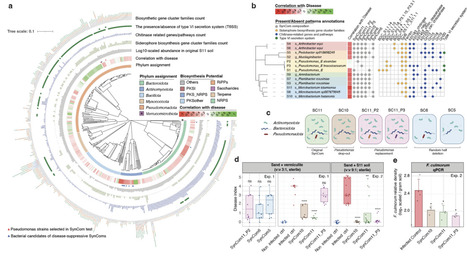
Across the biosphere, microbiomes play essential roles in shaping the health of their host. One notable example of such a microbiome-associated phenotype is disease-suppressive soils, where susceptible plant hosts enrich and activate specific rhizosphere microbial consortia for protection against fungal root pathogens. However, identifying and reconstructing microbial consortia responsible for host protection remains challenging, given the inherent taxonomic and functional complexity of microbiomes. Here, we integrated metagenomic profiling of disease-suppressive microbiomes perturbed by dilution-to-extinction (DTE) with comprehensive culturing and synthetic ecology to identify the key bacterial taxa conferring suppressiveness to the fungal wheat pathogen Fusarium culmorum. Metagenomics of wheat rhizosphere samples along the DTE trajectory revealed bacterial taxa and functions associated with the disease-suppressive phenotype. Crosslinking these DTE metagenome data with a genome-sequenced collection of 336 rhizobacterial isolates from the suppressive soil allowed the reconstruction of synthetic communities (SynComs) of 11 de-replicated strains negatively associated with disease severity. Upon re-introduction in sterilized suppressive soils, this SynCom consistently reproduced the disease-suppressive phenotype. Paired time-series metagenomics and metatranscriptomics of the SynComs pinpointed candidate biosynthetic gene clusters, including a novel non-alpha poly-amino-acid (NAPAA) gene cluster from Arthrobacter, upregulated in presence of F. culmorum. Chemically synthesized NAPAA variants Ɛ-poly-L-lysine and δ-poly-L-ornithine significantly inhibited F. culmorum hyphal growth. Collectively, our work establishes a transformative strategy for reconstructing microbial consortia that recapitulates beneficial microbiome-associated phenotypes in plant and animal kingdoms.

|
Scooped by
mhryu@live.com
April 11, 3:27 PM
|
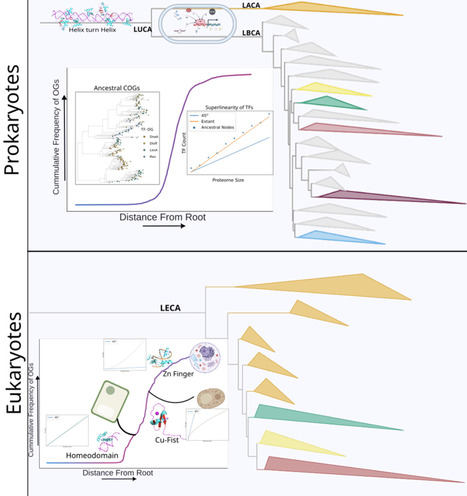
Transcription initiation is regulated by proteins called transcription factors (TFs). Though TFs help determine phenotype across the tree of life, they are nonessential for minimal cellular life and are often absent in endosymbiotic and parasitic organisms. Given this and the idea that it is a certain level of organism complexity that calls for specific transcription regulation, we traced the evolutionary history of TF repertoire on a bacterio-archaeal tree of life using a dataset of ∼500,000 TFs, grouped into ∼1,700 orthologous groups (OGs) across ∼3,000 species. The most ancestral prokaryotes encoded multiple TFs. Going by known extant functions of these TFs, they possibly regulated sugar-fermentation metabolism, sensed overall metabolic state and redox, responded to DNA damage or bound metals; many of which are consistent with some reconstructions of ancestral gene pools and physiologies. The number of TFs as well as their superfamily-level diversity, through evolutionary history, matches expectations against genome size derived from extant bacteria, suggesting pre-LUCA diversification of TF sequence families. Emergence of new TFs along the phylogeny largely followed a smooth cumulative distribution curve, suggesting steady innovation, early in prokaryote evolution, in contrast to eukaryotes, in which a majority of TF families emerged in a burst manner at the ancestors of multicellular lineages. Gains of TFs late in prokaryotic evolution predominantly featured recycling of protein families discovered elsewhere in the prokaryotic tree, consistent with the dominance of horizontal gene transfer in these organisms. We speculate on the difference between the evolutionary trajectory of prokaryotic TF repertoire and compare it with the eukaryotic TF repertoire trajectory. This helps us in understanding the manner in which their TF repertoires have evolved in two different super-kingdoms. The difference between the evolutionary dynamics of TF- repertoires might be due to how complexity is envisioned in these two different kingdoms.

|
Scooped by
mhryu@live.com
April 11, 2:57 PM
|
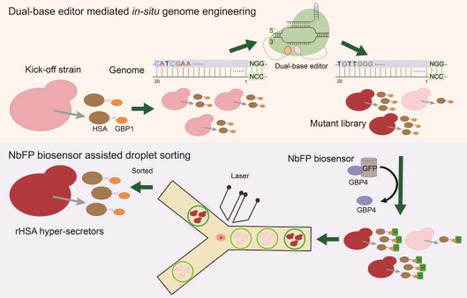
The disparity between the production and demand of recombinant proteins (r-proteins) has significantly hindered their commercial viability. Leveraging genomic resources offers substantial promise in enhancing our comprehension of metabolic and regulatory networks, thus facilitating the development of highly productive protein cell factories. However, the considerable gap between high-throughput strategies for monitoring r-protein secretion and genome perturbation in P. pastoris continues to obstruct the systematic linkage of genotype and phenotype, thereby limiting the optimization of production. Here, we developed a novel strategy combining dual-base editor-mediated in-situ genome engineering with nanobody-regulated biosensor-assisted droplet sorting to enhance r-protein secretion (BINDER) in P. pastoris. We successfully employed BINDER to screen recombinant human serum albumin (rHSA) hyper-producers and identified two critical SNVs conferring up to a 1.78-fold improved secretion titer from 113632 mutants, providing valuable insights into the secretion mechanism. Fed-batch cultivation of the engineered strain resulted in the highest reported rHSA titer, 23.43g/L, in P. pastoris, demonstrating its substantial potential for industrial applications. Given the high transferability of base editors and the novel biosensor's independence from the properties of the target protein, the strategy developed here might be expanded to a variety of microbial species and r-proteins.

|
Scooped by
mhryu@live.com
April 11, 2:18 PM
|
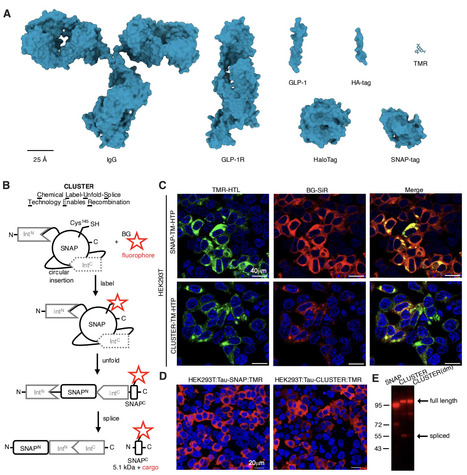
Precise and minimally perturbative protein labelling remains a key challenge for studying biomolecular function in living systems. Here, a minimal way to specifically label proteins based on SNAP-tag and intein-mediated protein splicing reaction is introduced. Termed CLUSTER (for Chemical Label-Unfold-Splice Technology Enables Recombination), this chimeric platform supports efficient labelling across diverse targets in living cells by retaining a fluorescent, 5 kDa sized peptide on a fusion protein of interest after splicing. A bacterial screening workflow was developed to optimize the reaction efficiency and construct design. Quantitative characterization using fluorescence polarization provides mechanistic insight into labelling efficiency and dynamics, while molecular dynamics simulations elucidate its stability, grasping the intricate nature of protein behavior upon covalent labelling. This bio-orthogonal labelling technology allows for a versatile and minimally invasive approach for protein labelling, providing a powerful tool to probe protein behavior in native cellular systems.
|
 Your new post is loading...
Your new post is loading...













Abudayyeh, Gootenberg, Proteins of interest go through iterative rounds of low-N screening. A foundational PLM generates embeddings for all mutants of a protein and the average embedding by pooling across all residues is used as input for the top-layer model. Each mutant’s activity is experimentally determined and used to train a domain expert top-layer model with PLM embedding as input. The top-layer model then nominates the top-N mutants for the next round of testing and the weights are updated iteratively in an active learning format.