 Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 12:27 PM
|
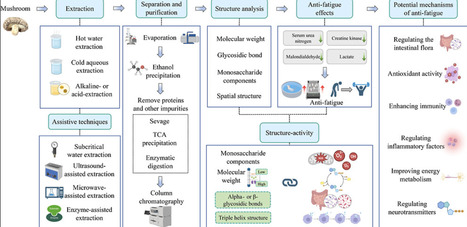
Exercise-induced fatigue is often accompanied by a poor mental state, reduced work efficiency, and decreased athletic performance. Mushroom polysaccharides have been shown in recent years to have a positive effect on alleviating exercise-induced fatigue. This review comprehensively analyzes the extraction methods, structure–activity relationships, and mechanisms of mushroom polysaccharides in alleviating exercise-induced fatigue from 1999 to 2025. Research has revealed that mushroom polysaccharides not only directly alleviate exercise-induced fatigue by regulating gut microbiota but also indirectly mitigate it by influencing oxidative stress, inflammatory factors, and energy metabolism. Moreover, the structure of mushroom polysaccharides is closely related to their biological activity, but there is no systematic consensus on the key structural feature responsible for their antifatigue effects. The purpose of this review is to provide new theoretical evidence for the relationship between the polysaccharide structure and activity and to pave the way for the development of antifatigue functional food ingredients based on mushroom polysaccharides.

|
Scooped by
mhryu@live.com
Today, 12:21 PM
|
Glycoside hydrolases, “glycosidases”, catalyze carbohydrate catabolism, remodeling, and signaling by accelerating glycosidic-bond cleavage by more than 17 orders of magnitude. Distributed across every kingdom of life and grouped into over 180 sequence-defined families, these enzymes exhibit exceptional diversity in fold, mechanism, and physiological function, and many also catalyze transglycosylation or phosphorolysis. The classical Koshland paradigms─stereochemical inversion, enzymatic nucleophile-assisted retention, and substrate-assisted retention─are analyzed with an emphasis on the conformational itineraries and oxocarbenium ion-like transition states revealed by kinetic isotope effects, linear free-energy relationships, and high-resolution three-dimensional structures. Attention then turns to noncanonical enzymes that employ NAD+-dependent redox hydrolysis or other cleavage mechanisms. Mechanistic insights have inspired the development of engineered glycosidase-derived catalysts for programmed bond construction as well as mechanism-based inhibitors, transition-state analogues, and activity-based probes that are driving advances in chemical biology, biotechnology, and drug discovery.

|
Scooped by
mhryu@live.com
Today, 12:08 PM
|
E. coli is a well-known host species for studies on recombinant protein expression, typically induced during the exponential phase. Nutrient depletion and the accumulation of toxic substances, which are common during the stationary phase, negatively impact recombinant protein expression, thereby limiting its suitability for efficient protein production. This study aimed to explore a physiological state distinct from conventional strategies for a complementary approach by investigating recombinant protein expression during the stationary phase. An unconventional stationary-like state was engineered using glucose to trigger acetate-mediated growth arrest and reduce tryptone to prevent regrowth. Under these conditions, the absolute and relative expression of signal peptide-fused human interferon gamma (mCsn2-SP-hIFN-γ) increased > 2.4 and 2.5-fold, respectively, compared with that in exponential-phase induction. The present findings suggest that regardless of the unfavorable conditions of the stationary phase for recombinant protein production, glucose utilization may enable the establishment of a stationary-phase model to address these limitations. Overall, these findings underscore the importance of the intracellular metabolic state and expression timing in optimizing recombinant protein production. Although only a single protein was examined in this study, the results highlight the potential of the unconventional stationary phase as an effective new strategy for enhancing recombinant protein expression.

|
Scooped by
mhryu@live.com
February 21, 11:53 PM
|
Bacterial conjugation is a major route of horizontal gene transfer and a key driver of the dissemination of antibiotic resistance, exacerbating the global health crisis. Central to this process is TraI, a bifunctional relaxase-helicase encoded by the iconic F-plasmid family. TraI initiates single-stranded DNA (ssDNA) transfer by covalently attaching to plasmid DNA through its trans-esterase (TE) activity and subsequently unwinds the plasmid via its helicase activity. Although genetic and biochemical studies have postulated that two TraI molecules are required for efficient conjugative transfer, how these molecules coordinate their activities in space and time - and how strand nicking is coupled to helicase loading - has remained unknown. Here, we capture the elusive, functionally asymmetric TraI homodimer and report its cryo-EM structure, revealing that the interaction between the two TraI molecules is mediated predominantly by ssDNA. We further show that TE-driven duplex melting at the origin of transfer (oriT) by one TraI molecule enables helicase loading by a second TraI molecule on the transfer strand of the unnicked plasmid DNA. Although the TE domain is intrinsically competent for strand nicking, its activity is inhibited by the host factor IHF, and this inhibition is relieved upon helicase loading. Together, these findings define a regulatory loop that coordinates helicase loading with strand nicking, providing mechanistic insight into the earliest stages of conjugative DNA processing.

|
Scooped by
mhryu@live.com
February 21, 2:05 PM
|
Periodontitis is a chronic inflammatory disease driven by subgingival dysbiosis. Despite progress in characterizing microbial diversity, functional heterogeneity at the single-cell level remains poorly understood, mainly due to challenges from low microbial biomass and host contamination. Here, we present a single-cell RNA sequencing framework for profiling subgingival bacteria, enabling high-resolution analysis of these communities in health and periodontitis. Using 16 subgingival samples, we generated an atlas spanning 133,458 cells across 285 species, including 57 core active species grouped into eight functional clusters. Health-associated subpopulations specializing in adhesion and polysaccharide degradation decline in periodontitis, coinciding with hypoxia and elevated amino acid availability. The keystone pathogens T. denticola and P. gingivalis display species-specific tendencies in amino acid metabolism-related transcriptional profiles, while P. intermedia harbors a proteolytic subpopulation enriched in periodontitis. These insights deepen our understanding of periodontitis pathogenesis and inform precision diagnostics and therapeutic strategies.

|
Scooped by
mhryu@live.com
February 21, 1:11 PM
|
Advancements in synthetic genetic circuits have enabled programmable and condition-dependent control of microbial cell growth. CRISPR-Cas9-based kill switches, genetic systems that program cells to lose viability in response to specific conditions, have recently been demonstrated for bacterial cell factories but not yet in yeast. Here, we present a foundational demonstration for a CRISPR-based kill switch in Saccharomyces cerevisiae, CRISPR KiSS. The CRISPR KiSS employs inducible CRISPR targeting essential genes to elicit growth inhibition. The activation of the KiSS system is achieved through conditional expression of a guide RNA (gRNA) upon anhydrotetracycline (ATc) induction, thereby activating CRISPR-mediated gene disruption. We demonstrate that targeting the essential genes (ERG13, PGA3, TPI1 or CDC19) leads to severe growth inhibition upon ATc induction. Still, the current set up does not allow complete killing of the cells due to system inactivation, e.g. escape from CRISPR based cutting. We studied reasons for system inactivation and substantially improved the system by simultaneous expression of two different gRNAs. Sequencing escape mutants revealed mutations in both the gRNA sequences and target genes as potential sources of system inactivation. This work highlights the potential of harnessing a CRISPR-based kill switch in S. cerevisiae. Cells expressing the system were able to escape growth inhibition through mutations and further optimization of the KiSS system is still needed for it to be used in various cell factory applications.

|
Scooped by
mhryu@live.com
February 21, 1:05 PM
|
Phages are a valuable resource for the genetic engineering of Streptomyces antibiotic-producing bacteria. Indeed, a few integrative vectors based on phage integrase are available to insert transgenes at specific genomic loci. Chromosome conformation captures previously demonstrated that the Streptomyces linear chromosome is organized in two spatial compartments: the central compartment encompassing the most conserved and highly expressed genes in exponential phase, and the terminal compartments enriched in poorly conserved sequences including specialized metabolite biosynthetic gene clusters. This study introduces a new integrative tool based on a recently described phage, Samy, which specifically targets the terminal compartment of its native host chromosome. Samy is related to PhiC31 phage and, like the latter, encodes a serine integrase. Whereas PhiC31 targets a site generally located near the origin of replication, the Samy integration site is one of the farthest known attB sites from it. We demonstrated that the Samy integrase efficiently mediates the specific integration of a non-replicating plasmid in six Streptomyces strains from distinct clades. Bioinformatic analyses revealed that the Samy-attB site is rather conserved and located in the terminal compartment of most Streptomyces chromosomes. Finally, heterologous expression of the albonoursin biosynthetic gene cluster from the Samy-, PhiC31-, and R4-attB sites yields quantitatively equivalent levels of production, though qualitative differences were observed. Altogether, these results demonstrate that the att-int Samy system expands Streptomyces genetic engineering tools by enabling targeted integration in the terminal chromosomal compartment.

|
Scooped by
mhryu@live.com
February 21, 12:56 PM
|
While extracellular vesicles (EVs) are established mediators of intra-species signaling, their contribution to cross-kingdom communication remains incompletely understood. Here, we investigate the EV-mediated interactions between human colon epithelial cells and both Gram-positive and Gram-negative gut bacteria. We show that bacterial EVs (BEVs) derived from Lacticaseibacillus casei, Enterococcus faecalis, and Proteus mirabilis induce distinct transcriptomic changes in Caco-2 cells depending on the bacterial species, with up to ~6,000 differentially expressed genes, including CCL20, CXCL8, or CXCL10. Transfection of BEV-derived RNA independently induces a subset of similar effects, indicating that the EV-mediated communication is partially driven by the RNA cargo. Conversely, we demonstrate that bacteria interact with Caco-2-derived EVs and miR-192-5p, which is highly abundant (~36.4-fold higher) in EVs isolated from conditioned medium compared with EVs from unconditioned medium, with modest effects on bacterial growth. Furthermore, we show that lipid-based packaging of miR-192-5p modulates its association with the bacteria. Our findings support a conceptual model in which EVs and their RNA cargo contribute to species-dependent host-microbe interactions. This study introduces a framework for understanding EVs as cross-kingdom regulators and underscores the importance of tailored, context-specific analyses for understanding the scope of EV-mediated interactions in microbiome-host homeostasis and disease.

|
Scooped by
mhryu@live.com
February 21, 12:30 PM
|
Cyanobacteria are known for their rich secondary metabolome with a long history of research directed toward the industrial and pharmaceutical applications of their natural products. Cyanobacterial metallophores (metal-chelating molecules), however, are understudied relative to metallophores from other phyla despite evidence suggesting that genes for metallophore biosynthesis are well-represented in cyanobacterial genomes. Many of the characterized cyanobacterial metallophores are formed from hybrid biosynthetic pathways and feature mixed coordinating functional groups, leading to enhanced structural and functional diversity. The few characterized metallophore families have intriguing properties including promiscuity of metal binding, photoreactivity, and amphiphilicity that are yet to be fully explored. Research suggests that these compounds are ecologically relevant and could guide community dynamics by controlling the availability of iron, detoxifying copper, and allelopathically inhibiting certain organisms. Cyanobacterial metallophores also have potential in the fields of therapeutic design, bioremediation, technology, and agriculture. In the past five years, the number of characterized metallophores from cyanobacteria has doubled, pointing to the great promise for future discoveries.

|
Scooped by
mhryu@live.com
February 21, 11:53 AM
|
Bacterial hypermutator strains drive rapid evolution of antibiotic resistance in chronic infections. Inspired by cancer therapy approaches that exploit synthetic lethality by targeting DNA repair deficiencies in hypermutator tumors, we tested whether pairing a conventional antibiotic with a secondary DNA-damaging agent could constrain hypermutator evolution in bacteria. Using high-throughput experimental evolution of E. coli repair-deficient strains, we evolved populations under carbapenem selection in combination with ciprofloxacin or mitomycin C. Strains lacking oxidative damage repair, double-strand break repair, or transcription-coupled repair showed significantly reduced evolvability, particularly under constant antibiotic pressure and increasing genotoxic stress. However, mismatch repair (MMR) hypermutators, the predominant clinical genotype, did not show reduced evolvability under these combination treatments. This is consistent with pathway orthogonality: MMR does not repair the structural DNA lesions induced by ciprofloxacin or mitomycin C, and the elevated mutation supply of MMR-deficient strains may allow rapid adaptation despite background DNA damage. Our findings demonstrate that combination strategies can constrain the evolvability of specific repair-deficient genotypes in vitro, but success requires matching DNA damage type to specific repair vulnerabilities. This work establishes proof of principle for genotype-directed antimicrobial strategies that exploit DNA repair vulnerabilities to constrain hypermutator evolution.

|
Scooped by
mhryu@live.com
February 21, 11:14 AM
|
Modeling protein conformational landscapes is essential for understanding dynamics, allostery, and drug discovery, yet existing resources lack diverse conformational coverage, energetic annotations, or benchmarking standards. ProteinConformers https://zhanggroup.org/ProteinConformers provides 2.7 million geometry-optimized conformations generated with a multi-seed molecular dynamics strategy, paired with 13.7 million energy evaluations and 5.5 million similarity annotations. It delivers continuous landscapes from non-native to near-native states, benchmarking framework for multi-conformation generators, and an interactive analysis platform.

|
Scooped by
mhryu@live.com
February 21, 10:57 AM
|
The choice between cell death (lysis) and viral dormancy (lysogeny) following bacteriophage infection serves as a founding paradigm for the emergence of cellular heterogeneity in a genetically uniform population. The determination of host fate arises through the stochastic transcription from multiple viral genomes present within each cell, but this activity remains hidden from empirical interrogation, which typically stops at the whole-cell level. Here we use parallel sequential fluorescence in situ hybridization (par-seqFISH), followed by spatial clustering of phage-encoded transcripts within each cell, to profile the transcriptional activity of individual phages during synchronized infection of E. coli by bacteriophage lambda. At the whole-cell level, transcription kinetics capture the developmental choice between lysis and lysogeny, and further demonstrate that viral replication is required for the emergence of diverging fate decisions. Zooming in to the single-phage level illuminates an individuality of viral activity during infection. We find that, while cells pursuing lysogeny display consensus activity of all inhabiting phages, lytic cells may contain phages that exhibit lysogenic activity. These findings support an earlier suggestion that consensus among coinfecting phages is required for cell dormancy. More broadly, our results highlight the need to identify how whole-cell behavior emerges from the activity of physically distinct copies of the same genetic circuit.

|
Scooped by
mhryu@live.com
February 20, 5:16 PM
|
Intracellular bacterial pathogens can cause high levels of morbidity and mortality in humans. Host immune responses that protect against these infections include pyroptosis, a form of lytic cell death caused by the insertion of large gasdermin (GSDM) pores into the host plasma membrane. Here we review recent advances in our understanding of how the five GSDM proteins, GSDMA–E, are activated by distinct signalling pathways. Pyroptosis can both eliminate intracellular niches and release cytosolic interleukin-1 family cytokines that further prime host immune responses against the invading pathogen. Because pyroptosis targets microbes, host-adapted intracellular pathogens have evolved strategies to efficiently subvert it. However, environmental pathogens fail to evade, making pyroptosis a potent barrier against infection. We summarize recent findings for the host-adapted bacterial pathogens Shigella flexneri, Salmonella enterica serovar Typhimurium and Mycobacterium tuberculosis, contrasted with the environmental bacteria Burkholderia thailandensis and Chromobacterium violaceum. This Review discusses recent advances in our understanding of how gasdermins are activated by distinct signalling pathways and summarizes recent findings from host-adapted bacterial pathogens contrasted with environmental bacteria.
|

|
Scooped by
mhryu@live.com
Today, 12:24 PM
|
Using cellulase to degrade cellulose for cellobiose production is an economically competitive pathway for the high-value utilization of lignocellulose. Cellobiohydrolases hydrolyze cellulose into cellobiose but suffer severe product inhibition, whereas endoglucanases (EGs) randomly cleave cellulose chains into oligosaccharides. Three heterologously expressed EGs from different sources (PcCel45A, BsCel5, and GtCel5A) were screened for their ability to degrade cellulose into cellobiose, and their hydrolysis processes were characterized. Using PcCel45A and BsCel5 to hydrolyze Avicel step-by-step at a protein ratio of 1:1 produced cellobiose with a purity of 90.99% and a yield of 1.2 g per 100 g of Avicel. Further studies revealed that PcCel45A and BsCel5 cannot degrade cellobiose, and their activities are not inhibited by the product, showing their advantage in cellobiose production. Additionally, adsorption kinetics and thermodynamic analyses indicated that PcCel45A and BsCel5 exhibited distinct adsorption mechanisms. This study provides a promising new strategy for the high-value utilization of lignocellulose.

|
Scooped by
mhryu@live.com
Today, 12:15 PM
|
High-yielding recombinant protein expression systems often face challenges due to the metabolic burden caused by the competition for cellular resources, resulting in reduced growth and, hence, limiting their industrial applicability. Furthermore, industrial recombinant protein production is also affected by the occurrence of oxygen gradients, which is a prevalent issue in large-scale bioreactors. These gradients create a heterogeneous environment in the bioreactor, which affects cell growth and metabolism, having severe consequences on the process performance. Both these factors alter cellular physiology and metabolism, thereby affecting recombinant protein yields. Understanding metabolic adaptations to these stress conditions is crucial for uncovering the underlying cellular mechanisms, which can direct further optimization of the recombinant strains. In this study, we aimed to explore the combined response of the central metabolism of E. coli to metabolic burden and microaerobic conditions. Two recombinant protein-producing E. coli BL21 strains carrying XylS/Pm vectors with low (A2-mCh) and medium plasmid copy numbers (A3-mCh), and producing mCherry protein, were studied by introducing oxygen limitation. Central metabolite pools were analyzed by three targeted LC–MS/MS methods, using the isotope dilution strategy for absolute quantification. Both recombinant strains exhibited different levels of metabolic burden, with the strain possessing a higher plasmid copy number showing more pronounced growth retardation and a stronger impact on metabolite pools. Both strains, however, showed a similar response to oxygen limitation, with significant adaptations in the central metabolite pools. The low plasmid copy number strain showed an increase in the concentration of lower glycolytic and tricarboxylic acid cycle metabolites, while the pools of upper glycolytic and pentose phosphate pathways and nucleoside phosphates were mostly unaffected. However, a more extreme response was seen in A3-mCh, where the majority of the metabolite pools were increased. Oxygen limitation caused lower metabolic activity, but the energy charge and redox balance were maintained, and no negative effect was observed on mCherry production rates. This study provides insights into metabolic adaptations in E. coli BL21 recombinant strains, having quite robust mechanisms to maintain intracellular metabolic homeostasis during both internal and external perturbations.

|
Scooped by
mhryu@live.com
Today, 11:48 AM
|
Anthropogenic nitrogen enrichment is widely expected to suppress symbiotic nitrogen fixation in terrestrial ecosystems. Nevertheless, observed symbiotic nitrogen fixation responses remain incompletely explained by exogenous nitrogen inputs, climate, and edaphic factors. In this meta-analysis, we integrate 908 globally distributed field measurements to identify the key predictors that improve simulation of symbiotic nitrogen fixation responses to nitrogen enrichment. On average, symbiotic nitrogen fixation declines by 33.0% upon nitrogen enrichment, with the reduction being more pronounced in non-croplands than croplands. Models considering only environmental factors overestimate symbiotic nitrogen fixation decline relative to observations. The better performance of plant traits like plant growth and biomass allocation (shoot:root ratio) partially buffer symbiotic nitrogen fixation suppression under nitrogen enrichment. Integrating both environmental factors and plant performance traits improves predictive accuracy of symbiotic nitrogen fixation responses by 42.7% and brings the simulated symbiotic nitrogen fixation reductions into close agreement with observations. The alterations in plant performance traits are thus critical for explaining variability in terrestrial symbiotic nitrogen fixation responses, and incorporating plant trait dynamics in Earth System Models can quantitatively partition the compensatory symbiotic nitrogen fixation supported by nitrogen-fixing plant growth from the direct negative impact of nitrogen inputs. Anthropogenic nitrogen enrichment reduces symbiotic nitrogen fixation, but environmental factors alone cannot explain this response. This meta-analysis shows that integrating both environmental factors and plant performance traits improves the predictive accuracy of symbiotic nitrogen fixation responses by 42.7%.

|
Scooped by
mhryu@live.com
February 21, 2:11 PM
|
Natural products are vital sources of medicines, and their analogues hold great promise as novel therapeutics to combat disease more effectively, including circumventing antibiotic resistance. Engineering biosynthetic pathways in microbial hosts enables efficient, cost-effective production of non-natural analogues — compounds structurally related to natural products but not necessarily derived through direct chemical modification. In this review, we highlight recent work on the in vivo production of novel natural product analogues, with an emphasis on combinatorial and precursor-directed biosynthesis, enzyme engineering, and retrobiosynthesis. We anticipate that further developments in artificial intelligence, particularly the use of machine learning models to understand enzymatic transformations and predict novel reactions, will significantly accelerate this field and drive forward its importance in drug discovery and related research.

|
Scooped by
mhryu@live.com
February 21, 1:33 PM
|
Antibiotic resistance is a growing global health threat. Although antibiotic activity is well studied in homogeneous liquid cultures, many infections are caused by spatially structured multicellular populations where consumption of scarce nutrients establishes strong spatial variations in their abundance. These nutrient variations have long been hypothesized to help bacterial populations tolerate antibiotics, since liquid culture studies link antibiotic tolerance to metabolic activity, and thus, local nutrient availability. Here, we test this hypothesis by visualizing cell death in structured E. coli populations exposed to select nutrients and antibiotics. We find that nutrient availability acts as a bottleneck to antibiotic killing, causing death to propagate through the population as a traveling front. By integrating our measurements with biophysical theory and simulations, we establish quantitative principles that explain how collective nutrient consumption can limit the progression of this “death front,” protecting a population from a nominally deadly antibiotic dose. While increasing nutrient supply can overcome this bottleneck, in some cases, excess nutrient unexpectedly promotes the regrowth of resistant cells. Altogether, this work provides a key step toward predicting and controlling antibiotic treatment of spatially structured bacterial populations, yielding biophysical insights into collective behavior and guiding strategies for effective antibiotic stewardship. In this work, authors provide evidence that bacteria in spatially structured populations protect each other from antibiotics through collective nutrient consumption, creating ‘death fronts’ that sweep through the colony– explaining why infections often survive treatments that work in lab tests.

|
Scooped by
mhryu@live.com
February 21, 1:08 PM
|
With antimicrobial resistance emerging as a top global health concern, new approaches to prevent and treat infections are urgently needed. Antimicrobial peptides are recognized as a promising alternative to conventional antibiotics. Considering the costly, labor-intensive, and time-consuming nature of wet lab screening, in recent years, proteomics research has increasingly relied on in silico methods to predict physicochemical, structural, and biological properties of candidate peptides. Here, we present the AMPSeek protocol to enable scientists to screen peptides and predict their potential antimicrobial activity, toxicity, and associated three-dimensional structure with a single command. The AMPSeek framework integrates AMPlify, tAMPer, and LocalColabFold, scales efficiently with increasing sample size, and produces an interactive HyperText Markup Language report to facilitate result interpretation. We provide step-by-step instructions for installing and running AMPSeek on a ten-peptide test set, along with two additional datasets (n = 20 and n = 100; mean peptide length ≈121.3) to demonstrate AMPSeek's near-linear scalability.

|
Scooped by
mhryu@live.com
February 21, 1:01 PM
|
The transition to low-carbon energy systems requires scalable and energy-efficient routes for producing liquid biofuels that are compatible with existing fuel infrastructures. This review focuses on bio-oil production from phototrophic microorganisms, highlighting their high biomass productivity, rapid growth, and inherent capacity for carbon dioxide fixation as key advantages over conventional biofuel feedstocks. Recent progress in thermochemical conversion technologies, particularly hydrothermal liquefaction (HTL) and fast pyrolysis, is critically assessed with respect to their suitability for wet and dry algal biomass, respectively. HTL enables direct processing of high-moisture biomass while avoiding energy-intensive drying, whereas fast pyrolysis offers high bio-oil yields from lipid-rich feedstocks. In parallel, catalytic upgrading strategies, including hydrodeoxygenation and related hydroprocessing routes, are discussed as essential steps for improving bio-oil stability, heating value, and fuel compatibility. Beyond conversion technologies, innovative biological and biotechnological strategies, such as strain optimization, stress induction, co-cultivation, and synthetic biology approaches, are examined for their role in tailoring biomass composition and enhancing bio-oil precursors. The integration of microalgal cultivation with wastewater utilization is briefly considered as a supporting strategy to reduce production costs and improve overall sustainability. Overall, this review emphasizes that the effective coupling of advanced thermochemical conversion with targeted biological optimization represents the most promising pathway for scalable bio-oil production from phototrophic microorganisms, positioning algal bio-oil as a viable contributor to future low-carbon energy systems.

|
Scooped by
mhryu@live.com
February 21, 12:50 PM
|
In Azotobacter vinelandii, the sigma factor RpoS is maintained at low levels in exponentially growing cells due to degradation mediated by the chaperone–protease complex ClpXP, while high levels are observed in the stationary phase. This study showed that degradation of RpoS by ClpXP is under the control of the Gac-Rsm signal transduction system, in which GacA, the transcriptional activator of the two-component system GacS/GacA, activates transcription of the small RNAs RsmZ1 and RsmZ2. These RNAs bind to the translational repressor protein RsmA to counteract its repressor activity on its target mRNAs. We found that in stationary-phase cells, compared with the WT, a gacA mutant exhibited lower RpoS levels due to reduced stability, while levels of the clpP and clpX mRNAs were higher. Furthermore, inactivation of the clpP or clpX genes in the gacA mutant restored the stability of RpoS, suggesting that the observed RpoS instability is due to degradation by ClpXP. We also showed that inactivation of rsmA in either the WT or the gacA mutant resulted in RpoS levels higher than in the WT in both stationary and exponential phases, while clpP and clpX transcript levels were significantly reduced. Taken together, these data reveal that in A. vinelandii, the GacA-Rsm system controls RpoS stability through RsmA, which acts as a positive regulator of ClpXP expression.

|
Scooped by
mhryu@live.com
February 21, 12:16 PM
|
Protein structure prediction using deep learning has revolutionized protein design. Yet, our understanding of protein function remains a key limitation for designing novel proteins that perform complex biological tasks. Here, we adopt a massively-parallel, function-first approach to rationally design synthetic proteins. Using genome-scale CRISPR activation, we overexpress ~19,000 human proteins and measure their impact on precise gene editing. We identify over 800 native proteins that promote homology-directed repair. Using top candidates, we then design synthetic genome editors, Targeted Repair fUsion Editors (TruEditors), by fusing full-length proteins or smaller core domains to the Cas9 nuclease. We develop 12 unique TruEditors that improve precise gene editing in diverse cell types and at genomic loci where existing methods for precise gene editing fail. Using affinity proteomics, we show that these synthetic proteins work by coordinating with endogenous DNA repair complexes. The delivery of TruEditors via mRNA more than doubles the rate of chimeric antigen receptor (CAR) insertion into the TRAC locus of primary human T cells, enhancing CAR T cell-directed tumor cell killing, and improves precise editing in human pluripotent stem cells more than three-fold. Overall, our study demonstrates that genome-wide protein overexpression screens can guide the rational design of synthetic proteins for specific biological tasks.

|
Scooped by
mhryu@live.com
February 21, 11:35 AM
|
Models of protein evolution are foundational to biology, underpinning essential techniques such as phylogenetic tree inference, ancestral sequence reconstruction, multiple sequence alignment, variant effect prediction, and protein design. Historically, for computational tractability, these models have relied on the simplifying -- but biologically unrealistic -- assumption that sites in a given protein evolve independently of each other. A crucial test of any evolutionary model is its ability to simulate realistic evolutionary trajectories, but the independent-sites assumption leads to simulations that poorly reflect the complexity of natural protein evolution. Here we introduce PEINT (Protein Evolution IN Time), a flexible and generalizable deep learning framework for modeling how the entire protein sequence evolves over time while incorporating complex interactions between sites. This framework enables learning realistic patterns of constrained evolutionary transitions directly from millions of protein sequences spanning diverse fold families. Furthermore, unlike classical models that require pre-aligned sequences, PEINT learns indel dynamics directly from raw, unaligned sequences, thereby eliminating potential biases from alignment errors that can lead to incorrect inference of evolutionary patterns. By capturing higher-order epistatic interactions and modeling insertion-deletion processes that classical models typically ignore, PEINT accurately reproduces key signatures of natural evolution, including conservation patterns and family-specific dynamics. When simulating evolution along phylogenetic trees, PEINT generates highly novel sequences that preserve protein function, which we validate through experimental characterization of simulated carbonic anhydrase variants that retain enzymatic activity. PEINT thus enables realistic simulation of protein evolution that explores new sequence space while respecting structural and functional constraints. This evolution-informed generative modeling framework offers a powerful new tool for advancing both phylogenetic inference and protein engineering.

|
Scooped by
mhryu@live.com
February 21, 11:13 AM
|
Polyploidy is a ubiquitous biological phenomenon that promotes genetic innovation and adaptation. In the budding yeast Saccharomyces cerevisiae, polyploid strains are widespread in natural, industrial, and clinical niches, yet their origin is unclear. Here, we identify a novel mechanism of stepwise polyploidization that involves a Sporulate-Endoreplicate-Mate (SEM) sequence. We show that spore endoreplication during germination doubles the genome while preserving mating competence, enabling incremental increases in ploidy through mating with a non-endoreplicated spore from the same ascus. We experimentally demonstrate the transition from diploidy to triploidy and from triploidy to tetraploidy. Stepwise polyploidization thus positions triploids as central intermediates in ploidy evolution rather than evolutionary dead-ends. We further show that spores from intact asci can spontaneously undergo one or two successive SEM cycles and generate novel triploid and tetraploid strains without genetic manipulation. Natural polyploids harbor specific genomic features that are consistent with the SEM mechanism, including the prevalence of triploidy, extensive aneuploidy, pervasive heterozygosity, and a strong association with heterothallism. Together, our findings establish stepwise polyploidization through iterative SEM cycles as the predominant natural route to polyploidy in S. cerevisiae.

|
Scooped by
mhryu@live.com
February 20, 5:21 PM
|
Intestinal motility is a function of the enteric nervous system involving secretion of the excitatory neurotransmitter acetylcholine (ACh). Although gut commensal bacteria are key regulators of intestinal physiology, the molecular mechanisms underlying microbial influence on intestinal peristalsis and constipation remain unclear. Here we report a link between microbial nitrogen metabolism and intestinal motility regulation via ammonia production. We observed compensatory elevation of intestinal ammonia levels and urease activity in mouse models of intestinal dysmotility, induced by ACh deficiency, and in patients with constipation. Ammonia supplementation or intervention in mice with the urease-positive Lysinibacillus fusiformis isolated from patient stool, or engineered urease-expressing bacteria, effectively restored colonic ACh levels. In vitro, ammonia upregulated the expression of voltage-gated calcium channels on enteric neurons, driving Ca2+ influx to potentiate ACh secretion. Our study reveals a microbial compensatory mechanism that responds to fluctuating ACh levels in the intestine and provides microbial targets for intestinal motility disorders. Intestinal motility defects caused by lower acetylcholine levels in the gut are compensated by bacteria that generate ammonia from urea which facilitates the release of acetylcholine via upregulation of voltage-gated channels on enteric neurons.
|
 Your new post is loading...
Your new post is loading...
































mode of regulation