Your new post is loading...

|
Scooped by
mhryu@live.com
Today, 11:35 PM
|
The biological conversion of methanol into valuable chemicals represents a promising route for achieving global carbon neutrality. However, the rational engineering of complex industrial traits in microbial cell factories is hindered by an incomplete understanding of cellular metabolism and regulation. In this study, we developed a Random Mutagenesis Platform for Accelerated Genome Evolution (RAMPAGE) for the yeast Pichia pastoris (Komagataella phaffii). This platform enables continuous genome-wide diversification and facilitates the rapid evolution of superior phenotypes under defined selective pressures. Using RAMPAGE, we evolved yeast strains with significantly enhanced tolerance to extreme industrial conditions, including growth in 18% methanol, survival at 40°C, and resistance to methanol and thermal stresses. Genome resequencing of these strains revealed a collection of genes associated with methanol tolerance and thermotolerance. In addition, we used RAMPAGE to evolve a glycoengineered yeast strain with improved growth and stress tolerance. Subsequent genome resequencing and functional validation identified novel genetic engineering targets to improve the cellular fitness of glycoengineered yeast strains. By providing a simple, efficient, and programmable tool for continuous genome evolution, RAMPAGE expands the synthetic biology toolkit for P. pastoris, enabling its broad application in industrial biotechnology.

|
Scooped by
mhryu@live.com
Today, 11:24 PM
|
Understanding how life emerged from non-living matter remains one of the most profound challenges in science. Empirical constraints and the scarcity of ancient evidence make this question particularly suitable for theoretical and computational approaches. Here, we review recent progress toward a systems-level understanding of life’s origins, focusing on how mathematical models describe the progressive emergence of complexity across three interconnected levels: chemical, informational, and ecological. At the chemical level, models of autocatalytic networks and protometabolic organization capture how self-sustaining reaction systems and feedback loops could arise spontaneously under out-of-equilibrium conditions. At the informational level, studies of polymerization and template-assisted replication of biopolymers shed light on how simple molecular systems could give rise to the emergence of catalytic RNA, genetic heritable information and error-prone molecular evolution. Finally, ecological and thermodynamic models illuminate how protocells and subsequent microbial consortia might have diversified, interacted, and self-organized into the first ecosystems. Together, these approaches highlight a sequence of transitions and bifurcations that call for the development of a coherent framework for studying life’s origins and early evolution, grounded in complex systems theory, prebiotic chemistry, and ecological dynamics. We believe that such an integrative modeling effort will be essential for identifying universal principles underlying the emergence of living systems, bridging the current gap between molecular and ecological levels of organization, and guiding future experimental and computational research in origins-of-life studies.

|
Scooped by
mhryu@live.com
Today, 9:54 PM
|
Dinoflagellates and diatoms are key marine phytoplankton, with ecological roles strongly influenced by their associated phycosphere bacteria. However, the ecophysiological functions of these bacteria remain enigmatic as a result of insufficient taxonomic and genomic characterization. Here, by combining single-cell isolation with a custom statistical pipeline, we profiled resident bacterial communities associated with 108 diatom and 86 dinoflagellate strains, collected across temperate and tropical oceans worldwide. We examined genomic traits of key bacterial populations through whole-genome sequencing of representative isolates. Taxonomic compositions of dinoflagellate- and diatom-associated microbiota were distinct, highlighting host-specific differences. Each microbiota harbored characteristic genera with adaptive traits reflecting host metabolic profiles. Dinoflagellate-associated bacteria were enriched in genes responsible for motility and sulfur-compound use, whereas diatom-associated bacteria specialized in glycan use. We identified ‘foundation’ genera, defined as taxa with high occupancy and community-level impact in both phycosphere types (for example, Marivita and Marinobacter), sharing host-specific traits with characteristic bacteria while universally excelling in environmental response and resistance. Notably, foundation bacteria were enriched in Type VI secretion systems T6SS, emerging as a universal hallmark of phycosphere bacteria across global oceans. Overall, this study provides insights into the taxonomic and metabolic nature of phycosphere bacteria, highlighting the profound influences of interspecific interactions on marine ecological processes. This study reveals that some bacteria living in the phycosphere of marine diatoms and dinoflagellates are host-specific with genomic functions matching their hosts, while some termed as ‘foundation’ bacteria can span both phycosphere types.

|
Scooped by
mhryu@live.com
Today, 6:13 PM
|
Generalist biological artificial intelligence (GBAI) represents a transformative approach to modeling the ‘language of life’—the flow of information from DNA to cellular function. This Review synthesizes rapid advances in biological AI to interpret and generate DNA, RNA, proteins and cellular systems. We chart a course toward comprehensive systems that can concurrently process and predict across these domains, performing several critical biological tasks simultaneously. Substantial opportunities lie in synergizing language and structural AI, leveraging specialized models and improving AI agents for autonomous discovery. After addressing challenges in data, biological complexity, scalability and experimental validation, GBAI has the potential to deepen our understanding of disease pathways and biomarkers, advance automated therapeutic design and evaluation, and integrate within virtual cells to meaningfully simulate biological activity. This Review discusses the promises and pitfalls of biological AI algorithms and presents a vision for generalist biological artificial intelligence, in which models can perform diverse tasks across biological domains.

|
Scooped by
mhryu@live.com
Today, 5:45 PM
|
Viral sequences in diverse environments remain largely uncharacterized, impeding our comprehension of their genetic makeup, biological interactions, and potential applications. This underscores an urgent need for innovative analytical methods. Here, we present the VirHost Hunter framework, which employs phage tails and lysins, bypassing the requirement for full genomes, for efficient and high-resolution host assignment. By harnessing Protein Language Models and Vision Transformers, VirHost Hunter captures protein functional homology despite sequence dissimilarity, significantly boosting prediction accuracy. In the scenario of disease-associated gut bacteria, the calibrated VirHost Hunter surpasses existing methods, doubling phage host assignments, expanding taxonomic reach, and revealing previously uncharacterized phages targeting gut bacteria, including Akkermansia and Prevotella. Therefore, we establish a gut phage lysin database, enabling the synthesis of a lysin that effectively and specifically targets an obesity-promoting bacterium. VirHost Hunter’s precision and scalability mark a significant leap forward in virome research and present a promising avenue for microbiome therapies. Here, the authors present VirHost Hunter, an AI-based approach to phage–host assignment using tail and lysin proteins, showing it improves host resolution, expands functional discovery of gut phage, and enables targeted lysin identification for microbiome research.

|
Scooped by
mhryu@live.com
Today, 5:36 PM
|
In biotechnological applications, it is often necessary to introduce genes or entire pathways into a host cell, which can create a significant metabolic burden on the host, limiting productivity. In this study, we systematically investigated the physiological stress responses of Pseudomonas putida during heterologous protein production using a modular monitoring system consisting of a plasmid encoding a heterologous protein fused to eGFP and a chromosomally integrated capacity reporter. Our findings reveal that translation is the main bottleneck, with translational capacity becoming saturated under high expression loads. While increasing the strength of the ribosome binding site improved protein production for non-burdensome proteins, this effect was not observed for larger fusion proteins. Variations in fusion protein size suggested that translational demand, rather than the overall mass of protein produced, determines metabolic burden. We further evaluated how resource availability affects protein expression by modifying the metabolic regime or supplementing with amino acids. While the carbon source affected cellular capacity, it did not significantly alter heterologous protein production. Amino acid supplementation alleviated the growth defects of MBPeGFP-producing cells and modestly improved protein production rates. Together, these findings emphasize that metabolic burden is influenced not only by the size of the produced protein but also by transcript architecture, resource allocation, and the physiological state of the host. Therefore, successful optimization of heterologous protein production requires a holistic approach integrating construct design with host physiology and cultivation strategies.

|
Scooped by
mhryu@live.com
Today, 5:07 PM
|
Proximity labeling (PL) has emerged as a powerful technology for mapping subcellular compositions and molecular interactions. This approach employs promiscuous enzymes that generate reactive species to tag endogenous biomolecules, which are then identified by mass spectrometry or sequencing. However, conventional PL methods—such as peroxidases, biotin ligases, and photocatalytic systems—face significant limitations for in vivo applications, hindering their use in native biological contexts. In this review, we first summarize the application of peroxidases and biotin ligases in living animals, highlighting how they have provided insights into cell surface proteomes and cell type-specific secretion, despite their constraints. We then explore recent advances in in vivo-compatible PL technologies, including novel enzymes like tyrosinase, laccase, and lipoic acid ligase, as well as innovative photocatalytic strategies activated by near-infrared light, ultrasound, or bioluminescence. These emerging tools hold great potential to expand spatial multi-omics from cellular systems to living organisms.

|
Scooped by
mhryu@live.com
Today, 4:49 PM
|
The generation of complex traits involves the coordinated interplay of multiple gene networks. Elucidating the function of transcriptional cis-regulatory elements (CREs) in regulating gene expression is crucial for understanding complex regulatory pathways and improving our ability to modify macro-phenotypes. While traditional bulk sequencing approaches rely on tissue or cell population aggregates, single-cell transcriptomics provides a more precise perspective by capturing cell-type-specific information. The integration of single-cell technology with genome-wide genetic screening, particularly through the single-cell CRISPR (scCRISPR) system, enables the identification of critical regulatory elements and provides novel insights into gene-expression control mechanisms. Here, we summarise recent advances in diverse strategies for functional genome analysis using the scCRISPR system, with an emphasis on its potential to revolutionise single-cell genetic screening of CREs. We also explore the challenges and opportunities for applying these approaches in plant research.

|
Scooped by
mhryu@live.com
Today, 4:06 PM
|
Although sequence-discrete species appear to dominate microbial communities, readily distinguishing between distinct populations of a species recovered from different short-read metagenomic samples is challenging due to technical limitations associated with read length. To close this gap, we developed a novel algorithm to evaluate which reads in a metagenome belong to a target population based on the distribution of sequence identities of reads aligned to a reference sequence, which are filtered using a Kernel density estimation (KDE) as a flexible alternative to the commonly used static 95% nucleotide identity cutoff. Subsequently, we employed the average nucleotide identity of reads (ANIr) aligning above the KDE threshold, and resampling techniques for estimating the confidence intervals of ANIr values, to quantify intra-population sequence diversity and compare populations across globally representative marine samples. Most populations showed high ANIr in only a few samples at similar depths, and decreased ANIr and increased gene-content difference between samples where a closely related population is detected [e.g., same 95% ANI-based species]. Accordingly, ANIr correlated with the physical distance between the samples, and only a few truly cosmopolitan populations were identified. Among the latter, Alteromonas macleodii (97% average amino-acid identity -or AAI- to the type genome) and Prochlorococcus marinus (79% AAI) showed high relative abundance in both surface (0-200m) and deep (>1000m) samples. These results suggest that microbial communities under different environmental conditions share very few identical and abundant populations, and provide a highly needed methodology to track such populations over space and time, in marine or other habitats.

|
Scooped by
mhryu@live.com
Today, 3:36 PM
|
Wound infections are becoming increasingly difficult to treat due to rising antibiotic-resistant bacteria. β-Lactamase–producing bacteria are among the most common pathogens implicated in these infections. Here, we report a bacterial enzyme-responsive hydrogel formulated with a cephalosporin-derived, β-lactamase–cleavable crosslinker that undergoes selective degradation in the presence of bacterial β-lactamases. This degradation triggers the on-demand release of encapsulated ciprofloxacin-loaded liposomes, ensuring that antibiotic delivery occurs only at the site of infection. This selective degradation and release was demonstrated in both ex vivo and in vivo models of Pseudomonas aeruginosa wound infections. In a murine skin abrasion infection model, a single application of the hydrogel led to complete bacterial eradication and enhanced wound healing, outperforming a commercial silver-based hydrogel wound dressing. These responsive hydrogels did not induce ciprofloxacin resistance in non–β-lactamase–producing bacteria. These findings demonstrate that β-lactamase–responsive hydrogels provide a precise, infection-triggered antibiotic delivery platform that can improve the treatment of wound infections and mitigate antimicrobial resistance.

|
Scooped by
mhryu@live.com
Today, 3:19 PM
|
The accelerating growth of plant science knowledge presents a major challenge for researchers seeking to extract accurate, up-to-date knowledge from an increasingly fragmented and domain-specific corpus. General-purpose large language models (LLMs), while powerful, often misinterpret plant science terminology and lack mechanisms for source traceability. We created PlantScience.ai, a virtual plant biology scientist powered by our automated scientific knowledge graph construction pipeline (AutoSKG). PlantScience.ai exhibits expert-level reasoning in plant biology and maintains scholarly rigor in its citations. Through continuous learning, it integrates the latest research, ensuring that its knowledge base remains current and scientifically robust. Apart from providing the answers to the scientific questions, PlantScience.ai can interact with human scientists, follow instructions, and retrieve information with citation awareness, grounding each response in primary sources to ensure accuracy and verifiability. PlantScience.ai marks a pivotal advance toward a collaborative scientific paradigm in which virtual and human plant scientists work synergistically to accelerate discovery while preserving the unique value of human insight. PlantScience.ai is available at https://plantscience.ai .

|
Scooped by
mhryu@live.com
Today, 3:02 PM
|
Plant root-associated anoxic microsites may influence the fate of nutrients and contaminants in the rhizosphere, but their dynamics remain relatively unknown. To examine the formation of root-induced anoxic microsites over space and time, we use microfluidic devices integrated with transparent, planar oxygen sensors in a wheat (Triticum aestivum) rhizosphere, with and without soil microorganisms. We found that suboxic (< 2% air saturation) conditions commonly establish at root tips and more rarely establish along more mature root segments, particularly in the presence of soil organic matter and complex microbial communities. Additionally, the distribution of oxygen, and thus root-induced anoxic microsites, depends on complex interactions among light–dark cycles, growth rate, and presence of microorganisms in the rhizosphere. This study provides real-time observations of the micron-scale oxygen dynamics around actively growing roots, thereby linking root physiology to anoxic microsite formation in the rhizosphere. Our work suggests a strong potential for root-driven anoxic microsite formation, prompting important questions about anoxic microsite impact on biogeochemical processes in natural rhizosphere soil.

|
Scooped by
mhryu@live.com
Today, 2:25 PM
|
Chlorophyll is one of the most abundant pigments on Earth. Although its degradation is well understood in plants, the role of prokaryotes in this process - despite their vast metabolic capabilities - remains unknown. Recent developments in the field of AI-predicted protein structures have opened new avenues for investigating functional homologies between evolutionary-distant organisms previously inaccessible through traditional sequence- or profile-based methods. Here, we present the first evidence of Chlorophyll a (Chl a) degradation by prokaryotes, discovered through a novel bioinformatic framework which bridges the gap across the domains of life via structural alignments of functionally characterised plant proteins, followed by structure similarity graph-based clustering. Metagenomic sequencing data was assembled and binned, yielding over 70,000 medium- to high-quality genomes in total, furthermore publicly available datasets containing genomes from prokaryotic isolates, metagenome-assembled genomes, as well as single-cell genomes were then mined for prokaryotic homologues of Chl a degradation genes. Our analysis revealed over 400 genomes from diverse taxonomic groups and habitats that possess a complete pathway, more than 50% stemming from isolates. Additionally, many other genomes harbor partial pathways, suggesting that Chl a degradation capabilities are globally widespread across diverse ecosystems. We then validated our in silico findings using the model organism Shewanella acanthi and confirmed its Chl a degradation capability via growth experiments, fluorescence spectroscopy and HPLC analyses. Our findings reveal a previously unrecognized pathway in prokaryotes, highlighting the power of structure-based remote homology detection for uncovering metabolic capabilities and evolutionary relationships.
|

|
Scooped by
mhryu@live.com
Today, 11:31 PM
|
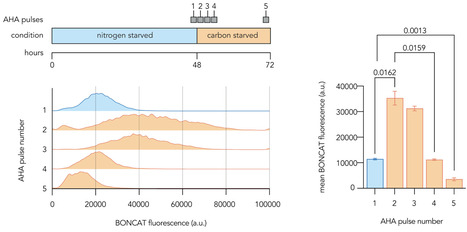
Heterotrophic bacteria rapidly deplete essential macronutrients during growth and must navigate subsequent periods of growth arrest imposed by starvation. Nutrient limitations can be dynamic in nature, requiring ongoing regulatory adjustments involving new protein synthesis despite total biosynthetic activities being dramatically lower than during growth. Here, we have characterized the responses of the opportunistic pathogen Pseudomonas aeruginosa to prolonged starvation for carbon or nitrogen sources and to transitions between these states. We find that most cells survive both types of starvation for more than a week and maintain low but robustly detectable levels of protein synthesis in the absence of growth. Nitrogen-starved cells are larger, make more proteins, and retain fewer ribosomes than carbon-starved cells, indicating that distinct physiological strategies are adopted during the two starvation types. We found that the newly synthesized proteomes of each starvation type are distinct although many of the most highly synthesized proteins are shared between both conditions. Interestingly, we observed a temporary burst of protein synthesis as cells were transitioned between the two starvation conditions, which may reflect active remodeling of the proteome during growth arrest. We also used transposon insertion sequencing to identify genes impacting fitness in both starvation conditions and during transitions between the two and found that a highly overlapping set of global regulators most strongly influenced survival. Combining these data sets, we highlight proteases and chaperones, flagellar motility, and the nitrogen-related phosphotransferase system as key fitness-impacting functions that are actively maintained by growth-arrested Pseudomonas aeruginosa.

|
Scooped by
mhryu@live.com
Today, 9:59 PM
|
The field of protein therapeutics has evolved from native biologics to precision-engineered multi–functional therapeutic agents, overcoming inherent limitations such as structural instability, rapid clearance, and immunogenicity. While early strategies, including PEGylation and albumin fusion, and the emergence of antibody–drug conjugates (ADCs), have demonstrated clinical value, challenges remain in improving the site-specificity of modifications, suppressing structural heterogeneity, adjusting linker stability, and broadening payload diversity for protein therapeutics. Recent advances in bioconjugation techniques, such as chemoselective reactions, enzymatic labeling, glycan engineering, and genetic code expansion, now enable more site-specific and homogeneous modifications, achieving enhancements in batch consistency, controlled payload release, and multi-functionality. This review highlights key developments over the past two years across three categories of modified protein therapeutics: ADCs, cytokines, and proximity-enabled covalent therapeutics. Looking forward, we outline future directions focused on scalable site-specific platforms, immunogenicity management, and delivery optimization, increasingly propelled by artificial intelligence-aided protein design and reaction manipulation.

|
Scooped by
mhryu@live.com
Today, 9:51 PM
|
Smart wearable and implantable biosensors enable continuous, real-time monitoring of biophysical and biochemical signals for personalized and preventive healthcare. Advances in flexible, stretchable, and biocompatible materials ensure long-term comfort and seamless body integration, while multimodal and multi-analyte sensing improves robustness. AI enhances signal processing and predictive insights, yet challenges remain in motion artifacts, energy autonomy, data privacy, and clinical interpretability. This review summarizes materials, device architectures, and AI-assisted strategies applications.

|
Scooped by
mhryu@live.com
Today, 6:11 PM
|
Group I introns are catalytic RNAs capable of self-splicing, yet structural insights into full-length precursor RNAs and post-splicing circularization have been limited. Here, we present cryo-EM structures of the Azoarcus pre-tRNAIle system across key catalytic stages, including the full-length precursor in Pre-1S conformations, the linear intron, and the circular introns, at 2.8-3.3 Å resolution. These structures reveal a preformed P1 helix, register shifts during splicing, and a conformational flip of G37 that stabilizes the circularization site. Biochemical analyses confirm a two-step circularization mechanism, generating a secondary circular product via an alternative ligation site. Together, our results provide an atomic-level view of a group I intron system through splicing and circularization. This work uncovers structural principles governing RNA conformational dynamics, catalysis, and circular RNA formation, with broad implications for ribozyme engineering. Cryo-EM structures of the Azoarcus group I intron capture its progression from pre-splicing to circular RNA products, revealing coordinated RNA rearrangements that drive catalysis and a two-step circularization pathway.

|
Scooped by
mhryu@live.com
Today, 5:38 PM
|
Peptide aggregation is a long-standing challenge in chemical peptide synthesis, limiting its efficiency and reliability. Although data-driven methods have enhanced our understanding of many sequence-based phenomena, no comprehensive approach addresses so-called non-random difficult couplings (generally linked to aggregation) during solid-phase peptide synthesis. Here we leverage existing peptide synthesis datasets, supplemented with further experimental data, to build a predictive model that deciphers the role of individual amino acids in triggering aggregation. We first identified and experimentally validated composition-dependent aggregation as a stronger predictor than sequence-based patterns. This insight enabled the development of a composition vector representation, allowing insights into the aggregation propensities of individual amino acids. Applying an ensemble of trained models, we predicted the aggregation properties of peptides and recommended the optimized use of aggregation-reducing tools. By elucidating each individual amino acid’s influence, this method holds the potential to accelerate synthesis optimization through existing data, offering a robust framework for understanding and controlling peptide aggregation. Aggregation during solid-phase peptide synthesis is a bottleneck, often rendering peptide synthesis costly and inefficient. Now the impacts of individual amino acids on aggregation have been characterized through machine learning and analysis of composition patterns. The generated models predict problematic peptides and optimize synthesis conditions, enabling chemists to use aggregation-avoiding routes.

|
Scooped by
mhryu@live.com
Today, 5:10 PM
|
Soils are heterogeneous and dynamic systems characterized by complex physical, chemical, and biological interactions. Understanding these interactions is critical, as they influence plant productivity, global biogeochemical cycles, and ecosystem resilience. While ecologists have long studied soils in field, greenhouse, and laboratory settings, their complexity and heterogeneity make it challenging to pinpoint key properties driving biological processes and derive mechanistic insights. Advancements in synthetic biology, which seeks to engineer and control biological processes in soils, have increased the demand for standardized and controllable experimental platforms. These platforms, referred to here as ‘synthetic soils’, are systems designed to reproduce selected physicochemical characteristics of natural soils in a simplified and defined format, allowing scientists to systematically change soil physicochemical properties (i.e. texture, mineralogy, pH) to study how biological components (i.e. microbes, plants, soil fauna, etc.) respond to, modify, or interact within these controlled environments. This review explores existing synthetic soils, their advantages, limitations, and applications in ecology and synthetic biology, and discusses potential directions for their future development.

|
Scooped by
mhryu@live.com
Today, 5:01 PM
|
RNAs exhibit complex dynamics in cells, including expression, splicing, localization, translation and degradation, and these processes are highly coordinated and tightly regulated both spatially and temporally. To better understand the biological function of diverse RNAs, approaches that allow monitoring of RNA with high spatiotemporal resolution are essential. Fluorescent RNAs (FRs), fluorescent protein–like entities consisting of RNA aptamers and their cognate fluorogenic dyes, have emerged as a promising approach for imaging RNA dynamics in live cells. We recently reported the development of several high-performance FRs, named Pepper, Clivia and Okra, that show advantageous properties, including high cellular brightness and photostability, low ion dependence and/or large Stokes shifts, and have been used to image diverse RNA species in live cells. In this protocol, we provide easy, efficient and generalizable strategies for using FRs to visualize different RNA species in bacteria and mammalian cells by expressing the RNA of interest tagged with one or more copies of the aptamer. We also provide a detailed procedure for multiplexed RNA imaging using orthogonal FRs and the steps to perform super-resolution live imaging of RNAs. The protocol typically takes 5–7 d, including cloning, transfection of mammalian cells or transformation of bacteria, live imaging and results analysis. This protocol is applicable to the real-time monitoring of the localization and dynamics of RNAs of interest in live cells. This protocol provides guidelines for using fluorescent RNAs, entities consisting of an RNA aptamer bound to its cognate fluorogenic dye, for live imaging of the localization and dynamics of different RNA species in bacteria and mammalian cells.

|
Scooped by
mhryu@live.com
Today, 4:45 PM
|
In this work, we describe an engineering approach that leverages ecological drift to generate Minimal Microbiomes; microbial consortia that are relatively simple, cohesive, and functionally complete. This process can be applied to any microbial ecosystem, provided that the target microbiome can be experimentally mimicked. Empirical support for this approach has emerged from multiple independent studies. We use simulations across diverse scenarios, significantly varying niche structures and biotic interactions, to explore the experimental conditions and source microbiome characteristics that favor successful outcomes, within a computational framework that also enables the study of microbial community assembly. Our results indicate that the effectiveness of this approach is constrained by several factors, and that perfect outcomes should not be routinely expected. Nevertheless, despite its drawbacks, this strategy remains a powerful tool for simplifying microbiomes and isolating key co-adapted populations, enabling the construction of low-diversity consortia that retain community function and present ecological cohesion.

|
Scooped by
mhryu@live.com
Today, 3:49 PM
|
Protein expression levels optimize cell fitness: Too low an expression level of essential proteins will slow growth by compromising essential processes, whereas overexpression slows growth by increasing the metabolic load. This trade-off naïvely predicts that cells maximize their fitness by sufficiency, expressing just enough of each essential protein for function. We test this prediction in the naturally competent bacterium Acinetobacter baylyi by characterizing the proliferation dynamics of essential-gene knockouts at a single-cell scale (by imaging) as well as at a genome-wide scale. In these experiments, cells proliferate for multiple generations as target protein levels are diluted from their endogenous levels. This approach facilitates a proteome-scale analysis of the fitness landscape with respect to protein abundance. We find that most essential proteins are subject to a threshold-like fitness landscape: Growth is independent of protein abundance above a critical threshold and arrests below that threshold. We have recently analyzed the implications of this landscape for growth robustness. Confirming signature predictions of this model, we find that (i) roughly 70% of essential proteins are overabundant, (ii) overabundance increases as the expression level decreases, and (iii) the lowest abundance proteins are in vast excess (>10×) of what is required for growth in the typical cell. These results reveal that robustness plays a fundamental role in determining the expression levels of essential genes and that overabundance is a key mechanism for ensuring robust growth.

|
Scooped by
mhryu@live.com
Today, 3:28 PM
|
Broad-host-range plasmids drive the spread of antibiotic resistance, particularly in surface-associated microbial systems prevalent in natural and host-associated environments. Predicting their realized host range is challenging because both transconjugant proliferation (vertical gene transfer, VGT) and conjugation (horizontal gene transfer, HGT) contribute to transconjugant diversity. Here, we hypothesized that the realized host range is determined by the interplay between VGT and HGT. We experimentally tested this hypothesis by analyzing transconjugant diversity under conditions that differ in their ability to support bacterial growth. Fast-growth conditions increased transconjugant abundance but reduced diversity, whereas slow-growth conditions supported fewer but more diverse transconjugants. We complemented these experiments with individual-based simulations that explicitly incorporated both VGT and HGT. Our results demonstrate that the realized host range is jointly governed by initial HGT events and subsequent VGT-driven expansion, highlighting the importance of integrating transfer and post-transfer dynamics when predicting plasmid-mediated antibiotic resistance spread.

|
Scooped by
mhryu@live.com
Today, 3:16 PM
|
Viral proteins interact with host proteins to hijack cellular pathways important for viral replication. Viral mimics are proteins whose structural similarity to host-mimicked proteins allows them to interact with mutual host targets. This mimicry poses a challenge for the host—how to avoid mimics without compromising essential interactions with host-mimicked proteins. Despite the prevalence of mimicry, the evolutionary dynamics between host and viral mimics remain largely unknown. We address this by integrating structural modeling, host–virus interaction networks, and comprehensive evolutionary analyses of host and viral proteins. We show that host proteins targeted by mimics and host-mimicked proteins are highly conserved, and that this is related to functional constraints imposed on host proteins. Host interface residues that interact with both mimics and host-mimicked proteins evolve slowly, while residues that exclusively interact with mimics evolve significantly faster. Surprisingly, viral mimics do not evolve rapidly, instead displaying complex evolutionary patterns. Our analysis reveals host’s limited capacity to escape mimicry and viral evolution to exploit this, and highlights how constraints lead to unexpectedly slow evolution of host–virus interaction networks.

|
Scooped by
mhryu@live.com
Today, 2:58 PM
|
Plant roots form symbioses with beneficial microorganisms to enhance nutrient acquisition. Most terrestrial plants form arbuscular mycorrhizal symbiosis (AMS) with obligate biotrophic Glomeromycotina fungi, which supply hosts with mineral nutrients in exchange for carbon through specialized symbiotic hyphal structures (arbuscules) that develop within root cortex cells. Legumes form root nodule symbiosis (RNS) with nitrogen-fixing rhizobia, which are housed as differentiated bacteroids within specialized symbiotic organs (nodules) and provide plants with ammonia in return for carbon. RNS exhibits high partner specificity, occurring only between compatible hosts and microbes. Conversely, AMS is less specific, although symbiosis outcomes are context-dependent and influenced by host and fungal genotype, environmental conditions, and microbial competition. In both cases, plants favor high-performing microsymbionts by recognizing them during symbiosis initiation or by punishing low-performing symbionts through postcolonization sanctions. Microbes, in turn, employ strategies to manipulate plants for their own benefit. Here, we review the molecular mechanisms underlying partner preference in beneficial plant–microbe interactions and discuss how host partner selection strategies maintain mutualistic stability in AMS and RNS, alongside microbial strategies to evade host control. Understanding the dynamic interplay of functionally diverse plant–microbe symbioses provides a basis for improving mutualisms in both natural and agricultural systems.
|
 Your new post is loading...
Your new post is loading...