 Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 1:11 AM
|
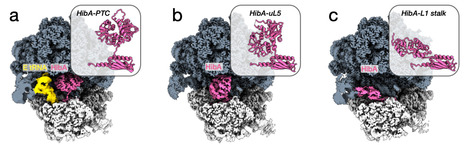
Ribosome hibernation preserves translation machinery during stress, yet its mechanisms in Archaea remain poorly defined. Using cryo-EM analysis, we studied hibernation pathways in Pyrococcus abyssi stressed cells. We identified HibA, a previously unrecognized family of hibernation factors widespread in Archaea. HibA consists of a bacterial-like HPF/RaiA domain fused to a Cystathionine Beta Synthase module. Unexpectedly, HibA binds to the ribosome in three different conformations, occupying the A, P and E sites of tRNAs, as well as that of mRNA, enhancing its ability to protect the ribosome from degradation. Idle ribosomes also frequently accumulate the archaeal homolog of eukaryotic ribosome maturation protein SBDS (aSBDS), suggesting that stressed archaeal cells may engage parallel hibernation routes in which aSBDS can complement HibA. Deletion of hibA in Thermococcus barophilus delays recovery from stationary phase and reduces 70S ribosome pools, establishing its role in ribosome preservation. Taxonomic profiling shows that many archaeal lineages encode distinct repertoires of ribosome-associated protection factors, underscoring the modular and multi-layered nature of archaeal hibernation systems. In addition, a comprehensive phylogenetic analysis highlights the evolutionary relationships between prevalent ribosome hibernation factors across Bacteria and Archaea. Ribosome hibernation preserves translation machinery during stress. Here, the authors identify a ribosome hibernation factor that is widespread in archaea and combines bacterial HPF/RaiA-like and cystathionine beta-synthase domains.

|
Scooped by
mhryu@live.com
Today, 12:55 AM
|
Monitoring molecular activities within single live cells is vital for understanding cellular differentiation, senescence, heterogeneity, and disease progression. However, conventional single-cell analyses often rely on micromanipulation or extraction followed by downstream measurements, which cannot capture in situ real-time dynamics. Fluorescent labeling and electrochemical methods provide temporal resolution but face limitations in labeling, substrate scope, and multiplexing. Here, we present a nanopore probe that enables real-time, multiplexed monitoring of intracellular activities in single live cells. The device integrates an aluminum oxide nanostraw membrane for molecular extraction and a glass nanopore membrane for single-channel electrical detection. Using a hippocampal neuron model of ischemia-hypoxia, we simultaneously tracked dynamic changes in intracellular glutamate, ascorbic acid, and adenosine triphosphate—three key molecules involved in oxygen-glucose deprivation-induced neuronal edema. Our findings establish this nanopore probe as a powerful platform for real-time, label-free molecular profiling at the single-cell level, opening opportunities for studying disease mechanisms and therapeutic responses.

|
Scooped by
mhryu@live.com
Today, 12:43 AM
|
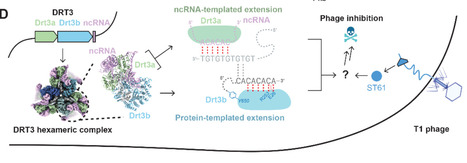
Defense-associated reverse transcriptases (DRTs) are widespread bacterial antiphage systems that use unconventional mechanisms of polynucleotide synthesis. We show that DRT3, which comprises two distinct RTs (Drt3a and Drt3b) and a noncoding RNA (ncRNA), synthesizes alternating poly(GT/AC) double-stranded DNA. Cryo–electron microscopy structures at 2.6 Å resolution reveal a D3-symmetric 6:6:6 complex of Drt3a, Drt3b, and ncRNA. Drt3a produces the poly(GT) strand using a conserved ACACAC template within the ncRNA. Notably, Drt3b synthesizes a complementary, protein-primed poly(AC) strand in the complete absence of a nucleic acid template, using conserved active site residues specific to Drt3b to enforce precise base alternation. These findings expand the functional landscape of nucleic acid polymerases, revealing a protein-templated mechanism for sequence-specific DNA synthesis.

|
Scooped by
mhryu@live.com
April 26, 12:41 PM
|
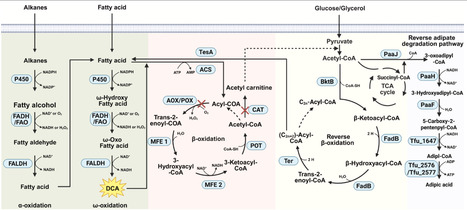
Medium- and long-chain dicarboxylic acids (M/LCDAs) are key monomers for the synthesis of nylons and high-performance engineering plastics. Compared to traditional chemical methods, microbial synthesis offers advantages such as environmental friendliness and high regioselectivity. However, its industrial application remains limited by bottlenecks, including low mass transfer efficiency on hydrophobic substrates, instability of key oxidase systems, and cellular metabolic imbalances. This review summarizes recent strategies leveraging enzyme engineering, systems metabolic engineering, and diverse synthetic biology approaches to overcome current limitations in the biosynthesis of M/LCDAs. We specifically highlight mechanisms for enhancing the transmembrane transport of hydrophobic substrates and the mining of novel transporters. Furthermore, we elaborate on protein engineering efforts targeting key enzymes (e.g. cytochrome P450s), covering rational design, fusion expression, and novel dimerization techniques. At the systems level, we discuss metabolic network regulation achieved through the construction of the reverse β-oxidation cycle (r-BOX) and the reprogramming of cofactor regeneration and energy metabolism. Finally, future perspectives on integrating AI-aided design and waste valorization are proposed to provide theoretical guidance for the efficient and sustainable biomanufacturing of M/LCDAs.

|
Scooped by
mhryu@live.com
April 26, 11:22 AM
|
Nitrate (NO3−) serves as a pivotal molecule with dual functions in nutrient supply and signaling during plant growth and development. Precise monitoring of its spatiotemporal dynamics in planta is therefore essential for dissecting the regulatory mechanisms underlying plant nitrogen metabolism. However, conventional nitrate detection methods suffer from inherent limitations, including destructive sampling, insufficient spatiotemporal resolution, and an inability to achieve real-time whole-plant monitoring. Here, we report a genetically encoded nitrate biosensor, designated NitNRCL1, constructed using a split firefly luciferase complementation system. Functional validation in both prokaryotic and eukaryotic systems demonstrates that NitNRCL1 responds to changes in nitrate availability and generates stable chemiluminescent signals in bacteria and diverse plant species. Importantly, NitNRCL1 enables non-invasive, real-time, and whole-plant monitoring of nitrate levels in living plants. Using NitNRCL1, we successfully imaged the spatiotemporal dynamics of nitrate signaling in Arabidopsis thaliana. Collectively, our findings establish NitNRCL1 as a robust and novel tool for investigating nitrate transport, signaling, and metabolic pathways in plants. This biosensor advances our mechanistic understanding of plant nitrate biology and provides a technical foundation for breeding nitrogen-use-efficient crops and developing precision fertilization strategies.

|
Scooped by
mhryu@live.com
April 26, 11:16 AM
|
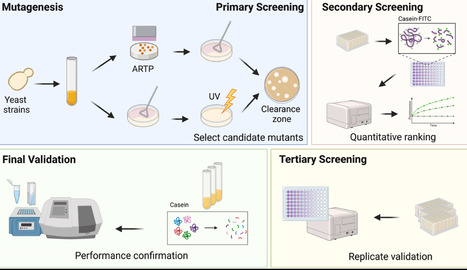
Efficient secretion of heterologous proteases in S. cerevisiae remains a major bottleneck due to host cytotoxicity and secretory stress. In this study, we developed a yeast chassis strain through three rounds of iterative evolution, combining ultraviolet (UV) and atmospheric and room temperature plasma (ARTP) mutagenesis with a fluorescence-based screening pipeline. The final strain, MA10Ah121, exhibited protease production nearly three times that of its parental strain LB184M and maintained genetic stability across passages. Importantly, MA10Ah121 supported enhanced expression of two structurally distinct collagenases, ColG (Class I) and ColH (Class II), demonstrating its broad utility for recombinant protease production. Whole-genome sequencing revealed substantial chromosomal remodeling, including reversion of a 151 kb duplication enriched in genes related to energy metabolism and protein synthesis, along with widespread structural variations. These changes reflect a genome-wide adaptation that rebalances biosynthetic capacity and stress tolerance. This work provides a robust and versatile S. cerevisiae chassis for the biosynthesis of proteases and other challenging recombinant enzymes in industrial biotechnology applications.

|
Scooped by
mhryu@live.com
April 25, 11:45 PM
|
Advances in live-cell fluorescence microscopy have enabled us to visualize single molecules (such as mRNAs and nascent proteins) in real time with high spatiotemporal resolution. However, these experiments generate large datasets that require complex computational processing pipelines to derive meaningful and quantitative information, which is a technical barrier for many researchers. Here, we introduce MicroLive, an open-source Python-based application for quantifying live-cell microscopy images. MicroLive provides an interactive Graphical User Interface (GUI) to perform key tasks, including cell segmentation, photobleaching correction, single-particle detection/tracking, spot intensity quantification, inter-channel colocalization, and time-series correlation analysis. As a ground-truth testing dataset, we used synthetic live-cell imaging data generated with the rSNAPed toolkit, demonstrating accurate extraction of biologically relevant parameters. Microscopy images of U-2 OS cells expressing a gene construct smHA-KDM5B-BoxB-MS2 were used to demonstrate the use of this software.

|
Scooped by
mhryu@live.com
April 25, 11:38 PM
|
Higher-order genetic interactions have profound implications for understanding the molecular mechanisms of phenotypic variation, yet they remain poorly characterized. Most studies focus on pairwise interactions because high-throughput screens over the vast combinatorial space are challenging. Here, we develop Dango, a computational method based on a self-attention hypergraph neural network, to predict higher-order genetic interactions among groups of genes. As a proof of concept, we provide predictions for over 400 million trigenic interactions in the yeast S. cerevisiae, greatly expanding their quantitative landscape. Dango accurately predicts trigenic interactions and reveals biological functions related to cell growth. We further incorporate protein embeddings and uncertainty estimation to improve biological relevance and interpretability. Moreover, predicted interactions serve as genetic markers for growth responses across diverse conditions. Together, Dango enables a more complete map of complex genetic interactions that shape phenotypic diversity.

|
Scooped by
mhryu@live.com
April 25, 11:27 PM
|
Freshwater lakes are globally significant sources of potent greenhouse gases (GHGs), but how their GHGs emissions respond to changing nutrient levels remains unclear. Here, we demonstrated that nitrous oxide (N2O) production pathways in lake sediments are tightly linked to trophic state, whereas methane (CH4) production appears to be multifactorial Through global metagenomics and controlled batch experiments. In eutrophic sediments, N2O is efficiently removed through complete denitrification, with nitrification serving as the main production pathway, whereas oligotrophic sediments produce N2O primarily via incomplete denitrification. By simulating nutrient transitions using a cross-inoculation experiment, we further revealed that lake sediments systematically shift between these N2O production pathways as their trophic state changes, from denitrification-driven to nitrification-dominated during eutrophication, with the inverse pattern during oligotrophication. Consequently, N2O emissions can be effectively mitigated by inhibiting nitrification in eutrophic lakes and restricting incomplete denitrification in oligotrophic ones. Our findings establish trophic status as a key driver of N2O production sources in lake sediments. Trophic status is a critical regulator of N2O dynamics in freshwater lake sediments, with implications for predicting GHG fluxes under global change scenarios.

|
Scooped by
mhryu@live.com
April 25, 11:15 PM
|
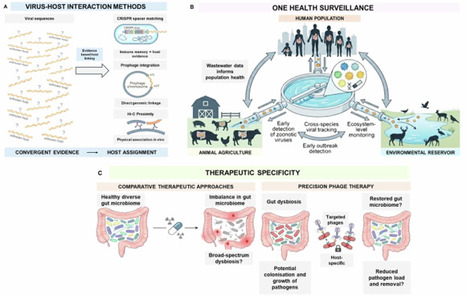
The Gut microbiome-virome dynamics and interactions Collection highlights gut viruses, mainly bacteriophages, as determinants of microbial community structure and host-relevant functions across human, animal, and environmental systems. The Collection welcomes studies that quantify virus-microbe interactions, evidence-based linking viruses to microbial hosts, characterise infection dynamics, and connect them to ecological or clinical outcomes. It prioritises methodological rigour and translational relevance in diagnostics, surveillance, and phage-based interventions.

|
Scooped by
mhryu@live.com
April 25, 4:03 PM
|
The gut-liver-brain axis is central to metabolic and neurological homeostasis and is mediated by host- and microbiota-derived metabolites. Disruptions in this axis contribute to complex disorders, underscoring the need for targeted, multi-metabolite interventions. Here, we engineered commensal Lactobacillus plantarum WCFS1 strains to specifically modulate metabolites dysregulated in hepatic encephalopathy (HE), a disorder driven by hyperammonemia and amino acid imbalance. One strain couples ammonia assimilation with branched-chain amino acid (BCAA) biosynthesis, whereas the other enhances L-glutamine utilization to suppress ammonia generation. In two preclinical HE models, these strains reduced systemic ammonia by up to 10-fold, restored BCAA and L-glutamine balance, and improved anxiety-like and cognitive behaviors. Notably, they outperformed rifaximin, a clinically used HE therapy, while preserving gut microbiota diversity. These findings establish engineered commensals as a modular, responsive platform for multi-metabolite modulation of host-microbiota metabolism, offering a programmable strategy to restore metabolic homeostasis in disorders of the gut-liver-brain axis.

|
Scooped by
mhryu@live.com
April 25, 3:50 PM
|
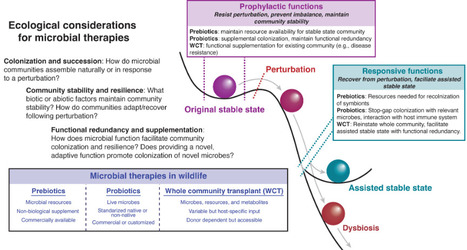
Microbial ecology is increasingly incorporated into human and animal medicine via the study and purposeful manipulation of host-associated microbiomes. Microbial therapies—treatments with the aim of beneficially modulating microbiomes—are a burgeoning area of research and industry. These microbial therapies include prebiotic dietary items, live probiotics, and whole microbiota transplants (e.g., fecal microbiota transplants). Although microbial therapies for humans and domestic animals are now widely produced for commercial use and application, evidence supporting the efficacy of commercial microbial therapies is mixed. We suggest that microbial therapies are most effective when paired with concepts from ecology and rigorous empirical research. This is particularly relevant for the development and use of microbial therapies in wildlife animal species, in which we see large-scale variation in microbial communities across hosts of varying ecologies. Identifying and developing microbial therapies that can simultaneously be accessible and effective in a variety of hosts poses a novel challenge for microbial ecologists, animal scientists, and human and animal medical professionals. In addition to pre- and probiotics, we suggest that whole microbiota transplants provide a method of microbial supplementation that may better align with species-specific microbial ecology. Moving forward, emerging methods used in human medicine such as machine learning, network analysis, and microbiome engineering using high-throughput culturomics will likely be key to identifying and applying functionally relevant (e.g., disease suppressive) microbial taxa for wildlife therapies.

|
Scooped by
mhryu@live.com
April 25, 3:42 PM
|
Gene expression analysis has evolved substantially over the past 25 years, from early transcript surveys using expressed sequence tags and microarrays to RNA sequencing, and more recently to single-cell and spatial transcriptomics. These successive waves have expanded measurement scale and resolution, enabling systematic discovery of transcriptional programmes, inference of gene regulatory networks, and increasingly direct links between transcriptomic insight and therapeutic strategies that modulate gene expression. In this Perspective, we synthesize major methodological milestones with bibliometric trends in leading bioinformatics journals to describe four revolutions that redefined gene expression analysis. We also map widely used computational tools onto a common timeline by analysing 70 78 831 open-access full-text articles, illustrating how enduring statistical frameworks coexist with rapidly growing end-to-end analysis ecosystems. We highlight current challenges and emerging directions in core bioinformatics approaches for gene expression analysis. Looking ahead, we argue that the next era will be defined less by generating new datasets and more by organizing, searching, and reusing transcriptomic and multimodal information at scale. We propose three future directions: consortium-scale searchable transcriptomic knowledgebases, foundation models for gene expression analysis, and programmable regulatory design for engineered control of gene expression. The landscape of gene expression analysis is shifting from descriptive measurement towards queryable, predictive, and programmable gene expression biology.
|

|
Scooped by
mhryu@live.com
Today, 1:01 AM
|
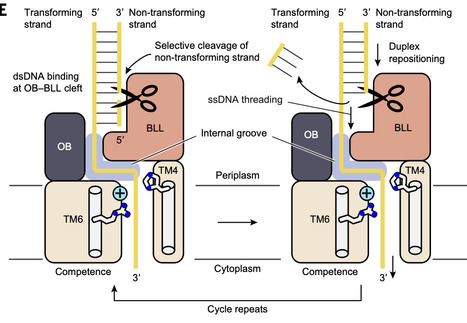
Natural transformation is one of the major pathways of horizontal gene transfer in bacteria, enabling the acquisition of extracellular DNA and its integration into the host genome. ComEC is a membrane protein responsible for DNA translocation in this process, yet its precise function and structure have remained elusive. Here, we report cryo–electron microscopy structures of ComEC in DNA-free, single-stranded DNA (ssDNA)–bound, and double-stranded DNA (dsDNA)–bound forms, together with biochemical analyses. These structures reveal that ComEC cleaves one strand of dsDNA at its extracellular domain and guides the remaining strand into a positively charged pore formed within the membrane domain. These findings provide a structural basis for the long-hypothesized roles of ComEC in both DNA processing and translocation across the inner membrane during natural transformation.

|
Scooped by
mhryu@live.com
Today, 12:51 AM
|
Bacteria often survive viral attack and environmental stress by sharing genes that enhance their defenses. The cholera pathogen Vibrio cholerae carries a sedentary chromosomal integron (SCI), a genetic element containing hundreds of mostly promoterless gene cassettes, about 10% of which encode antiviral systems. Cassettes are thought to reshuffle under stress to the favorable first array position, yet the SCI in pandemic V. cholerae has remained static for more than 60 years. In this study, we show that SCI diversification efficiently occurs by horizontal transfer linked to the genus’s aquatic lifestyle: DNA released from lysed cells is taken up by naturally competent vibrios and integrated into the first position of the SCI array, the primary site of strong expression, where it confers resistance to phage and potentially other threats.

|
Scooped by
mhryu@live.com
April 26, 11:40 PM
|
This study investigates the relationship between the gut microbiota and specific diseases. Data was collected from the Human Gut Microbiome Atlas, which examines regional variations across 20 countries on five continents, categorizing microbial species by taxonomy, from genus to species. The Atlas provides color-coded phylum classifications, numerical species counts within the same genus, and an analysis of dysbiosis-related associations with 23 diseases, as well as region-enriched species. The data stratified samples into distinct categories such as westernized, non-westernized, cancerous, and non-cancerous. The findings demonstrate that tree-based ensemble methods, such as Bagging and Boosting prediction methods, achieved the highest accuracies across all categories due to their robustness in handling the complex, high-dimensional data. The XGBoost model yielded the strongest predictive performance, achieving 91% accuracy for westernized cancer-associated samples, 84% accuracy for non-westernized cancer-associated samples, 92% accuracy for westernized samples, and 78% for non-westernized samples. Additionally, advanced topological data analysis was used to assess the global structure and underlying patterns within the dataset.

|
Scooped by
mhryu@live.com
April 26, 12:39 PM
|
The site-specific incorporation of noncanonical amino acids (ncAAs) has revolutionized protein research. However, synthesizing multifunctional biopolymers demands moving beyond single-type ncAA incorporation to simultaneously encoding multiple distinct ncAAs. This review summarizes transformative progress in the genetic encoding of multiple distinct ncAAs, including the development of mutually orthogonal translation systems and the expansion of codon repertoires. We highlight recent breakthroughs that have achieved the incorporation of multiple distinct ncAAs. Notably, integrating rare codon recoding with stop codon suppression in mammalian systems has recently enabled the genetic encoding of five distinct ncAAs. These innovations establish a robust platform for advanced biosensors, next-generation therapeutics, and synthetic biology with expanded chemical repertoires.

|
Scooped by
mhryu@live.com
April 26, 11:19 AM
|
Recent advances in Machine Learning have transformed antibody development through in silico models, accelerating therapeutic candidate identification. However, challenges persist: rapid adaptation of property predictors to laboratory-specific assays with incomplete datasets; batch effects introducing systematic bias; assay costs necessitating efficient unseen property prediction. We introduce a novel multimodal architecture featuring specialized tokenization and embedding projection that integrates text and protein language models (pLM) and a learning strategy to enable context-conditioned multi-property prediction without learning shortcuts. Our framework enables prompting without dictionary merging across modalities, creating a compact model capable of context-conditioned learning for multi-property prediction. The orchestrating model avoids pLM-to-text projection while enabling inference-time adaptation without retraining. Using 876,898 antibody heavy chain sequences with batch effect simulation, our architecture achieved Spearman’s ρ > 0.8 across multiple developability properties, significantly outperforming fine-tuned multimodal LLMs and showed the ability to leverage correlation between properties for prediction. This approach has the potential to address critical antibody development challenges.

|
Scooped by
mhryu@live.com
April 25, 11:56 PM
|
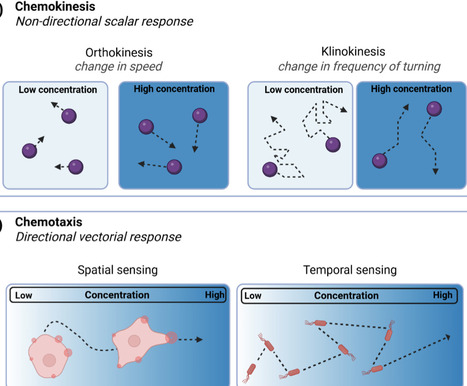
Chemotaxis is the directional motion of objects in response to chemical gradients, a process that drives navigation in micro/nanomotors, enabling a bio-inspired transition toward autonomous direction control. However, current studies lack consistent mechanistic justification, definition, and standardized experimental validation. This review summarizes key physical principles governing active chemotaxis, surveys experimental strategies for generating chemical gradients and quantifying responses, and examines how individual chemotactic mechanisms and long-range, anisotropic chemical interactions give rise to emergent collective behaviors, while outlining prospective applications. Together, these efforts establish a principled engineering framework for advancing chemotaxis as a robust functional navigation modality in synthetic micro/nanomotors. This Review Article examines artificial chemotaxis in micro/nanomotors, detailing mechanisms, experimental strategies, and collective behaviors. It highlights the need for standardized validation and explores applications in biomedical and industrial fields, emphasizing the challenges and potential of synthetic chemotaxis.

|
Scooped by
mhryu@live.com
April 25, 11:43 PM
|
Microbiomes perform critical functions for their hosts, and understanding microbiome variation is important for both basic and applied science. However, host traits alone cannot explain the entirety of microbiome variation, because, alongside host traits, microbiomes are shaped by multiple ecological processes. Researchers have thus turned to theories of island biology, conceptualising animal hosts as islands and animal microbiomes as metacommunities that assemble within and disperse between host islands. To develop realistic models, this host-as-island metaphor must be examined by explicitly comparing geological and host islands. Here, we critically examine the host-as-island metaphor by evaluating how microbiome variation is shaped by the four metacommunity processes that explain biodiversity on geological islands: local interspecies interactions, local selection, dispersal, and stochasticity. Key differences between host islands and geological islands include the complexity of microbiome transmission networks arising from host mobility and sociality and the capacity of hosts to evolve to control their microbiomes. We conclude with discussions of how eco-evolutionary dynamics differ between geological islands and host islands, and the reciprocal relevance of island biology and microbiome science.

|
Scooped by
mhryu@live.com
April 25, 11:29 PM
|
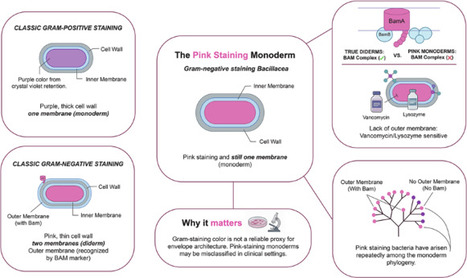
Gram staining has guided microbiology for over a century by coloring cells purple or pink, a read-out thought to distinguish monoderms (single membrane, thick peptidoglycan) from diderms (inner and outer membranes with thin wall). Here we show this rule fails repeatedly across Bacillaceae lineages historically deemed “Gram-positive”. By combining light and transmission-electron microscopy, antibiotic-sensitivity assays and comparative genomics across 57 strains, we identify “Gram-negative-staining monoderms” lacking outer membrane yet retaining thick peptidoglycan walls. These bacteria lack lipopolysaccharide- and β-barrel assembly-genes and remain highly susceptible to vancomycin and lysozyme (agents normally excluded by diderm envelopes), demonstrating functional monoderm status. Surprisingly, teichoic-acid biosynthetic pathways are patchily distributed and do not predict staining behavior. This discovery calls into question the textbook purple-or-pink dichotomy, decoupling stain color from membrane architecture. Clinically, misidentifying pink-staining Bacillaceae (including emerging pathogens such as Bacillus infantis) risks inappropriate therapy, whereas genome-guided diagnostics enable precise antibiotic stewardship. Genomic and microscopy evidence shows that the Bacillaceae, assumed to only stain Gram-positive, actually have some lineages staining Gram-negative, yet unexpectedly maintaining a thick wall and no outer membrane, challenging Gram staining assumptions.

|
Scooped by
mhryu@live.com
April 25, 11:20 PM
|
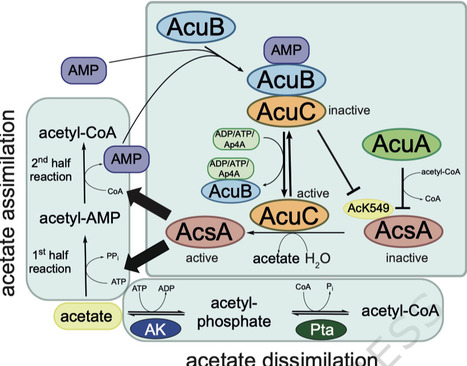
Bacteria adjust their metabolism to the cellular energy state. AMP-forming acetyl-CoA-synthetase AcsA generates acetyl-CoA from acetate, ATP and CoA. In Bacilli, including Bacillus subtilis and Geobacillus stearothermophilus, AcsA is reversely transcribed upstream of the acu-operon encoding for the proteins AcuA, AcuB and AcuC. Lysine-acetyltransferase AcuA uses acetyl-CoA to acetylate and inactivate AcsA, while AcuC re-activates AcsA activity by deacetylation. How the counteracting activities of AcuA and AcuC are regulated is not understood. Here, we close this gap of knowledge and perform a structure-function analyzes on AcuB. These reveal AcuB forming a scissor-shaped dimer with each monomer consisting of an N-terminal Bateman domain binding to adenine nucleotides and a C-terminal ACT domain. Structural and biochemical studies as well as molecular dynamics simulations support that AMP bound AcuB binds and inhibits AcuC. Our data describe another layer of regulation of AcsA activity in Firmicutes coordinating acetate assimilation and dissimilation by the energy sensor AcuB. Bacteria adjust their metabolism to the cellular energy state. Here, authors identify a layer of regulation of AMP-forming acetyl-CoA synthetase AcsA activity by AcuB acting as energy sensor inhibiting the AcsA deacetylase AcuC in presence of AMP.

|
Scooped by
mhryu@live.com
April 25, 11:15 PM
|
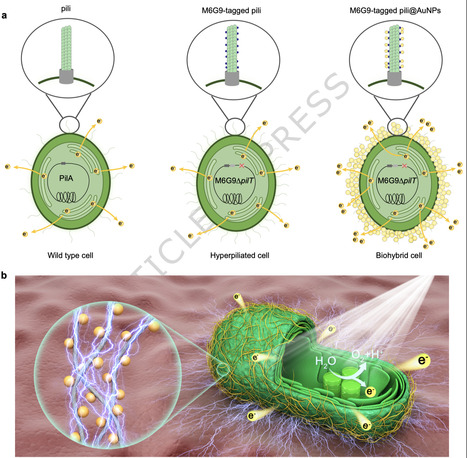
The efficient extraction of electrons from photosynthetic microorganisms remains a critical challenge in living biophotovoltaics (BPV). While nanomaterials can facilitate electron transport, their stochastic adsorption leads to inefficient material-wasteful interfaces. Here, we demonstrate a controllable approach to direct the targeted assembly of gold nanoparticles (AuNPs) onto the type IV pili of Synechocystis sp. PCC 6803 by using a genetically encoded gold-binding peptide. This approach creates a spatially precise conductive nano-bio interface on the cell envelope that serves as a dedicated electron conduit between photosynthetic electron transport chains (PETCs) and electrodes. This nano-bio interface enhances electron transfer through synergistic improvements in interfacial charge transfer and biofilm density, ultimately yielding a four-fold increase in photocurrent density, while using two orders of magnitude less gold than non-targeted strategies. Moreover, the AuNPs can be transferred from inactivated to fresh cells, indicating a potential pathway for long-term stability. This work establishes a generalizable strategy for the rational design of conductive interfaces on living cells, with implications for biophotovoltaics, microbial electrosynthesis, and next-generation biohybrid devices. Inefficient extracellular electron transfer of cyanobacteria impedes power output in biophotovoltaics. Here, the authors assemble a gold conductive layer on the cell surface by attaching gold-binding peptide to the pili structure, achieving a four-fold increase in photocurrent generation.

|
Scooped by
mhryu@live.com
April 25, 4:01 PM
|
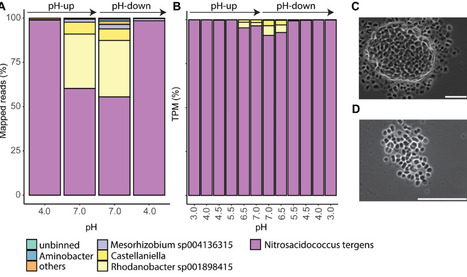
Acidophilic ammonia-oxidizing bacteria (AOB) have only recently been discovered. These organisms hold great promise for acidic wastewater treatment; however, their physiology remains poorly understood compared to that of neutrophilic AOB. Here, we investigated the physiology of the acidophilic AOB “Candidatus Nitrosacidococcus tergens” sp. RJ19 across a broad pH range (2.5–7.0) using a specialized bioreactor system. We monitored nitrogen (N) transformations combined with microbial community composition and transcriptomic profiles, focusing on nitrogen metabolism and proton stress responses. Our results show that above pH 6.0, “Ca. Na. tergens” performs complete and stoichiometric conversion of ammonium to nitrite, coinciding with isotopic fractionation effects specific for ammonia oxidation and increased expression of key ammonia oxidation genes. The apparent absence of nirK and cycA did not impede ammonia oxidation, suggesting that these genes are non-essential in this context. Below pH 6.0, nitric oxide and nitrate accumulated, and nitrous oxide (N₂O) levels, although negligible compared to the other N-compounds, peaked near pH 4.0. Stable isotope analysis, including the site-specific 15N-enrichment at the inner (α) and outer (β) nitrogen positions of the N2O molecule, indicated nitrifier-denitrification as the source of N₂O, supported by the highest norB expression at this pH. These findings provide new insights into the acid-tolerant physiology of “Ca. Na. tergens” and advance its potential application in engineered nitrogen removal systems under acidic conditions.

|
Scooped by
mhryu@live.com
April 25, 3:45 PM
|
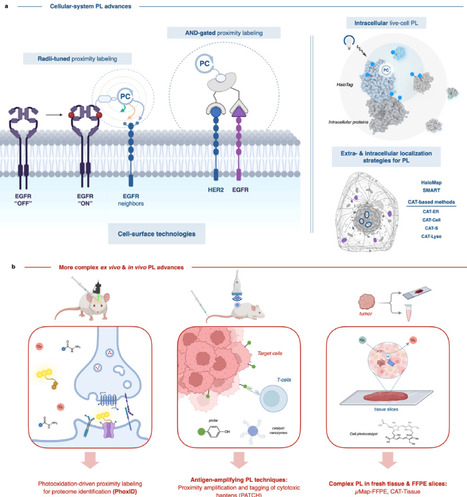
Protein–protein interactions (PPI) and spatially restricted molecular contacts govern cellular function, yet many are poorly captured by classical biochemical approaches that rely on cell lysis or stable complex isolation. Proximity labeling (PL) technologies have transformed interactome analysis by enabling covalent tagging of biomolecular neighborhoods directly within intact cells, tissues, and living organisms. By generating short-lived reactive species, PL provides spatially and temporally resolved snapshots of molecular organization under native conditions. Recent advances across enzymatic, chemical, and photocatalytic PL platforms have expanded control over labeling radius, kinetics, and activation, while reducing background and enabling microenvironment-specific targeting. Hybrid genetic-chemical and optogenetic strategies further extend PL beyond mapping toward proximity-based signal amplification and functional interrogation. This review focuses on the most significant methodological and conceptual advances in proximity labeling reported over the past two years, highlighting how these developments have enabled discovery of previously inaccessible interaction networks, including membrane assemblies, chromatin complexes, and in vivo protein microenvironments. We conclude by outlining key challenges and future opportunities for proximity labeling in interactome mapping and amplification.
|
 Your new post is loading...
Your new post is loading...





















droplet