 Your new post is loading...
 Your new post is loading...

|
Scooped by
Jean-Michel Ané
Today, 3:00 PM
|
In root nodule symbiosis, symbiosome compartments accommodate nitrogen-fixing rhizobia inside the plant cell. Differentiated into bacteroids, the rhizobia are surrounded by a peribacteroid space and a plant-derived peribacteroid membrane, which separates them from the plant cytoplasm but allows signal and nutrient exchange between host and microbe. The morphological features of symbiosomes are primarily determined by ultrastructural single focal plane imaging, with limited information about spatial details. This study combines 2D and 3D imaging, using transmission electron microscopy and focused ion beam scanning electron microscopy as complementary techniques to analyse the symbiosome ultrastructure and organisation in Lotus japonicus wild-type plants. The 3D model of a mature colonised root nodule cell region demonstrates a dense, puzzle-like arrangement of symbiosomes relative to one another and adjacent plant organelles. The symbiosome shape and size depends on the orientation and number of bacteroids within the compartment and features connective tubular structures. Furthermore, vesicular structures, some likely of bacterial origin, were present at the interface. The study presents a multi-angled analysis of symbiosome-related structures, highlighting their volumes, spatial distribution, and pronounced compactness. Interface associated vesicles, protrusions and connective structures hint towards a dynamic and flexible system that contributes to the plant-microbe crosstalk.

|
Scooped by
Jean-Michel Ané
Today, 2:52 PM
|
Development of arbuscular mycorrhiza (AM), a symbiosis between plants and beneficial Glomeromycotan fungi, is largely under plant control. Several genes, required for AM development, are proposed to be regulated by the karrikin signalling module, comprising the alpha/beta hydrolase receptor KARRIKIN INSENSITIVE 2 (KAI2), the F-box protein MORE AXILLARY GROWTH2 (MAX2) and the transcriptional repressor SUPPRESSOR OF MAX2 1 (SMAX1), which is ubiquitylated for proteasomal degradation upon KAI2-ligand-induced binding to the KAI2-MAX2 complex. Rice and Brachypodium distachyon kai2 mutants are incapable of forming AM. Here, we show that in Lotus japonicus, Pisum sativum, and Nicotiana benthamiana, KAI2 only quantitatively affects AM development, indicating angiosperms vary in their requirement for KAI2-signalling to support AM. Comparative transcriptomics of L. japonicus and B. distachyon roots after treatment with fungal signalling molecules revealed some AM-relevant genes respond KAI2-independently in L. japonicus but not in B. distachyon. Consistently we obtained

|
Scooped by
Jean-Michel Ané
Today, 2:43 PM
|
Nodules cystine-rich (NCR) peptides from legume nodules mediate bacteroid differentiation of Rhizobium sp. This study aimed to characterize NCR peptides of Phaseolus vulgaris, Trifolium repens, Medicago sativa, and Vicia faba, as relevant information on these legumes is still lacking. After exposing the root nodules of four leguminous plants to sonication, the peptides were separated according to their molecular weight by sodium dodecyl gel electrophoresis (SDS-PAGE). The NCR peptide was detected by high-performance liquid chromatography (HPLC), followed by the determination of the protein sequence and amino acid composition and their evolutionary history in terms of the phylogenetic tree by bioinformatics analysis. The results showed these four peptides had the same molecular weight of 60 kDa. The peaks of the curve varied with retention time depending on the legume from which the peptide was extracted. Bioinformatic analysis of the results showed that the amino acid substitutions varied in number and sequence for each species and that these peptides had three distinct wire, ribbon, and surface patterns within the structural modeling of their crystals. Side chains of amino acids in the loops and helices within the grid box may facilitate hydrophobic interactions via tyrosine, phenylalanine, or polar interaction sites via cysteine, histidine, and glutamine. These loops often serve as flexible regions to adapt to binding partners. In general, this study contributed to the characterization of NCR peptides that showed mismatches and differences in amino acid sequences and deletions of some amino acids at different locations depending on the type of legume from which the peptide was extracted.

|
Scooped by
Jean-Michel Ané
May 8, 10:21 AM
|
Plant-associated microbiomes are central to crop productivity, nutrient efficiency, and stress resilience, yet conventional microbiome manipulatio

|
Scooped by
Jean-Michel Ané
May 5, 8:55 PM
|
Background
Phosphate (Pi) is essential for plant growth, but its bioavailability in soil is often limited. Forming symbiotic associations with arbuscular mycorrhizal fungi can significantly enhance the host plant’s root Pi acquisition ability. PHT1 transporters, such as MtPT4, are involved in Pi uptake from arbuscules. It is unknown how Pi is exported from the mycorrhizal root to the vascular bundle.
Results
Here, we found the medicago vascular apoplastic Pi exporter PHO1 proteins not only mediate Pi transport in the direct Pi uptake pathway but also influence arbuscular mycorrhizal symbiosis. MtPHO1s showed Pi export activities when expressed in nicotiana leaves. The MtPHO1.3 tnt1 insertion mutant displayed significant growth inhibition and reduced shoot Pi content. Moreover, AM colonization was lower in the Mtpho1.3 mutant compared to the wild type. Consistent with this, the expression level of the strigolactone biosynthesis gene DWARF27 was suppressed in Mtpho1.3 mutant, and root exudates from the mutant induced less fungal hyphal branching. Furthermore, the Mtpho1.3 mutant exhibited increased arbuscule degeneration, suggesting that Pi levels in the plant are key to regulating arbuscule lifespan.
Conclusion
Our analysis shows that MtPHO1.3 also exerts function in mediating Pi translocation during arbuscular mycorrhizal symbiosis.

|
Scooped by
Jean-Michel Ané
May 4, 3:59 PM
|
Symbiotic nitrogen fixation (SNF) relies on aerobic respiration, yet the key enzyme, nitrogenase, is extremely oxygen labile. Leghemoglobin (Lb) resolves this “oxygen paradox” by buffering and facilitating O2 transport. However, the dynamic regulation of Lb during nodule development remains poorly understood. Earlier results from our laboratory demonstrated that site-specific serine phosphorylation of Lb reduces its oxygen sequestration capacity. Here, we investigated the spatio-temporal regulation of Lb with the progress of rhizobial load during SNF. Fluorescence immunohistochemistry (FIHC) using anti-Lb antibody revealed that its localization gradually shifted from the plasma membrane to the cytoplasm of infected cells as nodules mature. Using phospho-peptide (Lb) specific antibodies, we found that serine phosphorylation triggers this translocation. Furthermore, FIHC in conjunction with immunoprecipitation followed by immunoblotting with phospho- and non-phospho-peptide specific antibodies demonstrated that the non-phosphorylated form is detectable as early as 9 dpi, whereas the phosphorylated forms were first detected at 11 dpi and progressively accumulated during nodule maturation. This spatio-temporal transition coincides with increasing rhizobial colonization and is accompanied by a decline in the non-phosphorylated pool. Therefore, the increased cytoplasmic pool of phosphorylated Lb, which exhibits reduced oxygen sequestration capacity, likely functions in promoting oxygen transport to sustain elevated rhizobial respiration. Together, these findings demonstrate that site-specific serine phosphorylation represents one of the key regulatory mechanisms linking Lb localization dynamics with progression of rhizobial infection, thereby contributing to the maintenance of oxygen homeostasis during SNF.

|
Scooped by
Jean-Michel Ané
May 1, 11:55 AM
|
As the global population continues to rise and resources become increasingly limited, hunger remains a critical challenge, projected to worsen by 2050. Ensuring a sustainable and sufficient food supply requires innovative solutions, with microbiology playing a key role in enhancing food production, quality, safety, and nutrition, contributing to the United Nations Zero Hunger goal. Microorganisms are essential to agriculture, food processing, and health, offering ecofriendly alternatives to chemical-intensive methods. Beneficial microbes such as nitrogen-fixing bacteria, mycorrhizal fungi, and phosphate-solubilizing bacteria improve soil fertility, increasing crop yield and sustainability. Biopesticides and biofertilizers reduce reliance on synthetic chemicals, promoting healthier ecosystems. This chapter explores the diversity and applications of microorganisms in agriculture, food, and health, emphasizing their immense potential in solving global food challenges. By harnessing microbial breakthroughs, we can develop sustainable solutions to combat hunger and create a resilient global food system.

|
Scooped by
Jean-Michel Ané
April 28, 11:37 AM
|
Root nodule symbiosis and pathogen fungi infection represent contrasting biotic interactions that affect plant growth and demand precise cellular reprogramming in Phaseolus vulgaris. Inoculation with Rhizobium tropici promoted progressive nodule organogenesis, reaching their maximum nodule number and diameter at 21 and 30 days post-inoculation, respectively. This developmental progression was accompanied by significant reductions in plant height, leaf and root weight, alongside an increase in photosynthetic pigment content, consistent with the energetic requirements of symbiosis establishment. In contrast, infection by the necrotrophic fungus Sclerotinia sclerotiorum caused extensive foliar necrosis and high infection disease, confirming the susceptibility of P. vulgaris. To explore the molecular regulators associated with these processes, 11 ROP (Rho of Plants) GTPase genes were characterized in the P. vulgaris genome based on the Ras superfamily domain (PF00071) and the Rho-specific motif (PS51420). Structural and phylogenetic analyses grouped PvROPs into five conserved subfamilies, while promoter inspection revealed cis-regulatory elements associated with nodulation (OSE1ROOTNODULE and OSE2ROOTNODULE) and immune signaling (WRKY710S), suggesting context-dependent transcriptional control. Expression analyses demonstrated that PvROPs respond dynamically to both interactions. During early symbiosis, PvROP5, PvROP8, and PvROP10 were significantly upregulated at 7 dpi, indicating roles in infection thread formation and initial nodulation signaling, whereas PvROP1, PvROP7, and PvROP9 were upregulated at 21 dpi, likely contributing to nodule maturation and functionality. Conversely, PvROP1, PvROP3, PvROP7, and PvROP10 were strongly induced in leaves infected with S. sclerotiorum, suggesting a potential role in the plant defense response. Our results provide valuable insight into the role of PvROPs to understand how legumes balance both pathogenic and symbiotic interactions in P. vulgaris.

|
Scooped by
Jean-Michel Ané
April 28, 11:30 AM
|
Transcription factors play essential roles modulating gene expression during plant development and the adaptation to environmental cues through the control of morphogenetic programs. In the root nodule symbiosis between legumes and rhizobia, two coordinated morphogenetic programs are activated by the perception of bacterial signals: the organogenesis of the nodule, a lateral root organ specialized in nitrogen fixation, and the infection process that allows the bacteria to colonize the nodule. These programs are influenced by the action of phytohormones, mainly auxin, cytokinin, ethylene, gibberellin, and brassinosteroid, which act modulating the activity of different families of transcription factors. In the past years, significant advancements have been made in understanding how transcription factors of the NIN (Nodule Inception), GRAS (GIBBERELLIN-ACID INSENSITIVE (GAI), REPRESSOR of GA1 (RGA), and SCARECROW (SCR)), ERF (Ethylene Response Factor), ARF (Auxin Response Factor), LBD (Lateral Organ Boundaries Domain), and SHI/STY (SHORT INTERNODES/STYLISH) families function at different developmental stages of bacterial infection and nodule formation and differentiation. Here, we review recent advances of this hormonal-mediated modulation of transcription factors with key roles in the root nodule symbiosis and their evolutionary origin from other developmental programs, as well as their post-transcriptional regulation by small RNAs. We also provide a perspective on how epigenomic approaches can shed light on how these transcription factors influence chromatin remodeling at loci containing key symbiotic genes.

|
Scooped by
Jean-Michel Ané
April 26, 4:50 PM
|
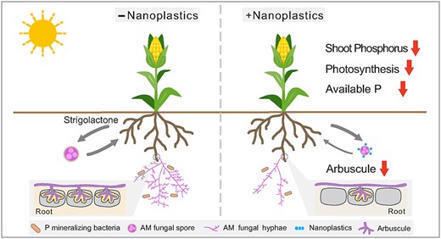
More than 80% of land plants form symbiotic relationships with arbuscular mycorrhizal (AM) fungi for nutrient uptake. As emerging soil pollutants, nanoplastics (NPs) accumulate in both crop and AM fungal tissue, posing non-negligible toxicity and health risks. However, whether and how NPs can impact plant-AM fungal partnership throughout the symbiotic process remains poorly understood. Here, using axenic root-fungal culture, fluorescence NP tracking, and real-time symbiotic signal monitoring, we show that during pre-colonization phase, NPs reduced spore germination rates (−48%) due to the NP accumulation on spore surface, hindering symbiotic signal perception. During the colonization phase, NPs entered fungal cells, disrupted organelles, and accelerated hyphal senescence, consequently reducing hyphal branching length (−22%) and secondary spore production (−32%). In real-world soil, inoculation of secondary spores (reproduced under NPs) formed fewer arbuscule structures (−46%) within maize roots with reduced carbon allocation to AM fungus, leading to lower hyphal length density (HLD) (−24%). During the post-colonization phase, lower HLD impaired the well-known function of phosphorus (P) mineralization by hyphae-enriched bacteria, reduced soil available P (−5.7%) and maize shoot P (−20%), eventually resulting in compromised plant performance. Our findings reveal an integrated yet largely underexplored mechanism of how NPs hinder plant performance by disrupting the dynamic relationship between plants and their symbiotic partners. In a broader context, understanding the alteration of plant-microbial interaction (rather than separately) under emerging stress can provide ecologically relevant implications for sustaining agricultural and terrestrial ecosystems.

|
Scooped by
Jean-Michel Ané
April 25, 4:47 PM
|
Mutualism in the symbiosis between arbuscular mycorrhizal fungi and plants is based upon the exchange of carbon for soil minerals, with phosphate being of central importance. The exchange of nutrients occurs when the fungus transiently colonises root cells, producing hyphal structures called arbuscules. The movement of phosphate from fungus to plant is well established, however its coordination and regulation at the ephemeral arbuscules remains elusive. Here, non-invasive imaging captures the complete growth and collapse of the arbuscules in unprecedented resolution, revealing heterogeneity in arbuscule development. Tracking the dynamics of rice PHosphate Transporter 1;11 (OsPHT1;11/ PT11) as a proxy for symbiotic phosphate transport shows consistent localisation across diverse arbuscules. However, we uncover phosphate-responsive variability in PT11 abundance, representing an essential, cellular-level layer of nutrient regulation. Such plasticity in arbuscule phosphate uptake capacity evidences uncoupling of arbuscule presence and arbuscule function, thereby demonstrating that arbuscules are not identical units of nutrient exchange.

|
Scooped by
Jean-Michel Ané
April 24, 5:05 PM
|
The search for life now extends beyond the traditional habitable zone to include the icy moons of Jupiter and Saturn. These moons feature ice-covered surfaces overlying substantial oceans formed primarily of liquid water and other potential constituents, such as ammonia. On Earth, ammonia supports biochemistry at low concentrations by providing nitrogen but becomes disruptive at higher concentrations. Ammonia could therefore influence the habitability of extraterrestrial oceans, yet this topic has received limited attention in the literature. This review synthesises current research on ammonia in Saturn’s icy moons, Enceladus and Titan, and its effects on terrestrial life. We summarize the celestial incorporation, speciation, and phase behaviour of ammonia and review data on its occurrence and concentration in icy moon oceans. We examine the role of ammonia in prebiotic chemistry, biochemistry, and toxicity. Focusing on bacteria, we compare known survival limits in ammonia to estimated ammonia concentrations on Enceladus and Titan. We find that bacterial survival limits exceed concentrations estimated on Enceladus, but are below those estimated on Titan, and propose that ammonia measurements are crucial for assessing extraterrestrial habitability. Finally, we highlight outstanding knowledge gaps and challenges that influence our understanding of how ammonia shapes the potential for life beyond Earth.

|
Scooped by
Jean-Michel Ané
April 21, 10:12 AM
|
Root exudation mediates the delivery of plant primary and secondary metabolites into soil, where they regulate plant–microbe interactions and terrestrial carbon cycling. Conventional exudate analyses quantify total root-released carbon yet obscure the spatial origin and rhizosphere influence of individual compounds. Here, we develop a rhizobacterial biosensor platform, named Suc-MAPP, to map local exudate profiles along the surface of colonized root tissues. Focusing on sucrose, we engineered sfGFP-based, sucrose-responsive gene circuits in Pseudomonas putida KT2440 for live imaging of exudate concentrations in the micromolar range. These biosensors reveal spatially structured sucrose exudation patterns across eudicots and monocots and implicate photoassimilated source–sink dynamics as a major determinant. We further apply this platform to phenotype exudation modulated by synthetic gene circuitry in Arabidopsis thaliana, identifying genetic design rules for graded sucrose release and quantifying how engineered export sculpts rhizosphere assembly of a defined bacterial community. Together, these results establish programmable rhizobacterial biosensors as tools to spatially resolve plant–environment carbon exchange in situ and provide a framework for extending this approach to diverse exudate targets.
|

|
Scooped by
Jean-Michel Ané
Today, 2:59 PM
|
While nitrogen fertilizers are widely used in agricultural production, their application incurs significant environmental and energetic costs. In contrast, some crops are less dependent on these fertilizers because they engage in symbioses with rhizobia, nitrogen-fixing bacteria provide ammonium to the plant in exchange for carbon. However, the carbon cost associated with nitrogen fixation can negatively impact crop yields. Improving the efficiency of this metabolic process could alleviate this impact on crop productivity. Mathematical models can help us quantitatively explore metabolic behavior and identify potential targets for metabolic engineering. In this work, we developed a kinetic model of determinate root nodule metabolism, where this symbiotic exchange of carbon from the plant and nitrogen from the bacteria occurs.
We used this model to evaluate how the predicted metabolic behavior differs between inefficient and efficient nodules, and to identify potential engineering targets for improving nitrogen fixation efficiency and rate. We show that the enzymes phosphoenolpyruvate carboxylase and pyruvate kinase have significant influence on the predicted rate and efficiency of nitrogen fixation, especially when their expression is varied in combination with oxidative Pentose Phosphate Pathway enzymes like glucose-6-phosphate dehydrogenase and 6-phosphogluconolactonase. The model predicts that pairing a 3-fold decrease in glucose-6-phosphate dehydrogenase activity along with either a 3-fold increase in phosphoenolpyruvate carboxylase activity or decrease in pyruvate kinase activity could increase nitrogen fixation rate by 5.51% while improving nitrogen fixation efficiency by 7.74%.

|
Scooped by
Jean-Michel Ané
Today, 2:51 PM
|
Quorum-sensing (QS) systems based on acyl-homoserine lactones (AHLs) regulate gene expression in response to cell density in many bacteria, including Rhizobium. These systems, typically composed of LuxI-like synthases and LuxR-like regulators, control processes such as plasmid conjugation, biofilm formation, and plant interactions. However, their evolutionary dynamics and genomic distribution in Rhizobium remain poorly understood. We analyzed 142 complete Rhizobium genomes using comparative genomics, phylogenetic reconstruction, and genomic context analysis. LuxI/LuxR homologs were identified based on sequence similarity and Pfam domain architecture, and their genomic contexts were examined. Phylogenetic relationships and coevolution between LuxI/LuxR pairs were assessed using cophylogenetic approaches. QS systems showed a highly heterogeneous distribution across Rhizobium genomes: some strains lacked canonical systems, whereas others encoded one or multiple systems in chromosomes and/or plasmids. Chromosomal QS systems were associated with multiple distinct genomic contexts, supporting at least seven independent acquisition events. In contrast, plasmid-encoded systems exhibited substantially greater diversity in both sequence and genomic organization. Phylogenetic and comparative analyses revealed dynamic gains and losses of QS systems, variable coevolution among LuxI/LuxR pairs, and evidence of partner recruitment. Notably, plasmids appear to act as major reservoirs of QS systems and likely sources of their transfer to chromosomes. These findings indicate that QS systems in Rhizobium evolve through a combination of horizontal gene transfer, genomic rearrangement, and differential retention across replicons. The higher diversity and mobility of plasmid-encoded systems highlight their central role in shaping QS evolution and functional innovation. Overall, this study provides a comprehensive framework for understanding the diversification and evolutionary trajectories of QS systems in complex multipartite bacterial genomes.

|
Scooped by
Jean-Michel Ané
Today, 2:42 PM
|
The relationship between plants and microorganisms has ancient roots. However, only recently we have begun to appreciate the true complexity of the holobiont and its intricate interactions. Fungi, bacteria and viruses residing within or around plants play a crucial role in enhancing their resilience to biotic and abiotic stressful factors. The impact of these interactions is not limited to stress resilience but are strictly correlated with the plant’s life cycle. The recognition of specific effects on plant growth and resilience has driven research towards utilizing microorganisms as sustainable solutions for agricultural challenges. The application of selected microorganisms, combined with the development of SynCom consortia, marks the beginning of a promising strategy for achieving sustainability in agriculture. This chapter explores the hidden world of plant-associated microorganisms, emphasizing their diverse beneficial impacts and providing evidence to support their potential applications in modern agriculture.

|
Scooped by
Jean-Michel Ané
May 8, 10:16 AM
|
Arbuscular mycorrhizal fungi (AMF) represent a key biological strategy for enhancing agricultural resilience under extreme climatic events such as drought

|
Scooped by
Jean-Michel Ané
May 5, 1:47 PM
|
Improving the effectiveness of microbial inoculants for soybean is essential to enhance biological nitrogen fixation and reduce fertilizer dependence; however, inoculated Bradyrhizobium strains frequently display inconsistent field performance. Inoculation is usually carried out with single-strain formulations, overlooking the possible influence of the native soil microbiota on nodulation success. This limitation may be addressed by formulating inoculants with consortia that include selected members of the soil microbiota. To this end, a synthetic microbial community (SynCom) was developed through a host-mediated microbiome engineering approach guided by leaf chlorophyll content as a rapid, non-destructive plant trait. The experiment was initiated by inoculating soybean plants with a consortium of 9 Bradyrhizobium spp. and 14 non-rhizobial soil isolates. Across eight consecutive selection rounds under gnotobiotic conditions, rhizosphere communities associated with superior plant performance were pooled and propagated. Recurrent selection induced significant shifts in community composition, consistently favoring Bradyrhizobium diazoefficiens as the dominant nodulating member and enriching taxa from Pseudomonadales, Burkholderiales, and Sphingomonadales. Sequencing-based profiling and network analysis suggested the emergence of a cohesive and functionally enriched community, with increased potential for nitrogen transformations and organic matter turnover. When evaluated in non-sterile soil, the SynCom derived from the sixth selection round increased nodule number and biomass relative to an uninoculated control and a commercial inoculant strain. These results suggest that plant-guided selection can steer rhizosphere community assembly toward beneficial configurations and support the development of improved soybean bioinoculants.

|
Scooped by
Jean-Michel Ané
May 4, 3:56 PM
|
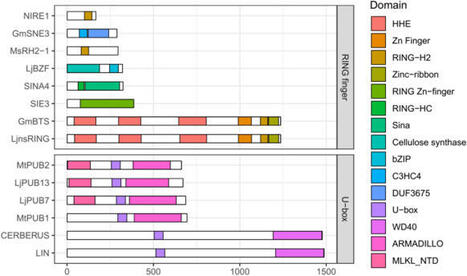
Symbiotic nitrogen fixation (SNF) by legumes is essential for sustainable agriculture, providing plant-available nitrogen while reducing reliance on synthetic fertilizers. The establishment of legume-rhizobium symbiosis requires tightly regulated host signaling to coordinate rhizobia infection, nodule development, and nitrogen fixation, while preventing excessive colonization or immune activation. Accumulating evidence indicates that ubiquitination, mediated by E1, E2, E3 ubiquitin ligases and deubiquitinating enzymes, plays a central role in controlling multiple stages of this process. In this review, we summarize current knowledge on ubiquitination-mediated regulation of symbiotic nitrogen fixation, with a focus on early symbiotic signaling and nodule development. We highlight key E3 ligases that modulate Nod factor receptor homeostasis, receptor-associated kinases, transcription factors, and infection thread growth, and discuss how ubiquitination interfaces with nutrient and stress signaling pathways. Finally, we outline key knowledge gaps and discuss the potential of manipulating ubiquitination pathways to improve nodulation efficiency and nitrogen use efficiency in crops.

|
Scooped by
Jean-Michel Ané
April 30, 5:32 PM
|
The rhizobia-legume symbiosis involves nodulation and nitrogen fixation, which are vital bioprocesses for the sustenance of agriculture, land and ecosystem, and the global nitrogen cycle. Legume plants and rhizobia interact intricately, resulting in the formation of specialized structures known as root nodules. Root nodules enhance soil fertility and promote more vigorous plant growth by converting atmospheric nitrogen into a form that plants can utilize. This symbiosis is regulated by complex and finely tuned molecular pathways that involve signal exchange during host recognition, nodulation, and nitrogen fixation, ensuring the successful initiation and maintenance of the symbiosis. This interaction comes with specific difficulties. This review explores the intricate dynamics of the symbiotic relationship between legumes and rhizobia. It involves the molecular, genetic, and ecological factors that govern this connection, providing a thorough understanding of its advantages and disadvantages. By examining these facets, this review aims to illuminate rhizobia’s potential to enhance legume yields and promote more environmentally conscious farming practices, while acknowledging the obstacles that must be overcome to maximize this mutualistic association.

|
Scooped by
Jean-Michel Ané
April 28, 11:32 AM
|
Plant–microbe interactions shape key agronomic traits, yet breeding strategies to harness them remain unclear.
We propose distinguishing the varying population, which supplies heritable variation, from the carrying population, where the selected property is expressed.
In traditional breeding, the varying and carrying populations coincide within a single organism, whereas in novel strategies, they are distributed among plants and microorganisms, with one partner varying and the other carrying the selected property.
This distinction defines holobiont breeding as a strategy in which both plants and microorganisms are combined to form a varying population, enabling the selection of interspecific indirect genetic effects.
Our framework builds on the replicator–interactor distinction by localizing these roles within plant–microorganism associations, thereby making it operational for breeding program design.

|
Scooped by
Jean-Michel Ané
April 26, 4:52 PM
|
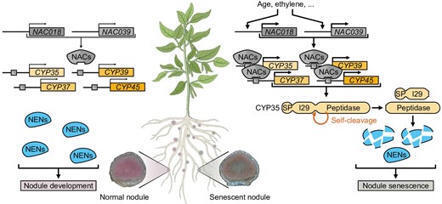
In legumes, symbiotic root nodules undergo senescence in response to developmental or environmental cues. This process determines the maintenance and nitrogen-fixing capacity of the root nodules, but the molecular mechanisms underlying its initiation are poorly understood. The cysteine protease CYP35 is a positive regulator of nodule senescence in soybean (Glycine max), but its substrates remain unknown. Here, we demonstrate that CYP35 promotes nodule senescence by cleaving a subset of Nodule-Enriched Nodulin proteins (NENs). Sequence and phylogenetic analyses indicate that CYP35 is a cathepsin L-like cysteine protease, with Cys149 as a key catalytic residue. CYP35 physically interacts with a distinct subfamily of eight NENs, NEN1–8. Soybean quadruple and quintuple nen mutants obtained by multiplex gene editing develop nodules with accelerated senescence and reduced nitrogenase activity, whereas over-expression of NEN2 or NEN5 delays senescence and enhances nodule function. CYP35 proteolyzes NEN2, NEN5, NEN6, and NEN7 in vitro and cleaves NEN2 in vivo in a Cys149-dependent manner. Our findings establish a direct molecular link between cysteine protease–mediated Nodulin cleavage and the onset of nodule senescence in soybean, providing insights into the regulation of nodule lifespan and nitrogen fixation.

|
Scooped by
Jean-Michel Ané
April 26, 4:43 PM
|
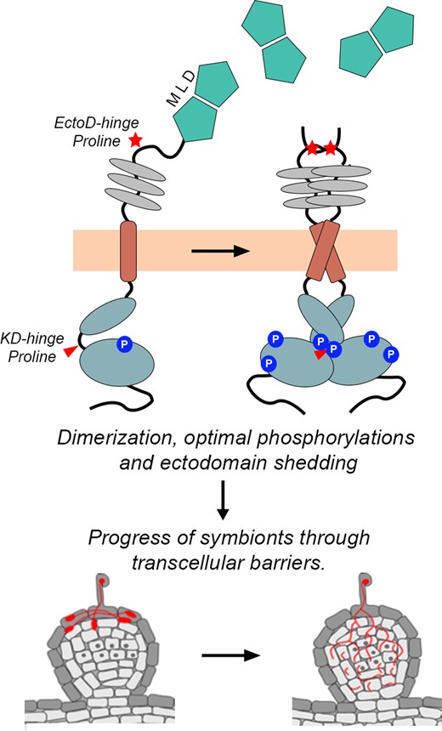
Symbiosis receptor kinase (SYMRK), a malectin-like-domain/leucine-rich-repeat receptor-like-kinase (MLD-LRR-RLK), is the upstream-most component in the common-symbiosis-signaling-pathway. We highlight 2 proline residues that were distinctly acquired by SYMRK orthologs in its hinge regions to constitute a signaling module for allowing progress of symbionts across transcellular barriers during rhizobia-legume symbiosis. Within the ectodomain hinge (EctoD-hinge), all MLD-LRR-RLKs have a conserved W1xnGDPCxnW2×4C motif, where SYMRK orthologs within legumes have a distinct signature with a proline preceding W2 enabling cleavage of SYMRK ectodomain for releasing MLD. Within the kinase hinge (KD-hinge) at gatekeeper + 1 position, a conserved glutamate in MLD-LRR-RLKs is replaced by proline in all SYMRK orthologs that enabled its dual-specific kinase activity for ensuring ectodomain cleavage. Substitution of either proline restricted cortical progression of symbionts, forming infection patches in the nodule apex without affecting epidermal invasion and nodule organogenesis. This halt was entirely overcome by ectopic expression of free MLD, demonstrating the released MLD has an active role in progress of symbionts. Overall, we show that conservation of distinct prolines in hinge regions of SYMRK orthologs in legumes generates a signaling module involving dimerization and optimal phosphorylation of SYMRK for releasing MLD as an active transducer of symbiosis signaling.

|
Scooped by
Jean-Michel Ané
April 25, 12:36 PM
|
Legume-rhizobial symbiosis, involving the intracellular accommodation of rhizobia within host cells for nitrogen fixation, represents a unique model among plant-microbe interactions. While this symbiosis requires sophisticated regulatory networks, direct host control over symbiont proteins remains largely elusive. Here, we demonstrate that soybean Nodulation Receptor Kinase (GmNORK) interacts with and directly phosphorylates the Sinorhizobium fredii HH103 effector NopZ. This host-mediated phosphorylation promotes NopZ nuclear/perinuclear enrichment and is associated with enhanced NopZ interaction with the nucleoporin GmNENA at the nuclear envelope/nuclear pore complex, establishing a key signaling step that links membrane-proximal symbiotic signaling to the nuclear pore. Expression of phosphomimetic NopZ variants enhances soybean nodulation, whereas GmNENA-silencing in soybean roots reduces nodule numbers. Our findings reveal a previously unknown model for plant-microbe interactions, where the direct phosphorylation of a bacterial effector by a host receptor kinase provides an essential regulatory mechanism to direct effector localization and promote the symbiotic program.

|
Scooped by
Jean-Michel Ané
April 21, 10:21 AM
|
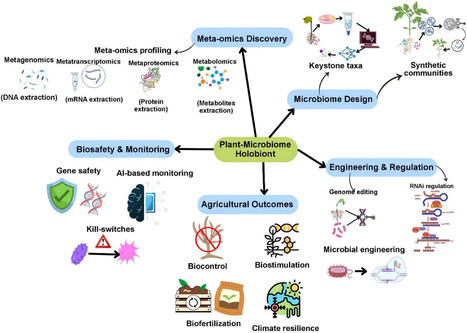
Plants exist as complex holobionts, whose health and productivity depend on dynamic interactions with diverse microbial partners. This chapter explores the ecological, molecular, and biotechnological dimensions of plant-microbe relationships, tracing the continuum from natural symbioses to their translation into microbe-based agricultural applications. It examines how plants actively structure their microbial communities across distinct yet interconnected compartments, such as the rhizosphere, phyllosphere, and endosphere, and how these microorganisms in turn modulate plant physiology, nutrient acquisition, and defense. Moving beyond the study of individual strains, we discuss the growing relevance of community-level approaches and synthetic microbial consortia as frameworks to decipher and replicate the ecological principles that sustain beneficial interactions. Finally, this chapter reflects on how the integration of ecological knowledge with emerging technologies such as synthetic biology, systems modeling, and artificial intelligence offers new opportunities to design resilient, predictable, and sustainable microbe-based solutions. By learning from these naturally optimized alliances, we can guide the transition to more adaptive and sustainable agricultural systems while sustaining plant health and productivity.
|





 Your new post is loading...
Your new post is loading...














Awesome work in this preprint