 Your new post is loading...
 Your new post is loading...

|
Scooped by
Jean-Michel Ané
June 2, 12:46 PM
|
The stabilization of rhizobia in specialized organelles, called symbiosomes, is an evolutionary hallmark for maintaining high rates of nitrogen fixation in legumes. This is achieved by releasing thousands of bacteria from nodular infection threads within in a single plant cell, which poses a great challenge for the host to keep control over these differentiated bacteria. Considering the importance of proteolytic degradation of antimicrobial proteins or generation of symbiosis-promoting peptides, proteolytic activity may represent a key regulatory system. Indeed, we identified the Medicago truncatula subtilisin-like protease (subtilase, SBT) 12a acting as a novel regulator of symbiosome functionality and maintenance. Loss-of-function mutations in SBT12a led to severe symbiotic defects with nodules of sbt12a being characterized by high level induction of defense/senescence-related genes. Using untargeted proteomic High-efficiency Undecanal-based N-Termini EnRichment (HUNTER) we identified and individually confirmed specific SBT12a target proteins that are involved in plant defense responses and symbiosome maintenance. This positions SBT12a as a central host factor controlling symbiosome performance.

|
Scooped by
Jean-Michel Ané
May 21, 11:40 AM
|
Arbuscular mycorrhizal fungi (AMF) have facilitated the colonization of land by plants some 470 million years ago. The vast majority of land plants have maintained a symbiotic association with these fungi to facilitate the uptake of mineral nutrients, such as phosphorus, at the cost of photosynthates delivered to the fungus in the form of lipids and sugars. Despite their importance for plant nutrient status, plants can refuse AMF if soil nutrient conditions are such that it is less costly for the plants to take up the nutrients by themselves or if environmental conditions are not appropriate. Recently, Hong et al. (2026) revealed how multiple signaling pathways in rice converge on a transcription factor complex involving the GRAS transcription factors NSP1 and NSP2 to control AM colonization. In this spotlight we highlight recent insights into the molecular mechanisms that control the permissiveness of plants to allow AMF entry into their roots (Figure 1fig1).

|
Scooped by
Jean-Michel Ané
May 18, 6:28 PM
|
Nutrient enrichment alters the functioning of grassland ecosystems, but the community structure and functions of microbes associated with the hyphosphere of arbuscular mycorrhizal (AM) fungi under nitrogen (N) and phosphorus (P) amendments remain poorly understood. Using a compartmented microcosm system and 16S rRNA gene and metagenomic sequencing, we studied the effects of AM fungal hyphae on soil microbial community composition, carbohydrate metabolism, and P cycling in four soils subjected to long-term N and/or P amendments. In long-term N-amended soils, AM fungal hyphae markedly altered the composition of the microbial community, improved P uptake and transport, and enriched genes associated with amino acid and secondary metabolite metabolism. Conversely, in long-term P-amended soils, the hyphae significantly reduced the concentration of available P in the soil and decreased the relative abundance of glucosyl transferases. Under combined NP amendments, the hyphae also induced significant changes in the composition of the microbial community and decreased the concentration of available P in the soil. In addition, AM fungal hyphae selectively modulated the abundance of specific genes involved in carbohydrate metabolism and P cycling, with variable effects depending on the soil. These results show that long-term nutrient amendments reshape AM fungal regulation of hyphosphere microbial communities and functions.

|
Scooped by
Jean-Michel Ané
May 17, 4:11 PM
|
Intercropping is widely practiced on sloping red-soil farmland in Southwest China and helps mitigate soil erosion. However, the mechanisms by which intercropping influences soil aggregate stability remain poorly understood. Based on an 11-year (2013–2024) fixed-location experiment in Yunnan Province, this study examined the effects of maize monoculture (MM), soybean monoculture (SS), and maize–soybean intercropping (IM/IS) on soil aggregate stability and explored how rhizospheric arbuscular mycorrhizal fungi (AMF) dynamics were associated with aggregate stability under different cropping systems.
Results indicated that: (1) compared with monocultures, intercropping significantly increased AMF colonization rates and hyphal density by 8.3–34.9% (P < 0.01) and 0.51–1.20-fold (P < 0.01), respectively, and increased the abundance of Glomus in maize rhizospheres. (2) Intercropping significantly increased glomalin-related soil protein (GRSP), with the increase in easily extractable GRSP (EE-GRSP; 13.11–29.91%) exceeding that of total GRSP (T-GRSP; 2.51–6.85%). GRSP contents were significantly higher at tasseling than at maturity (P < 0.05). (3) Compared with monocultures, intercropping significantly increased xylanase (XYL), N-acetyl-β-D-glucosaminidase (NAG), and acid phosphatase (PHOS) activities by 43.38–61.84%, 39.28–96.03%, and 42.57–101%, respectively (P < 0.05). These biochemical improvements were accompanied by improved soil structure, with >2 mm macroaggregates and mean weight diameter (MWD) increasing by 24.37–38.84% and 11.24–25.26%, respectively (P < 0.05). Structural equation modeling further indicated that hyphal density may indirectly improve soil aggregate stability by increasing EE-GRSP content and hydrolase activity. Overall, maize–soybean intercropping enhanced aggregate formation and stability through AMF-related GRSP and hydrolase pathways, supporting improved acidic red-soil structure on sloping farmland.

|
Scooped by
Jean-Michel Ané
May 17, 3:30 PM
|
Dramatic cytoskeleton modifications are involved in establishing mutualistic bacterial and fungal associations with plants, as new specialized symbiotic structures are formed. A recent study in Science1 explores actin remodeling during the inception of these interactions, potentially creating opportunities to increase mycorrhizal and nodulation efficiency and reduce our need for fertilizers.

|
Scooped by
Jean-Michel Ané
May 17, 3:15 PM
|
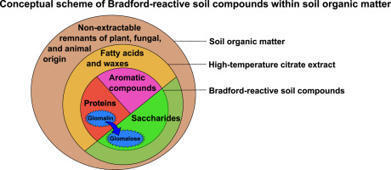
This review critically examines the chemical nature of Glomalin-related soil proteins (GRSP), the biases introduced by non-mycorrhizal sources, and emphasizes the need to avoid misinterpreting GRSP as an arbuscular mycorrhizal fungi (AMF) marker. GRSP have been widely used in agricultural science as indicators of soil health and as a quantitative indicator of the accumulation of decomposed biomass of AMF in soil. GRSP is typically extracted by repeated autoclaving of soil in a citrate buffer, followed by protein quantification using the Bradford assay. However, recent discussions on the composition, structure, and interpretation of GRSP raise concerns about its specificity as an AMF marker, and we have serious doubts about the protein dominance of this mixture. A key limitation is that conventionally measured GRSP concentrations also correlate with plant litter decomposition and organic matter inputs, indicating that GRSP reflects general organic accumulation rather than AMF-specific processes. This underscores the need for more direct and representative measurements of mycorrhizal interactions. To address this issue, we propose renaming GRSP as Bradford-reactive soil compounds (BRSC), a term that more accurately reflects the chemical heterogeneity of the fraction and avoids implying any specific association with AMF.

|
Scooped by
Jean-Michel Ané
May 14, 6:24 PM
|
Genome-based taxonomy offers a powerful means to resolve long-standing ambiguities in the classifications of agrobacteria and rhizobia, two bacterial groups with major ecological and agricultural significance. We applied a robust phylogenomic framework to genomes of the families Bartonellaceae and Rhizobiaceae to comprehensively reassess evolutionary relationships. Species trees were constructed using nucleotide sequences of 92 conserved genes and amino acid sequences of 120 ubiquitous proteins, clarifying relationships that were previously obscured by marker-limited historical classifications. These analyses demonstrated several instances of taxonomic inconsistencies across genera, most notably within Mesorhizobium, which forms a paraphyletic assemblage spanning multiple divergent lineages. These findings were further reinforced by the genome-wide similarity metric, average amino acid identity (AAI), which supported reclassifications at both genus and species levels, reflecting natural discontinuities between lineages. We propose the reclassification of eight species and five new genera and the validation of their names primarily under the Code of Nomenclature of Prokaryotes Described from Sequence Data (SeqCode). As genome-based resources expand, and with the availability of new nomenclatural frameworks such as the SeqCode, genome-informed taxonomy offers a powerful approach to delineate taxa into biologically meaningful groups that can be formally recognised. The revised taxonomy presented here brings greater coherence to the systematics of agrobacteria and rhizobia and provides a framework for future evolutionary investigations into these agriculturally and ecologically significant bacterial groups.

|
Scooped by
Jean-Michel Ané
May 12, 8:45 PM
|
•
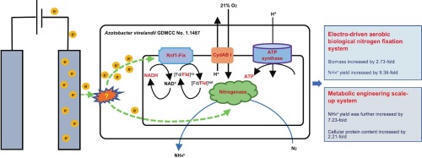
First aerobic (21% O2) electro-driven biological nitrogen fixation system.
•
Exogenous electricity boosts NH4+ and biomass via metabolic reprogramming.
•
Dual electron transfer: main Rnf1-NifF pathway plus a newly discovered bypass.
•
Metabolic engineering for enhanced electro-driven protein and NH4+ production.

|
Scooped by
Jean-Michel Ané
May 12, 6:33 PM
|
Intracellular infection by beneficial microorganisms enables endosymbiosis and remains a challenge in synthetic biology. A new study by Qiao et al. reveals that a formin orchestrates both rhizobial infection in legumes and mycorrhization in diverse plants. Harnessing this mechanism could enable the engineering of microbial entry into crops for sustainable nutrient acquisition.

|
Scooped by
Jean-Michel Ané
May 9, 3:00 PM
|
In root nodule symbiosis, symbiosome compartments accommodate nitrogen-fixing rhizobia inside the plant cell. Differentiated into bacteroids, the rhizobia are surrounded by a peribacteroid space and a plant-derived peribacteroid membrane, which separates them from the plant cytoplasm but allows signal and nutrient exchange between host and microbe. The morphological features of symbiosomes are primarily determined by ultrastructural single focal plane imaging, with limited information about spatial details. This study combines 2D and 3D imaging, using transmission electron microscopy and focused ion beam scanning electron microscopy as complementary techniques to analyse the symbiosome ultrastructure and organisation in Lotus japonicus wild-type plants. The 3D model of a mature colonised root nodule cell region demonstrates a dense, puzzle-like arrangement of symbiosomes relative to one another and adjacent plant organelles. The symbiosome shape and size depends on the orientation and number of bacteroids within the compartment and features connective tubular structures. Furthermore, vesicular structures, some likely of bacterial origin, were present at the interface. The study presents a multi-angled analysis of symbiosome-related structures, highlighting their volumes, spatial distribution, and pronounced compactness. Interface associated vesicles, protrusions and connective structures hint towards a dynamic and flexible system that contributes to the plant-microbe crosstalk.

|
Scooped by
Jean-Michel Ané
May 9, 2:52 PM
|
Development of arbuscular mycorrhiza (AM), a symbiosis between plants and beneficial Glomeromycotan fungi, is largely under plant control. Several genes, required for AM development, are proposed to be regulated by the karrikin signalling module, comprising the alpha/beta hydrolase receptor KARRIKIN INSENSITIVE 2 (KAI2), the F-box protein MORE AXILLARY GROWTH2 (MAX2) and the transcriptional repressor SUPPRESSOR OF MAX2 1 (SMAX1), which is ubiquitylated for proteasomal degradation upon KAI2-ligand-induced binding to the KAI2-MAX2 complex. Rice and Brachypodium distachyon kai2 mutants are incapable of forming AM. Here, we show that in Lotus japonicus, Pisum sativum, and Nicotiana benthamiana, KAI2 only quantitatively affects AM development, indicating angiosperms vary in their requirement for KAI2-signalling to support AM. Comparative transcriptomics of L. japonicus and B. distachyon roots after treatment with fungal signalling molecules revealed some AM-relevant genes respond KAI2-independently in L. japonicus but not in B. distachyon. Consistently we obtained

|
Scooped by
Jean-Michel Ané
May 9, 2:43 PM
|
Nodules cystine-rich (NCR) peptides from legume nodules mediate bacteroid differentiation of Rhizobium sp. This study aimed to characterize NCR peptides of Phaseolus vulgaris, Trifolium repens, Medicago sativa, and Vicia faba, as relevant information on these legumes is still lacking. After exposing the root nodules of four leguminous plants to sonication, the peptides were separated according to their molecular weight by sodium dodecyl gel electrophoresis (SDS-PAGE). The NCR peptide was detected by high-performance liquid chromatography (HPLC), followed by the determination of the protein sequence and amino acid composition and their evolutionary history in terms of the phylogenetic tree by bioinformatics analysis. The results showed these four peptides had the same molecular weight of 60 kDa. The peaks of the curve varied with retention time depending on the legume from which the peptide was extracted. Bioinformatic analysis of the results showed that the amino acid substitutions varied in number and sequence for each species and that these peptides had three distinct wire, ribbon, and surface patterns within the structural modeling of their crystals. Side chains of amino acids in the loops and helices within the grid box may facilitate hydrophobic interactions via tyrosine, phenylalanine, or polar interaction sites via cysteine, histidine, and glutamine. These loops often serve as flexible regions to adapt to binding partners. In general, this study contributed to the characterization of NCR peptides that showed mismatches and differences in amino acid sequences and deletions of some amino acids at different locations depending on the type of legume from which the peptide was extracted.

|
Scooped by
Jean-Michel Ané
May 8, 10:21 AM
|
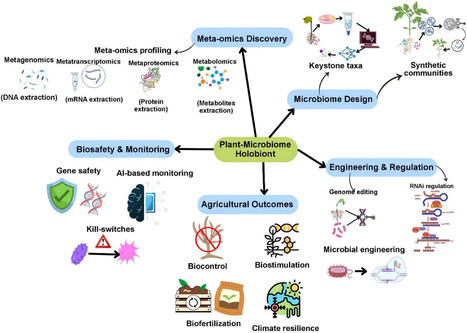
Plant-associated microbiomes are central to crop productivity, nutrient efficiency, and stress resilience, yet conventional microbiome manipulatio
|

|
Scooped by
Jean-Michel Ané
June 1, 6:21 PM
|
The advent of endosymbiosis underlies evolutionary innovation and ecosystem function. However, whether free-living partners tend to benefit or exploit each other during the early stages of novel endosymbiosis remains a dilemma. Rhizobia soil bacteria can initiate root nodules and fix nitrogen for host plants as endosymbionts due to genes carried on mobile genetic elements such as the symbiosis island (SI). We conjugated marked SIs into the genomes of non-nodulating strains, which was sufficient to generate de novo root nodule-forming endosymbionts. Most novel endosymbionts originated as commensals that incurred no detectable costs to host plants, in contrast to predictions of exploitation. In fact, a third of novel endosymbionts originated as nitrogen-fixing mutualists. Consistent with phylogenetic limits to transfer of mobile genetic element function, novel endosymbionts derived from more closely related SI donor and recipient strains showed greater nitrogen fixation. However, consistent with selection on the SI for broad horizontal transfer, we did not detect phylogenetic limits to SI transmission, and the SI was able to displace other genomic elements residing at its characteristic tRNA gene insertion site. We thus provide genetic, genomic, and functional evidence of how mobile genetic elements can potentiate and constrain major evolutionary transitions to expand bacterial niches, with cascading impacts on the fitness of host organisms.

|
Scooped by
Jean-Michel Ané
May 18, 6:44 PM
|
The trait‑based root economics space provides a fundamental framework for understanding plant belowground adaptation to environmental change, yet it typically omits root exudation that sustains plant resource acquisition and health. Results from a ~3500 km north‑south forest transect demonstrate that arbuscular mycorrhizal (AM) trees have higher root exudation rates than ectomycorrhizal (ECM) trees, with stronger increases under warmer and wetter climates. Root exudation rate is positively related to specific root length and negatively related to root diameter for both mycorrhizal types, but is related to tissue density and nitrogen content only in AM trees. These findings suggest that root exudation may represent an alternative collaboration strategy, facilitating organic nutrient acquisition more strongly for AM species and functioning as a relatively redundant nutrient acquisition mechanism for ECM species. Moreover, mycorrhizal type modulates how exudation rates relate to the “fast–slow” strategy. Overall, our findings refine the conceptual framework of the root economics space and enhance understanding of fine‑root strategies.

|
Scooped by
Jean-Michel Ané
May 17, 4:18 PM
|
Synthetic microbial communities (SynComs) are emerging as promising alternatives to single-strain inoculants in agriculture, offering greater functional robustness and environmental adaptability. However, transforming conceptual studies into engineerable and scalable agricultural practices remains challenging. In this opinion article, we synthesize current research on plant SynComs through a framework that moves from strain-centered assembly toward system-level design, linking the identification of truly stable coexisting communities in natural microbiomes to the elucidation of plant–microbe–soil interaction mechanisms, the development of dynamical models, and the integration of these models into platform-based design and production pipelines. We focus on recent advances that integrate generalized Lotka–Volterra and consumer-resource models with multi-omics data and other system-level constraints, with the aim of introducing model-driven concepts of SynCom design and promoting their large-scale application in agriculture.

|
Scooped by
Jean-Michel Ané
May 17, 4:06 PM
|
Tobacco Fusarium root rot is a widespread soil-borne root-stem disease affecting tobacco in China. Arbuscular mycorrhizal fungi (AMF) can enhance the disease resistance of host plants. A systematic study of tobacco's response to AMF under pathogen attack can lay the foundation for the application of AMF in alleviating tobacco root rot. This study aimed to clarify the effects of pre-inoculation with Claroideoglomus lamellosum (Cl) on tobacco growth, physiology, and root rot resistance, and to identify key AMF-responsive genes in tobacco leaves under Fusarium oxysporum (F) infection. Pot experiments with tobacco K326 included four treatments: control (CK), single Cl inoculation, single F inoculation, and dual Cl + F inoculation. Disease incidence, growth indices, and antioxidant enzyme activities were measured to assess Cl-induced resistance. Transcriptome analysis revealed key gene expression changes in response to Cl under F stress. Compared with CK, Cl formed a stable symbiosis with tobacco and promoted plant growth, as evidenced by increases in total biomass (77.71%), shoot biomass (83.5%), root biomass (107.71%), plant height (33.22%), root length (38.78%), stem diameter (18.67%), and effective leaf number (66.67%). Cl also enhanced leaf activities of superoxide dismutase (30.48%), peroxidase (71.81%), phenylalanine ammonia-lyase (169.25%), polyphenol oxidase (320.22%), and catalase (125.44%). Relative to pathogen treatment alone (F), the Cl + F combination reduced root rot incidence by 57.14% and disease index by 70.61%. In contrast, pathogen infection alone suppressed tobacco growth (6.65–27.64%) and decreased antioxidant enzyme activities (0.65–51.46%) compared with CK. Transcriptome analysis identified 20 050 differentially expressed genes (DEGs), with 2365 DEGs specifically responsive to AMF under pathogen challenge as key genes for AMF-induced resistance. GO and KEGG enrichment indicated that AMF enhanced resistance via plant-pathogen interaction and MAPK signaling pathways, while peroxisome, fatty acid elongation, and other key metabolic pathways also contributed significantly. Cl is an effective strain for enhancing tobacco resistance against root rot. Inoculation with Cl significantly promoted tobacco growth and increased physiological enzyme activities in tobacco leaves. At the transcriptomic level, it was confirmed that the effect of Cl inoculation on tobacco resistance against root rot involves multiple pathways and metabolic routes, forming a tripartite "signaling-membrane transport-metabolism" defense network model, thereby improving tobacco resistance to root rot.

|
Scooped by
Jean-Michel Ané
May 17, 3:23 PM
|
Calcium (Ca²⁺) functions as an essential intracellular second messenger in plants, mediating processes such as pathogen defence, stress adaptation and enzyme activation. Plants possess multiple families of calcium‑binding proteins, and among these, calcium/calmodulin‑dependent protein kinases (CCaMKs) remain among the least characterized. Also, their dual capacity to bind both Ca²⁺ and calmodulin makes them a subject of significant research interest. Usually, there is one CCaMK gene per plant genome, but when we cloned the full-length VaCCaMK cDNA sequence, we found several transcripts with missing exons, so it is possible that alternative splicing increases the overall diversity of CCaMK sequences. This study investigated the role of CCaMKs in Vitis amurensis Rupr. under abiotic stress conditions using grapevine cell cultures overexpressing the VaCCaMK1 gene and its alternatives VaCCaMK1-s1, -s2. Results demonstrated that VaCCaMK1‑overexpressing cultures did not exhibit increased tolerance to salt, osmotic, cold, or heat stress. Additionally, the content of secondary metabolites, as represented by stilbenes mainly produced by used grapevine cells, remained largely unchanged. However, a significant increase in both fresh and dry cell mass was observed compared with the control group: fresh biomass increased 1.1–1.7‑fold, and dry biomass 1.1–1.8‑fold. These findings indicate that VaCCaMK genes do not affect plant cell susceptibility to the tested abiotic stresses. VaCCaMK1-s1 or -s2 the overexpression had a similar but weaker effect on grape cells. Nevertheless, VaCCaMK appear to act as a positive regulator of cell growth and development in V. amurensis, suggesting their potential role in enhancing biomass accumulation.

|
Scooped by
Jean-Michel Ané
May 14, 6:26 PM
|
Long-term field experiments were established in 1969 at the gleyic fluvisol to assess the effects of soil management on soil organic matter an

|
Scooped by
Jean-Michel Ané
May 12, 8:46 PM
|
The uptake of nitrogen-fixing bacteria into living plant cells and the intracellular accommodation of arbuscular mycorrhiza (AM) fungi requires the plasma membrane-localised Symbiosis Receptor-like Kinase (SymRK). AM is widespread across terrestrial vascular plant lineages, while the nitrogen-fixing root nodule symbiosis (RNS) is restricted to one clade within the eurosids. This distribution led to the concept that SymRK was adopted during evolution to mediate RNS. Comparative analyses revealed that SymRK orthologs from the eurosid clade support RNS while SymRK from the phylogenetically distant species Solanum lycopersicum (tomato) does not. To dissect the molecular basis for this different functionality, we carried out complementation analyses of the Lotus japonicus symrk-3 mutant which is unable to form AM or RNS. Domains swap chimera from the tomato and L. japonicus SymRK orthologs revealed that the intracellular domain of L. japonicus SymRK is necessary and for cortical infection thread (IT) and symbiosome development at 21 days post inoculation. Notably, this signalling specificity could be overcome by ectopic expression of tomato SymRK, pointing to altered protein dosage as a potential determinant of function. Consistent with this idea, SINA family E3 ubiquitin ligases interacted with and ubiquitinylated L. japonicus SymRK, but not tomato SymRK. In yeast two hybrid analysis, the interaction of SymRK with SINA2 and SINA4 depended on the C-terminal intrinsically disordered tail region of L. japonicus SymRK. We conclude that the SymRK intracellular domain evolved interaction capabilities with SINA E3 ligases which correlates with its ability to support RNS.

|
Scooped by
Jean-Michel Ané
May 12, 8:45 PM
|
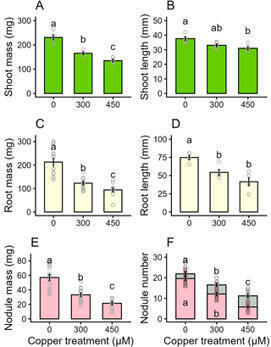
Copper can be a soil contaminant at concentrations that are toxic to plants, particularly because of copper-induced oxidative stress. The legume-rhizobia partnership that allows for biological nitrogen fixation is sensitive to oxidative stress, and this study investigates if copper acts directly on the machinery of nitrogen fixation, or indirectly via toxicity to the entire plant. When Lotus japonicus inoculated with its rhizobial partner Mesorhizobium loti was exposed to 300 or 450 µM of copper, biomass was reduced by 30–40% in the shoots, 40–55% in the roots, and 40–60% in the nodules relative to control. While concentrations of copper in shoots and roots increased in proportion to the amount of copper in the growth medium, concentrations of copper in nodules did not vary in response to copper treatment. Malondialdehyde, a marker of oxidative damage, in the nodule similarly did not vary with copper treatment. However, nitrogen fixation and ascorbate peroxidase activity decreased by 40–45% and 40–60%, respectively, which can be indicators of early nodule senescence. This would suggest that copper-induced reduction in nodule activity is not directly due to oxidative stress in the nodules; it is due to stress on the host plant that limits its ability to support its symbionts.

|
Scooped by
Jean-Michel Ané
May 11, 10:36 AM
|
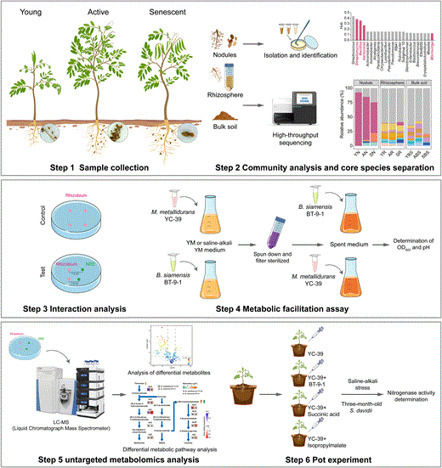
The root nodules formed by rhizobia and leguminous plants are specialized structures for nitrogen fixation. However, a large number of non-rhizobial endophytes also coexist within the nodules, and their contribution to nitrogen fixation under abiotic stress conditions remains unclear. Here, using the wild leguminous shrub Sophora davidii as model system, we identified an important NRE (Bacillus siamensis BT-9-1) by analyzing keystone taxa within the bacterial cooccurrence network of root nodules. This strain could improve the survival of Mesorhizobium metallidurans YC-39 under saline–alkali stress. A mechanistic investigation revealed that the expression of ilvA, ilvH, and ilvD was downregulated, and the contents of (2S)-isopropylmalate and succinic acid decreased in M. metallidurans YC-39 under saline–alkali conditions, whereas B. siamensis BT-9-1 presented increased accumulation of these metabolites. These findings indicate that B. siamensis BT-9-1 cross-feeds M. metallidurans YC-39 with these metabolites, rescuing the compromised branched-chain amino acid synthesis pathway and the tricarboxylic acid cycle in saline–alkali environments. Eventually, coinoculation with B. siamensis BT-9-1 and M. metallidurans YC-39, along with (2S)-isopropylmalate and succinic acid supplementation, increased nitrogenase activity of the symbionts. Our study reveals a novel mechanism by which non-rhizobial endophyte Bacillus species enhances the growth and nitrogen fixation efficiency of M. metallidurans under saline–alkali stress through the delivery of key metabolites.

|
Scooped by
Jean-Michel Ané
May 9, 2:59 PM
|
While nitrogen fertilizers are widely used in agricultural production, their application incurs significant environmental and energetic costs. In contrast, some crops are less dependent on these fertilizers because they engage in symbioses with rhizobia, nitrogen-fixing bacteria provide ammonium to the plant in exchange for carbon. However, the carbon cost associated with nitrogen fixation can negatively impact crop yields. Improving the efficiency of this metabolic process could alleviate this impact on crop productivity. Mathematical models can help us quantitatively explore metabolic behavior and identify potential targets for metabolic engineering. In this work, we developed a kinetic model of determinate root nodule metabolism, where this symbiotic exchange of carbon from the plant and nitrogen from the bacteria occurs.
We used this model to evaluate how the predicted metabolic behavior differs between inefficient and efficient nodules, and to identify potential engineering targets for improving nitrogen fixation efficiency and rate. We show that the enzymes phosphoenolpyruvate carboxylase and pyruvate kinase have significant influence on the predicted rate and efficiency of nitrogen fixation, especially when their expression is varied in combination with oxidative Pentose Phosphate Pathway enzymes like glucose-6-phosphate dehydrogenase and 6-phosphogluconolactonase. The model predicts that pairing a 3-fold decrease in glucose-6-phosphate dehydrogenase activity along with either a 3-fold increase in phosphoenolpyruvate carboxylase activity or decrease in pyruvate kinase activity could increase nitrogen fixation rate by 5.51% while improving nitrogen fixation efficiency by 7.74%.

|
Scooped by
Jean-Michel Ané
May 9, 2:51 PM
|
Quorum-sensing (QS) systems based on acyl-homoserine lactones (AHLs) regulate gene expression in response to cell density in many bacteria, including Rhizobium. These systems, typically composed of LuxI-like synthases and LuxR-like regulators, control processes such as plasmid conjugation, biofilm formation, and plant interactions. However, their evolutionary dynamics and genomic distribution in Rhizobium remain poorly understood. We analyzed 142 complete Rhizobium genomes using comparative genomics, phylogenetic reconstruction, and genomic context analysis. LuxI/LuxR homologs were identified based on sequence similarity and Pfam domain architecture, and their genomic contexts were examined. Phylogenetic relationships and coevolution between LuxI/LuxR pairs were assessed using cophylogenetic approaches. QS systems showed a highly heterogeneous distribution across Rhizobium genomes: some strains lacked canonical systems, whereas others encoded one or multiple systems in chromosomes and/or plasmids. Chromosomal QS systems were associated with multiple distinct genomic contexts, supporting at least seven independent acquisition events. In contrast, plasmid-encoded systems exhibited substantially greater diversity in both sequence and genomic organization. Phylogenetic and comparative analyses revealed dynamic gains and losses of QS systems, variable coevolution among LuxI/LuxR pairs, and evidence of partner recruitment. Notably, plasmids appear to act as major reservoirs of QS systems and likely sources of their transfer to chromosomes. These findings indicate that QS systems in Rhizobium evolve through a combination of horizontal gene transfer, genomic rearrangement, and differential retention across replicons. The higher diversity and mobility of plasmid-encoded systems highlight their central role in shaping QS evolution and functional innovation. Overall, this study provides a comprehensive framework for understanding the diversification and evolutionary trajectories of QS systems in complex multipartite bacterial genomes.

|
Scooped by
Jean-Michel Ané
May 9, 2:42 PM
|
The relationship between plants and microorganisms has ancient roots. However, only recently we have begun to appreciate the true complexity of the holobiont and its intricate interactions. Fungi, bacteria and viruses residing within or around plants play a crucial role in enhancing their resilience to biotic and abiotic stressful factors. The impact of these interactions is not limited to stress resilience but are strictly correlated with the plant’s life cycle. The recognition of specific effects on plant growth and resilience has driven research towards utilizing microorganisms as sustainable solutions for agricultural challenges. The application of selected microorganisms, combined with the development of SynCom consortia, marks the beginning of a promising strategy for achieving sustainability in agriculture. This chapter explores the hidden world of plant-associated microorganisms, emphasizing their diverse beneficial impacts and providing evidence to support their potential applications in modern agriculture.
|






 Your new post is loading...
Your new post is loading...










Merry Christmas!