Your new post is loading...
 Your new post is loading...

|
Scooped by
Gilbert C FAURE
August 18, 2020 1:53 PM
|
(2020). Development of SARS-CoV-2 vaccines: should we focus on mucosal immunity? Expert Opinion on Biological Therapy: Vol. 20, No. 8, pp. 831-836.

|
Scooped by
Gilbert C FAURE
August 9, 2020 6:46 AM
|
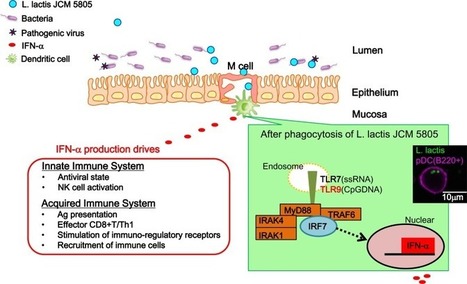
Toll-Like Receptor (TLR) 9 stimulation is required for induction of potent immune responses against pathogen invasion. The use of unmethylated CpG as adjuvants in vaccines provides an excellent means of stimulating adaptive immunity.

|
Scooped by
Gilbert C FAURE
July 14, 2020 8:29 AM
|
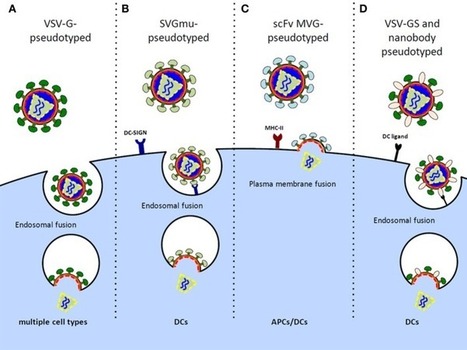
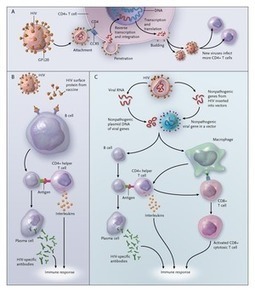
The development of an effective HIV vaccine to prevent and/or cure HIV remains a global health priority. Given their central role in the initiation of adaptive immune responses, dendritic cell (DC)-based vaccines are being increasingly explored as immunotherapeutic ...

|
Scooped by
Gilbert C FAURE
May 4, 2020 4:29 AM
|
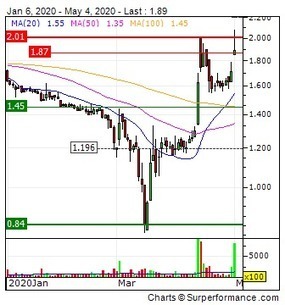
Regulatory News:
Vaxxel SAS, a French start-up, developing vaccines against respiratory viral infections, announces the acquisition of Transgene’s proprietary DuckCelt®-T17 cell line....| May 4, 2020...

|
Scooped by
Gilbert C FAURE
January 4, 2020 2:16 AM
|
The immune system mounts robust responses to infections, vaccines and cancer, but only now have scientists fully begun to unravel how non-circulating populations of T cells that reside in the body's "mucosal barrier tissues" ...

|
Scooped by
Gilbert C FAURE
December 5, 2019 9:01 AM
|
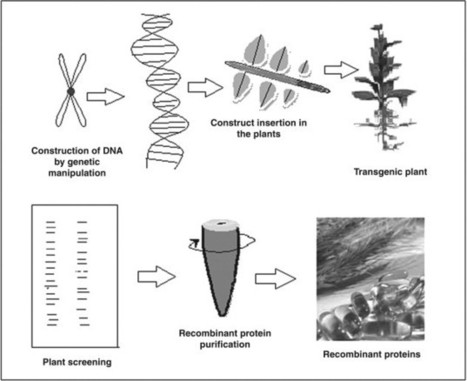
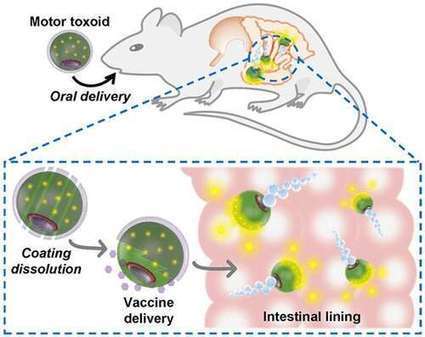
Edible Vaccine Edible vaccines are subunit preparations, do not involve attenuated pathogens, and improve the safety of individuals as compared to traditional vaccine since there is no possibility of proteins reforming into infectious organisms. From: Modern Applications of Plant Biotechnology in Pharmaceutical Sciences, 2015 Related terms: View all Topics Learn more about Edible Vaccine Edible Vaccines Saurabh Bhatia, Randhir Dahiya, in Modern Applications of Plant Biotechnology in Pharmaceutical Sciences, 2015 9.4.1 Advantages of Edible Vaccine • Edible vaccines are effective as a delivery vehicle for immunization because adjuvants that enhance the immune response are not required. • Edible vaccine can elicit mucosal immunity, which is not observed in traditional vaccines. • Edible vaccines are also cost effective in availability, storage, preparation, production, and transportation. Vaccines produced by biotechnological methods are stable at room temperature, unlike traditional vaccine, which needs cold chain storage, which multiplies the yearly cost to preserve vaccines. Moreover, the seeds of transgenic plants could be dried as there is less moisture content in seeds and the plants with oil or their aqueous extracts possess more storage opportunities. Manufacturing cost is low as there is no need for special premises to manufacture them. Edible vaccine can be easily produced at mass level in comparison to an animal system. • Edible vaccines are well tolerated, as they do not require administration by injection unlike traditional vaccines. Thus, there is also a reduced need for medical personnel and risk of contamination is low. The feasibility of oral administration compared to injection is also an advantage. • Plant-derived vaccines could be the source for new vaccines combining numerous antigens. These multicomponent vaccines are called second generation vaccines as they allow for several antigens to approach M-cells simultaneously. • Edible vaccines are subunit preparations, do not involve attenuated pathogens, and improve the safety of individuals as compared to traditional vaccine since there is no possibility of proteins reforming into infectious organisms. • The separation and purification of vaccines from plant materials is very easy and pathogenic contamination from animal cells can be effectively prevented. Read full chapter Purchase book Vaccines in Theory and Practice In Immunology for Pharmacy, 2012 Plant Vaccines Experimental edible vaccines, which offer protection against diarrheal disease, have been developed by using potatoes, rice, and bananas as vaccinating agents. To prepare a vaccine, microbial antigen genes are inserted into a Ti plasmid isolated from Agrobacterium tumefaciens. A modified Ti plasmid is capable of integrating into the plant cell genome and transforming the plant. The mature, transformed plant produces glycosylated microbial proteins in the edible parts of the plant. After the plant part is ingested, antigens stimulate local immunity, systemic immunity, or both. The benefits of edible vaccines are enormous. Inexpensive vaccines can be grown locally and administration of these vaccines does not require invasive medical procedures. Read full chapter Purchase book Vaccines and Clinical Immunization Tak W. Mak, Mary E. Saunders, in The Immune Response, 2006 One of the more intangible difficulties with edible vaccines is that these genetically engineered plants are negatively viewed by some as “frankenfoods,” or genetically modified organisms (GMOs) that may be harmful. Of course, these types of plants should be grown under strictly controlled conditions that limit their unintended spread. One technology that may alleviate concerns about the latter possibility is chloroplast transformation. Like mitochondria in mammals, chloroplasts in most plant species contain their own genome and are inherited maternally. Thus, exogenous genes introduced into the chloroplast genome stay with the transgenic plant and are not packaged and distributed in its pollen. The risk of transmission of the transgene beyond its prescribed borders is thus substantially reduced. Hopefully, sufficient clinical trial data can soon be accumulated that will demonstrate the efficacy and safety of edible vaccines, allowing us to finally achieve the worthy goal of vaccinating all the world's children against a wide spectrum of devastating diseases both cheaply and painlessly. Read full chapter Purchase book Plant-Based Biotechnological Products With Their Production Host, Modes of Delivery Systems, and Stability Testing Saurabh Bhatia, Randhir Dahiya, in Modern Applications of Plant Biotechnology in Pharmaceutical Sciences, 2015 8.2.1.1.6 Vaccines There has been considerable interest in developing low-cost, edible (i.e., oral) vaccines. Traditional edible vaccines, as for polio, use whole, attenuated organisms or semipurified materials to induce both systemic (Ig-G-mediated) and local membrane (Ig-A-mediated) immunity. Plant-based vaccines cover various proteins in form of antigens obtained from DNA encoded with antigenic sequences from pathogenic viruses, bacteria, and parasites. Key immunogenic proteins or antigenic sequences can be synthesized in plant tissues and subsequently ingested as edible subunit vaccines. The mucosal immune system can induce protective immune responses against pathogens or toxins, and may also be useful to induce tolerance to ingested or inhaled antigens. The production of secretory Ig-A (sIg-A) and provocation of specific immune lymphocytes can occur in mucosal regions, and these regions take on special importance in the development of edible vaccines. Aside from intrinsic low production cost, plant-based vaccines offer a number of unique advantages, including increased safety, stability, versatility, and efficacy. Plant produced vaccines can be grown locally where needed, avoiding storage and transportation costs. Relevant antigens are naturally stored in plant tissue, and oral vaccines can be effectively administered directly in the food product in which they are grown, eliminating purification costs. In many instances, it appears that refrigeration will not be needed to preserve vaccine efficacy, removing a major impediment to international vaccination efforts of the past. Plants engineered to express only select antigenic portions of the relevant pathogen may reduce immunotoxicity and other adverse effects, and plant-derived vaccines are free of contamination with mammalian viruses. Finally, the development of multicomponent vaccines is possible by insertion of multiple genetic elements or through cross-breeding of transgenic lines expressing antigens from various pathogenic organisms. There are, however, some limitations associated with the use of transgenic plants for vaccine production. A major limitation of the expression of recombinant antigens in transgenic plants is obtaining a protein concentration adequate to confer total immunity, given varying protein expression among and within the various plant species. Tight control of expression yields will likely be necessary to reduce variability and assure consistent, effective immunization. During the last decade, nearly a dozen vaccine antigens have been expressed in plants (Table 8.6). Transgenic potatoes can produce antigens of enterotoxigenic E. coli heat labile enterotoxin B subunit, and is effective in immunizing against viruses and bacteria that cause diarrhea. Still other “edible vaccines” are under development for rabies, foot and mouth disease (veterinary), cholera, and autoimmune diabetes. Transgenic lupin and lettuce plants can express hepatitis B surface antigen. Efforts are under way to develop an “edible vaccine” against the measles virus using the tobacco plant. A plant-based oral subunit vaccine for the respiratory syncytial virus (RSV) using either the apple or the tomato is under development. The plant species to be used for the production and delivery of an oral vaccine can be specifically selected to achieve desired goals. A large number of food plants (e.g., alfalfa, apple, asparagus, banana, barley, cabbage, canola, cantaloupe, carrots, cauliflower, cranberry, cucumber, eggplant, flax, grape, kiwi, lettuce, lupin, maize, melon, papaya, pea, peanut, pepper, plum, potato, raspberry, rice, service berry, soybean, squash, strawberry, sugar beet, sugarcane, sunflower, sweet potato, tomato, walnut, and wheat) have been transformed. Many of the high volume, high acreage plants such as corn, soybean, rice, and wheat may offer advantages. Corn, since it is a major component in the diet of the domestic animal, is a good candidate for vaccine production. In humans, particularly infants, the plant of choice to produce the vaccine might be the banana. Bananas are a common component of many infant diets and can be consumed uncooked, thus eliminating the possibility of protein denaturation due to high temperatures. Unfortunately, it is relatively difficult to create transgenic bananas and the production time is longer than for certain other food crops. Cereals and other edible plants are advantageous for vaccine production over plant species such as tobacco because of the lower levels of toxic metabolites. It is evident that there are numerous opportunities to identify and develop low-cost plant-derived vaccine materials, including edible plant-based vaccines [19]. Table 8.6. Recombinant Vaccines Expressed in Plants Year Vaccine antigen Species 1992 Hepatitis virus B surface antigen Tobacco 1995 Malaria parasite antigen Virus particle 1995 Rabies virus glycoprotein Tomato 1995 E. coli heat-labile Tobacco, enterotoxin, potato 1996 Human rhinovirus 14 (HRV-14) and human immunodeficiency virus type (HIV-1) epitopes Virus particle 1996 Norwalk virus capsid protein Tobacco, potato 1997 Diabetes-associated autoantigen Tobacco, potato 1997 Hepatitis B surface proteins Potato 1997 Mink enteritis virus epitope Virus particle 1997 Rabies and HIV epitopes Virus particle 1998 Foot and mouth disease virus VP1 structural protein Arabidopsis 1998 E. coli heat-labile enterotoxin Potato 1998 E. coli heat-labile enterotoxin Potato 1998 Rabies virus Virus particle 1998 Cholera toxin B subunit Potato 1998 Human insulin-cholera toxin B subunit fusion protein Potato 1999 Foot and mouth disease virus VP1 structural protein Alfalfa 1999 Hepatitis B virus surface antigen Yellow lupin, lettuce 1999 Human cytomegalovirus glycoprotein B Tobacco 1999 Dental caries (S. mutans) Tobacco 1999 Diabetes-associated autoantigen Tobacco, carrot 2002 Respiratory syncytial virus Tomato Read full chapter Purchase book Vaccines against Bacterial Enteric Infections Jan Holmgren, Myron M. Levine, in Mucosal Immunology (Fourth Edition), 2015 Plant-Based Vaccines An innovative live vector strategy in the 1990s was the concept of expressing protective vaccine antigens in transgenic plants for use as “edible vaccines” with the potential for generating affordable vaccines that would be easy to administer orally for impoverished populations in the developing world (see also Chapter 66). Various plants such as potatoes, tomatoes, lettuce, bananas, corn, and rice were used to express toxin antigens from V. cholerae and ETEC as well as antigens from Norwalk virus, hepatitis B virus, and rotavirus (Arntzen et al., 2005; Lugade et al., 2010). In early phase 1 clinical trials (Tacket, 2009), oral immunization with transgenic plant vaccines consisting of E. coli heat-labile enterotoxin B subunit expressed in potato (Tacket et al., 1998) or corn (Tacket et al., 2004a) induced toxin-neutralizing serum antibodies as well as intestine-derived IgA antibody-secreting cells and fecal IgA against the heat-labile toxin. Likewise, oral vaccination of human volunteers with potatoes expressing Norwalk virus capsid protein induced vaccine-specific IgA antibody-secreting cells as well as serum IgG antibodies. However, it is now generally accepted that the first-generation easy-to-make transgenic plant-based edible vaccines are unlikely to meet requirements for licensure; it remains to be seen if in future such plants, with increasing sophistication of expression of vaccine antigens, may still have usefulness for large-scale production of selected vaccine antigens. Read full chapter Purchase book History and Scope of Plant Biotechnology Saurabh Bhatia, in Modern Applications of Plant Biotechnology in Pharmaceutical Sciences, 2015 1.3.1 Biotechnology in Pharmaceutical Sciences Biotechnology in pharmaceutical sciences has brought about the production of monoclonal antibody, DNA, RNA probes for the diagnosis of various diseases; valuable drugs; edible vaccines like human hepatitis B; therapeutic drugs such as alkaloids, glycosides, steroids, flavonoids, tannins, proteins, enzymes, antibiotics, metabolites, etc. Interference with the plant genotype leads to the expression of various recombinant proteins, which forms antibodies, vaccines, and several other proteins having various pharmaceutical applications. Development of hairy root culture by means of Agrobacterium infection makes plants less dependent on growth hormones for their future growth. This genetic transformation of tumor in plants also gives a better yield of secondary metabolites. Even today, a variety of pharmaceutical drugs and chemicals are being produced by genetic engineering with better quality and increased quantity. Thus, plant biotechnology has provided us with a very efficient and economic technique for the production of a variety of biochemicals [133–136]. In industrial applications, plant biotechnology is used for the production of transgenic drugs. The major benefits are expected in medical, pharmaceutical, and health sciences. In medical sciences, it is used for the production of antibiotics, insulin, growth hormone, interferon, clotting factor VIII, vaccines, probes for infectious and gene therapy, etc. A major breakthrough in plant biotechnology was through rDNA technology, which led to the production of therapeutic recombinant proteins. The basis of the production of recombinant proteins is molecular pharming of therapeutic plants by rDNA technology, which is depicted in Fig 1.7. Genetic manipulation of DNA to form the final DNA construct is the initial step of rDNA technology. Further transfer of DNA construct in respective plants to conduct trangenesis is the second step of rDNA technology. This transfer is possible by using a suitable vector (medium) such as Agrobacterium sp. Successful transfer may lead to production of various transgenes. This transgenesis is followed by screening of plants. In this step, plants having the suitable gene expression for the desired recombinant protein are selected. Finally, recombinant proteins are purified to form various biopharmaceuticals and vaccines. Some of the popular plant-derived biopharmaceuticals are human growth hormone, enkephalin, IgG, human lactoferrin (antimicrobial), human serum albumin, human α- and β-interferon, human α1-antitrypsin, erythropoietin, hirudin, human α and β hemoglobin, etc. Some important vaccines such as envelope surface protein (hepatitis b virus (humans), glycoprotein (rabies virus), malarial B-cell epitope (malaria), and Escherichia coli Lt-B toxin (enterotoxigenic E. coli)) are also produced by rDNA technology [133–136]. Read full chapter Purchase book Mucosal Vaccines from Plant Biotechnology Hugh S. Mason, ... Tsafrir Mor, in Mucosal Immunology (Fourth Edition), 2015 Abstract The use of plants for production of recombinant proteins has evolved over the past 25 years. The first plant-based vaccines were expressed in stably transgenic plants, with the idea to conveniently deliver “edible vaccines” by ingestion of the antigen-containing plant material. These systems provided a proof of concept that oral delivery of vaccines in crude plant material could stimulate antigen-specific serum and mucosal antibodies. Transgenic grains like rice in particular provide a stable and robust vehicle for antigen delivery. However, some issues exist with stably transgenic plants, including relatively low expression levels and regulatory issues. Thus, many recent studies use transient expression with plant viral vectors to achieve rapid high expression in Nicotiana benthamiana, followed by purification of antigen and intranasal delivery for effective stimulation of mucosal immune responses. Read full chapter Purchase book Transgenic Plants for Mucosal Vaccines Hugh S. Mason, ... Charles J. Arntzen, in Mucosal Immunology (Third Edition), 2005 Viral diarrhea: Norwalk virus The Norwalk virus and related Norwalk-like viruses are responsible for 42% of outbreaks of acute epidemic gastroenteritis in the United States. The Norwalk virus capsid protein (NVCP) was the antigen chosen to develop an oral edible vaccine, since when expressed in insect cells it assembled into 38-nm Norwalk virus–like particles (VLPs) and reacted with serum of infected humans (Jiang et al., 1992). Tobacco and potato plants were transformed with constructs harboring the NVCP sequence; the plant recombinant protein assembled into VLPs identical to the insect cell–derived antigen (Mason et al., 1996). Mice that were gavaged with partially purified VLPs from tobacco leaf or fed with transgenic tubers developed serum IgG and fecal IgA antibodies specific for NVCP. A clinical trial was performed with the same potatoes used for the preclinical study (Tacket et al., 2000). Of 20 adult volunteers, 10 received two doses (days 0 and 7) and 10 received three doses (days 0, 7, and 21) of 150 g of raw transgenic potato tubers containing NVCP at 215 to 750 μg/dose. It is important to note that tuber expression was quite variable, and at most only half of NVCP in these potatoes was assembled as VLP; thus, the effective dose of potato vaccine was ∼325 μg/dose. Unassembled subunits are likely to be much less stable in the GI tract and thus less immunogenic. However, 19 of 20 subjects in the experimental group showed significant increases in the numbers of IgA antibody–forming cells (AFCs), ranging from 6 to 280 per 106 peripheral blood mononuclear cells (PBMCs), and 6 of 20 subjects in this group developed increases in IgG AFCs. Four volunteers showed increases in serum IgG anti-NVCP antibody titers, 4 had increased serum IgM, and 6 showed increased IgA in their stool samples (17-fold mean increase). Although the antibody responses were less impressive than those obtained with LT-B, the study showed that a plant-derived protein other than LT-B and CT-B can stimulate human immune responses after oral delivery. Insect cell–derived 250-µg doses of purified Norwalk VLP provided more effective seroconversion (Ball et al., 1999); thus it is likely that part of the potato-delivered NVCP was unavailable for uptake in the GI tract. More recent studies in transgenic tomato fruits with a plant-optimized NVCP gene resulted in higher expression and more potent immune responses in mice fed freeze-dried tomatoes (X. Zhang and H.S. Mason, unpublished results). A clinical trial is planned in which dried tomato powder formulated in gelatin capsules will be used to evaluate safety and immunogenicity (D. Kirk, H.S. Mason, and C.J. Arntzen, trial investigators). Read full chapter Purchase book Viruses as Tools for Vaccine Development Boriana Marintcheva, in Harnessing the Power of Viruses, 2018 8.6.2 Edible Vaccines A very attractive idea for alternative vaccine production and delivery is genetically engineering plants to produce vaccines that would be delivered to the human body as part of our diet, i.e., by eating traditional fruits and vegetables. Vaccine production in plants is already a fact due to advances of molecular farming (Chapter 4). However, the available vaccines are not edible, but rather traditional injectable component vaccines manufactured in plants. The bait rabies vaccine used to vaccinate wildlife is technically an edible vaccine; however, it contains attenuated vaccinia virus strain genetically modified to display rabies surface glycoprotein, i.e., newer generation subunit vaccine delivered in an edible packaging. The latter is effective because it uses the infectivity of the vaccinia virus to penetrate the animal body and is not limited by the so-called oral tolerance of our immune system. Oral tolerance essentially allows us to eat without detrimental immunological reaction to components of our food. Once a mechanism to overcome oral tolerance is found, it is envisioned that fruits and vegetables from our diet will be used to produce the vaccines. It is envisioned that plant material will be dried and packaged in capsules for oral delivery. It is hoped that the edible vaccines will not require refrigeration and will be significantly cheaper to produce. A huge hurdle in the process is the limited number of plants that can be easily manipulated by the tools of genetic engineering. The best candidates so far are tomatoes and potatoes, which are part of the human diet worldwide and happen to be relatives of tobacco, one of the most genetically amenable systems, but unfortunately, not edible due to toxicity. Progress has been made in genetically engineering bananas. Another problem to be solved is the delivery of consistent biologically active dose. Most likely, we are decades away from mass production of edible vaccines. Read full chapter Purchase book Plant-Based Vaccines Aboul-Ata E. Aboul-Ata, ... Pasquale Piazzolla, in Advances in Virus Research, 2014 3 Conclusion Constructed chimeric virus has to be inoculated, transfected, and/or infiltrated, using advanced methodologies, that is, nanoparticles and chitosan for transient expression through bioreactor plants (Dhama et al., 2013). Moreover, chimeric virus constructs are being commercially available (Yusibov & Rabindran, 2008), which makes edible vaccine development easy. Manns et al., (2001) have stated that sustained virological response (SVR) rate was 42% when peginterferon group was used after adjusting ribavirin. This type of therapeutics leads to using plant-based vaccines. Expression of potentially immunogenic peptides, either in transgenic plants or on the outer surface of genetically engineered chimeric viruses (Lico, Chen, & Santi, 2008; Tiwari, Verma, Singh, & Tuli, 2009), could offer remarkable advantages (Tacket & Mason, 1999). Specifically, the plant viruses are particularly attractive for producing oral vaccines because of their ability to infect edible crops. Plant components (fruits, leaves, and roots) can be eaten, providing an easy and inexpensive route of antigen (Ag) administration. In addition, edible plants are used as vehicles for delivering vaccines. This could protect these vaccines from degradation by gastric and intestinal fluids (Daniel, Streatfield, & Wyckoff, 2001; Webster, Thomas, Strugnell, Dry, & Wesselingh, 2002), because Ag delivery by plant cells protects the Ag during passage through the acid environment of the stomach. Finally, plant-derived vaccines eliminate the risk of contamination by zoonotic infections (Fischer, Stoger, Schillberg, Christou, & Twyman, 2004) such as virus or prion proteins, thereby diminishing the safety concerns associated with the use of many currently available types of vaccines. The use of plant viruses as nanoparticle platforms for producing a vaccine might have important clinical implications in oral vaccination, supporting the feasibility of producing a plant-derived Ag-presenting system. Read full chapter Purchase book

|
Scooped by
Gilbert C FAURE
November 12, 2019 1:54 PM
|
KEY POINTS VLP enhances antinorovirus IgA recall responses in humanized mice. Particulate structure is required for IgA enhancement. VLP-driven IgA responses are functionally superior to IgG responses. Abstract Virus-like particles (VLPs) provide a well-established vaccine platform; however, the immunogenic properties acquired by VLP structure remain poorly understood. In this study, we showed that systemic vaccination with norovirus VLP recalls human IgA responses at higher magnitudes than IgG responses under a humanized mouse model that was established by introducing human PBMCs in severely immunodeficient mice. The recall responses elicited by VLP vaccines depended on VLP structure and the disruption of VLP attenuated recall responses, with a more profound reduction being observed in IgA responses. The IgA-focusing property was also conserved in a murine norovirus-primed model under which murine IgA responses were recalled in a manner dependent on VLP structure. Importantly, the VLP-driven IgA response preferentially targeted virus-neutralizing epitopes located in the receptor-binding domain. Consequently, VLP-driven IgA responses were qualitatively superior to IgG responses in terms of the virus-neutralizing activity in vitro. Furthermore, the IgA in mucosa obtained remarkable protective function toward orally administrated virus in vivo. Thus, our results indicate the immune-focusing properties of the VLP vaccine that improve the quality/quantity of mucosal IgA responses, a finding with important implications for developing mucosal vaccines. Footnotes This work was partly supported by Grants-in-Aid for Scientific Research (C) 17K08895 and 25860377 from the Japan Society for the Promotion of Science and by the Research Program on Emerging and Re-emerging Infectious Disease from the Japan Agency for Medical Research and Development (Grant JP18fk0108051). The online version of this article contains supplemental material. Received April 29, 2019. Accepted October 8, 2019. Copyright © 2019 by The American Association of Immunologists, Inc. This article is distributed under The American Association of Immunologists, Inc., Reuse Terms and Conditions for Author Choice articles.

|
Scooped by
Gilbert C FAURE
September 27, 2019 2:36 PM
|
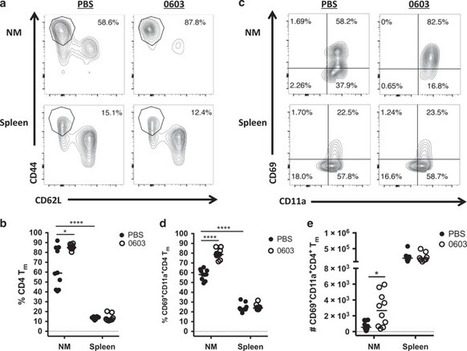
Scientists have identified a pair of molecules critical for T cells, part of the immune system, to travel to and populate the lungs. A potential application could be strengthening vaccines against respiratory pathogens such ...

|
Scooped by
Gilbert C FAURE
September 14, 2019 1:15 PM
|
KEY POINTS Trout have limited IgM and IgT repertoire diversity in NALT. Intranasal vaccination in trout triggers systemic and mucosal Ig response. IgM and IgT respond to i.p. and intranasal bacterin vaccination. Abstract Bony fish represent the most basal vertebrate branch with a dedicated mucosal immune system, which comprises immunologically heterogeneous microenvironments armed with innate and adaptive components. In rainbow trout (Oncorhynchus mykiss), a nasopharynx-associated lymphoid tissue (NALT) was recently described as a diffuse network of myeloid and lymphoid cells located in the olfactory organ of fish. Several studies have demonstrated high levels of protection conferred by nasal vaccines against viral and bacterial pathogens; however, the mechanisms underlying the observed protection are not well understood. We applied 5′RACE and a deep sequencing–based approach to investigate the clonal structure of the systemic and mucosal rainbow trout B cell repertoire. The analysis of Ig repertoire in control trout suggests different structures of IgM and IgT spleen and NALT repertoires, with restricted repertoire diversity in NALT. Nasal and injection vaccination with a bacterial vaccine revealed unique dynamics of IgM and IgT repertoires at systemic and mucosal sites and the remarkable ability of nasal vaccines to induce spleen Ig responses. Our findings provide an important immunological basis for the effectiveness of nasal vaccination in fish and other vertebrate animals and will help the design of future nasal vaccination strategies. Footnotes This work was supported by USDA AFRI Grant 2DN70-2RDN7 (to I.S.). S.M. also received funding from the People Programme (Marie Curie Actions) of the European Union’s Seventh Framework Programme (FP7/2007-2013) under REA Grant Agreement 600391. The dataset presented in this article has been submitted to the National Center for Biotechnology Information (https://www.ncbi.nlm.nih.gov/sra/) under accession number PRJNA551127. The online version of this article contains supplemental material. Abbreviations used in this article: D-NALT diffuse NALT ERM enteric red mouth HSD honestly significant difference HSR highly shared responding IMGT ImMunoGeneTics i.n. intranasally MID molecular identifier NALT nasopharynx-associated lymphoid tissue O-MALT MALT with well-organized SLOs O-NALT organized NALT PBT PBS pH 7.4 with 0.05% Tween 20 SLO secondary lymphoid organ UID unique molecular identifier. Received February 7, 2019. Accepted July 9, 2019. Copyright © 2019 by The American Association of Immunologists, Inc.

|
Suggested by
Société Francaise d'Immunologie
July 25, 2019 1:54 AM
|
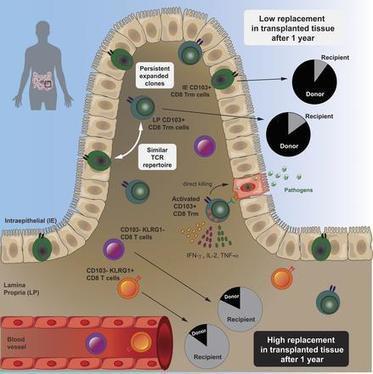
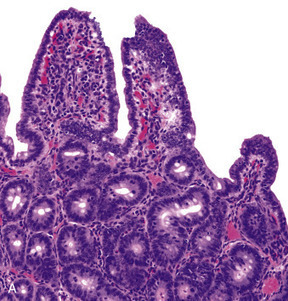
Graphical Abstract Abstract Resident memory CD8 T (Trm) cells have been shown to provide effective protective responses in the small intestine (SI) in mice. A better understanding of the generation and persistence of SI CD8 Trm cells in humans may have implications for intestinal immune-mediated diseases and vaccine development. Analyzing normal and transplanted human SI, we demonstrated that the majority of SI CD8 T cells were bona fide CD8 Trm cells that survived for >1 yr in the graft. Intraepithelial and lamina propria CD8 Trm cells showed a high clonal overlap and a repertoire dominated by expanded clones, conserved both spatially in the intestine and over time. Functionally, lamina propria CD8 Trm cells were potent cytokine producers, exhibiting a polyfunctional (IFN-γ+ IL-2+ TNF-α+) profile, and efficiently expressed cytotoxic mediators after stimulation. These results suggest that SI CD8 Trm cells could be relevant targets for future oral vaccines and therapeutic strategies for gut disorders. Submitted: 6 March 2019 Revision received 13 May 2019 Accepted: 20 June 2019 http://www.rupress.org/terms https://creativecommons.org/licenses/by-nc-sa/4.0/ This article is distributed under the terms of an Attribution–Noncommercial–Share Alike–No Mirror Sites license for the first six months after the publication date (see http://www.rupress.org/terms). After six months it is available under a Creative Commons License (Attribution–Noncommercial–Share Alike 4.0 International license, as described at https://creativecommons.org/licenses/by-nc-sa/4.0/).

|
Suggested by
Société Francaise d'Immunologie
April 20, 2019 3:01 AM
|
Research ArticlePulmonologyStem cells Free access | 10.1172/JCI125014 Yap/Taz regulate alveolar regeneration and resolution of lung inflammation Ryan LaCanna,1 Daniela Liccardo,1 Peggy Zhang,1 Lauren Tragesser,1 Yan Wang,2 Tongtong Cao,1 Harold A. Chapman,3 Edward E. Morrisey,4 Hao Shen,2 Walter J. Koch,1 Beata Kosmider,5 Marla R. Wolfson,5 and Ying Tian1 First published April 15, 2019 - More info Abstract Alveolar epithelium plays a pivotal role in protecting the lungs from inhaled infectious agents. Therefore, the regenerative capacity of the alveolar epithelium is critical for recovery from these insults in order to rebuild the epithelial barrier and restore pulmonary functions. Here, we show that sublethal infection of mice with Streptococcus pneumoniae, the most common pathogen of community-acquired pneumonia, led to exclusive damage in lung alveoli, followed by alveolar epithelial regeneration and resolution of lung inflammation. We show that surfactant protein C–expressing (SPC-expressing) alveolar epithelial type II cells (AECIIs) underwent proliferation and differentiation after infection, which contributed to the newly formed alveolar epithelium. This increase in AECII activities was correlated with increased nuclear expression of Yap and Taz, the mediators of the Hippo pathway. Mice that lacked Yap/Taz in AECIIs exhibited prolonged inflammatory responses in the lung and were delayed in alveolar epithelial regeneration during bacterial pneumonia. This impaired alveolar epithelial regeneration was paralleled by a failure to upregulate IκBa, the molecule that terminates NF-κB–mediated inflammatory responses. These results demonstrate that signals governing resolution of lung inflammation were altered in Yap/Taz mutant mice, which prevented the development of a proper regenerative niche, delaying repair and regeneration of alveolar epithelium during bacterial pneumonia. Graphical Abstract Introduction Bacterial pneumonia is a leading cause of mortality and morbidity worldwide. Despite effective antibiotics and vaccines, there is still substantial pulmonary morbidity in susceptible individuals (1). Infection by Streptococcus pneumoniae (the pneumococcus), the most common pathogen of community-acquired pneumonia, is characterized by acute elevation of bacterial number in the lung, diffuse alveolar epithelial damage, and a robust alveolar influx of leukocytes (2). While the virulence of the pneumococcus and host factors that contribute to bacterial pneumonia are becoming clear, up to now, little has been known about the types of damaged lung cells induced by bacterial pneumonia and the extent of lung regeneration following injury. Models of lung damage in mice have demonstrated that the regenerative processes in the lung involve local stem/progenitor cell populations (3). Alveolar epithelial type II cells (AECIIs), which express surfactant protein C (SPC), have been considered to be stem/progenitor cells for the alveolar epithelium (3). Lineage tracing of SPC+ AECIIs demonstrates their capacities of self-renewal and differentiation into AECIs during both the homeostatic state and regeneration after injury (4). Besides their stem/progenitor capacities, AECIIs have immune-modulatory functions. AECIIs release cytokines and chemokines that recruit leukocytes to the sites of injury/infection and activate these cells to mount an immune response (5, 6). Dysregulated immune responses have been implicated in a variety of inflammatory and fibrotic conditions affecting humans. In some types of pulmonary fibrosis, patients have abnormal alveolar structure with a loss of AECIs and accumulation of immune cells (7–9). The mechanisms for the failure of AECI regeneration are not clear, however. One explanation for the loss of AECIs is the loss of local stem/progenitor cells, the sources for generating new AECIs, after injury. However, this stem/progenitor cell loss cannot be the sole reason for the impaired regeneration, as lung injuries that do not cause loss of stem/progenitor cells can still lead to failed AECI regeneration in animal models and patients with pulmonary fibrosis (10). Those findings imply that non–stem/progenitor cell loss-specific mechanisms are responsible for the impaired lung regeneration that occurs in fibrotic lungs. The molecular mechanisms underlying lung regeneration have been studied in adult mice following pneumonectomy (11). Regeneration of lung alveoli depends on an increased inflammatory state in the lung, a process that requires recruitment of monocytes and macrophages into lung alveoli. It is unclear how inflammatory responses that normally lead to lung regeneration in acute injury are absent in fibrotic lungs with persistent inflammatory responses. Here, we show that mice infected with S. pneumoniae strain T4 (SpT4) have injuries exclusively in lung alveoli, with loss of AECIs and AECIIs and increased infiltration of immune cells. This is followed by rapid resolution of lung inflammation and alveolar epithelial regeneration via proliferation of preexisting SPC+ AECIIs and their differentiation into AECIs. We show that nuclear expression of Yap and Taz, the mediators of the Hippo signaling pathway, were markedly increased in AECIIs following SpT4 infection. We investigate the role of Yap and Taz in AECIIs during alveolar epithelial regeneration and resolution of lung inflammation in response to bacterial pneumonia–induced lung injury. Results Alveolar epithelial injury and recovery following S. pneumoniae–induced bacterial pneumonia in mice. We used SpT4 to infect mice by intranasal inhalation under anesthetization with a dose of approximately 5 × 106 CFU. SpT4 infection causes direct infection of the lower respiratory tract and acute bacterial pneumonia in mice, as indicated in previous reports (12, 13). Lungs were harvested at 2, 4, 7, and 14 days post infection (dpi) and analyzed by histology and flow cytometry. The levels of pneumococcal bacteria and cell apoptosis were highest in the alveolar spaces within the first 2 days after infection, as indicated by immunostaining with antibody specific for SpT4 capsule and TUNEL staining, individually (Figure 1, A–C). In contrast, their levels were reduced after 2 dpi, as seen by the gradual decrease in SpT4+ and TUNEL+ numbers within the alveolar region by 14 dpi. To determine pneumococci clearance in the lung, bacterial loads were measured by plating lung homogenate. A transient increase of bacterial load was detected in mouse lung, with the highest level at 2 dpi and 4 dpi, followed by a significant drop at 7 dpi (Figure 1B). Substantial destruction of alveolar epithelial cells was observed, as evidenced by significant loss of cell type–specific markers for AECIs (T1a) and AECIIs (SPC) at 2, 4, and 7 dpi (Figure 1A). By 14 dpi, the number of those cell populations returned to the basal level (Figure 1, D and E). The damage and recovery of lung epithelium during bacterial pneumonia were specific to the alveolar region, as there were no significant changes in the expression of airway epithelial cells, including club cells (CC10), ciliated cells (bTubulin IV), and basal cells (p63) (Supplemental Figure 1A; supplemental material available online with this article; https://doi.org/10.1172/JCI125014DS1). Figure 1 Alveolar epithelial injury and recovery in SpT4-infected mice. Lung tissues were collected at 0, 2, 4, 7, and 14 dpi with SpT4. (A) Immunostaining on lung sections with antibodies to the type 4 capsule of SpT4, T1a, and pro-SPC (SPC). Cell apoptosis was measured by TUNEL staining. Cell nuclei were stained with DAPI (blue). Scale bars: 50 μm. (B) Mouse lungs were homogenized, and lung lysates were plated for quantitative culture of colonizing pneumococci. (C) Quantification of cell apoptosis by counting TUNEL+ cells on lung sections. (D) Lung cells were dissociated, and T1a+ cells were quantified as percentage of total CD45– cells by flow cytometry. (E) Quantification of the number of SPC+ cells on lung sections. n ≥ 10 randomly selected fields per animal (C and E); n = 3–9 per group (B–E). **P < 0.01; ***P < 0.001; ****P < 0.0001, 1-way ANOVA. Alveolar epithelial regeneration and inflammatory resolution during bacterial pneumonia. We first examined AECII proliferation after SpT4-induced lung injury. To determine DNA synthesis in AECIIs, mice received a single i.p. injection of EdU and were sacrificed after a 3-hour labeling period. The frequency of EdU incorporation was determined on sectioned lungs by colabeling with antibody against AECII (SPC) (Figure 2A). A transient increase of DNA synthesis was observed in AECIIs, with a peak labeling index of 7% ± 0.3% occurring at 4 dpi (Figure 2B). Figure 2 Alveolar epithelial regeneration and Yap/Taz expression in mouse lungs during bacterial pneumonia. (A) Schematic of experimental design (left) and confocal images of lung sections at 0 dpi and 4 dpi. AECIIs in DNA synthesis phase were detected using Click-iT EdU Alexa Fluor (green) and coimmunostaining with antibody against pro-SPC (SPC) (red). Cell nuclei were stained with DAPI (blue). (B) Quantification of EdU+SPC+ cells as percentage of total SPC+ cells analyzed (~2200 SPC+ cells per animal). (C) Confocal images of lung sections of SPC-CreERT2, Rosa26-mTmG mice at 0, 7, and 14 dpi. Mice were administrated with 3 doses of tamoxifen to label SPC+ AECIIs. Fourteen days after the last tamoxifen treatment, mice were infected with SpT4. AECII-to-AECI differentiation was visualized by coimmunostaining with antibodies against GFP (lineage-labeled AECIIs) and T1a (AECIs). Arrowheads point to regions double-positive for GFP and T1a. (D) Quantification of percentage of GFP+T1a+ area of total GFP+ area per field using ImageJ. (E) Flow cytometry analysis of dissociated lung cells showing the percentage of GFP+T1a+ cells of total T1a+ cells at indicated time points. (F) Confocal images of lung sections of SPC-CreERT2, Rosa26-mTmG mice. Immunostaining with antibodies against GFP (lineage-labeled AECIIs) and Yap and Taz. Cell nuclei were stained with DAPI (blue). (G) Western blot using lung tissue lysates at 0 dpi or purified AECIIs at 0 and 7 dpi, blotted with anti-YAP, anti-pYAP (Ser127), anti-Taz, anti-pTaz (S89), and anti–β-actin. Histograms showed average of total YAP or TAZ normalized to β-actin (loading control), together with average ratio of pYAP/YAP and pTAZ/TAZ. n ≥ 4 per group (B, D, E); n = 3 per group (G). *P < 0.05; **P < 0.01; ****P < 0.0001, 1-way ANOVA (B, D, E) and Student’s t test (G). Scale bars: 10 μm. We next performed genetic lineage tracing of adult SPC+ AECIIs using the SPC-CreERT2, Rosa26-mTmG mouse line. Mice were administrated with 3 i.p. injections of tamoxifen at 0.2 mg/g per dose to induce GFP expression in SPC+ AECIIs. Mice were then infected with SpT4 at 14 days after the last tamoxifen treatment. Lungs were analyzed by immunohistochemistry and flow cytometry. Differentiation of AECIIs to AECIs was measured by coimmunostaining for lineage-labeled AECII marker (GFP) and AECI marker (T1a) (Figure 2C). We examined AECII-to-AECI differentiation by quantifying the percentage of GFP+ alveolar surface area covered by AECII-derived AECIs (GFP+T1a+) on sectioned lungs. There were no significant changes between noninfected and SpT4-infected lungs within 6 dpi. However, we observed significant increases in the levels of AECII-to-AECI differentiation (GFP+T1a+) at 7 dpi and onwards (Figure 2D and Supplemental Figure 1B). Consistent with this finding, FACS analysis showed the significant increase in the percentage of AECIs derived from preexisting AECIIs (GFP+T1a+) at 7 dpi compared with that in noninfected lungs. At 14 dpi, the level of GFP+T1a+ was elevated by 4.7 ± 0.4-fold compared with that at 7 dpi (Figure 2E). It has been postulated that inflammatory cells are important drivers of tissue repair and regeneration (14, 15). However, correlations between persistent lung inflammation and decreased lung regeneration have been observed in both various animal injury models and human patients (8, 10). To assess the lung inflammatory responses and their association with alveolar epithelial regeneration, we characterized the immune phenotypes in mouse lungs during bacterial pneumonia. Consistent with previous reports (12, 13), high levels of inflammatory responses were observed in the lung, as evidenced by the significant increases in the number of immune cells (CD45+) within 7 dpi, including lymphocytes (CD3+), macrophages (CD11c+CD64+), and neutrophils (Ly6G+) (Supplemental Figure 2, A–E). Yet the accumulation of inflammatory cells in the lung waned over time, as seen by the decrease in the number of those cell populations by 14 dpi. Mouse lungs exhibited profibrotic lesions at 7 dpi, but recovered from SpT4 infection without the acquisition of lung fibrosis at 14 dpi, as evidenced by Masson’s trichrome staining and hydroxyproline assay on lung tissue lysates (Supplemental Figure 2, F and G). The transient accumulation of lung inflammatory cells was accompanied by the loss of AECIs and the increase of profibrotic lesions, while the resolution of inflammatory cells coincided with the time frame of AECI recovery, AECII-to-AECI differentiation, and regression of profibrotic lesions in the lung (Supplemental Figure 2, H–J), implying that resolution of lung inflammation was closely associated with alveolar epithelial regeneration. Yap and Taz expression in AECIIs during bacterial pneumonia. To understand the mechanisms underlying alveolar epithelial regeneration following SpT4-induced lung injury, we performed microarray analysis on lineage-labeled AECIIs (GFP+) isolated from SPC-CreERT2, Rosa26-mTmG mouse lungs before SpT4 infection and at 8 dpi with SpT4. After normalization of data sets, we used ToppGene to identify pathways that were induced (FDR, <5% and >1.5-fold expression) in GFP+ AECIIs of SpT4-infected mice. Pathways associated with Hippo signaling, T cell receptor signaling, JAK/STAT, and p38 signaling were all induced in SpT4-infected mice (Supplemental Table 1). Quantitative reverse transcriptase PCR (qRT-PCR) analysis confirmed increased expression of Yap and Taz, the 2 paralogous transcriptional coactivators of the Hippo pathway (16, 17), as well as their target genes including Ctgf, Cyr61, and Birc5 in GFP+ AECIIs of SpT4-infected mice at 7 dpi compared with those of mice without infection (0 dpi) (Supplemental Figure 3A). Inhibition of Hippo signaling leads to nuclear Yap/Taz localization and activity (16, 17). Nuclear Yap/Taz expression was not detected in AECIIs of noninfected lungs (Figure 2F). However, we observed increases in nuclear Yap/Taz protein levels in AECIIs before (4 dpi) and during (7 dpi) differentiation toward AECIs (Figure 2F). Western blot confirmed higher levels of Yap/Taz protein in AECIIs purified from lungs at 7 dpi compared with those from noninfected lungs (Figure 2G). Moreover, nuclear locations of Yap/Taz proteins were also elevated in AECIIs at 7 dpi compared with noninfected AECIIs, as quantification of relative expression of phosphorylated Yap (p-Yap, Ser127) versus total Yap and p-Taz (Ser89) versus total Taz revealed lower levels of p-Yap/Yap and p-Taz/Taz in AECIIs at 7dpi than in noninfected AECIIs (Figure 2G). Yap/Taz deletion in AECIIs impairs alveolar epithelial regeneration and causes prolonged fibrotic lesions during bacterial pneumonia. The nuclear localization of Yap and Taz in AECIIs in response to SpT4-induced lung injury could potentially augment alveolar regeneration by enhancing their targeting of regeneration-related genes. We tested this in the adult mouse lung with tamoxifen-inducible Yap/Taz gene deletion in SPC+ AECIIs. SPC-CreERT2, Yapfl/fl, Tazfl/fl, Rosa26-mTmG mice were administrated with tamoxifen i.p. to delete Yap and Taz in SPC+ AECIIs (referred as Yap/Taz mutant mice). Yap/Taz deletion and reduced expression of their target genes were verified by qRT-PCR analysis performed on purified AECIIs 14 days after tamoxifen administration (Supplemental Figure 3B). To gain insight into the role of Yap/Taz in AECIIs in the steady-state adult lung, we quantified the survival, self-renewal, and differentiation of lineage-labeled SPC+ AECIIs at 9 days and 12 weeks after tamoxifen administration. We lineage labeled AECIIs by generating SPC-CreERT2, floxed Yap/Taz, Rosa26-mTmG mice so that a GFP reporter could be used to label and trace AECIIs in which Yap/Taz had been deleted. We found no detectable changes in GFP+ cell apoptosis, as assessed by TUNEL staining at 9 days and 12 weeks after tamoxifen administration (Supplemental Figure 3, C and D). A transient decrease of DNA synthesis was observed in the lineage-labeled AECIIs from Yap/Taz mutant mice compared with SPC-CreERT2, Rosa26-mTmG control mice at 9 days after tamoxifen administration (0.49% ± 0.10% vs. 1.45% ± 0.10%; P < 0.001; Supplemental Figure 3E). The decreased AECII cell-cycle progression in Yap/Taz mutant mice was also observed by immunostaining of sectioned lungs for the cell cycle marker Ki67 (0.30% ± 0.09% vs. 1.80% ± 0.09%; Yap/Taz mutant vs. SPC-CreERT2, Rosa26-mTmG control, respectively; P < 0.001; Supplemental Figure 3F). By 12 weeks after tamoxifen administration, there were no significant differences in the percentage of lineage-labeled AECIIs that were EdU+ (EdU+GFP+) between Yap/Taz mutant and control mice (0.16% ± 0.02%. vs 0.18 % ± 0.08%; Supplemental Figure 3E). However, the level of Ki67+GFP+ was still lower in Yap/Taz mutant mice than that in control mice (0.76% ± 0.09% vs.1.44% ± 0.4%, P < 0.01; Supplemental Figure 3F). Quantification of AECII-to-AECI differentiation by immunostaining and flow cytometry showed that Yap/Taz mutant mice exhibited reduced percentages of lineage-labeled AECIIs that had differentiated into AECIs (GFP+T1a+) at 12 weeks after tamoxifen administration compared with control mice (1.68% ± 0.25% vs. 2.70% ± 0.10%; P < 0.05; Supplemental Figure 3G). Histological analysis showed Yap/Taz mutant mice had normal alveolar structures at 4 weeks after tamoxifen administration, but exhibited patchy distribution of subtle interstitial infiltrates in the alveolar septa at 12 weeks after tamoxifen administration (Supplemental Figure 3H). FACS analysis revealed that Yap/Taz mutant lungs at 12 weeks after tamoxifen administration had increased numbers of inflammatory cells (CD45+) compared with SPC-CreERT2 control lungs (Supplemental Figure 3I). These data indicated that deletion of Yap/Taz in adult SPC+ AECIIs led to decreased AECII proliferation and AECII-to-AECI differentiation during steady-state tissue maintenance. Long-term deletion of Yap/Taz in SPC+ AECIIs caused increased accumulation of inflammatory cells in the lung. To determine whether Yap/Taz contributed to alveolar epithelial regeneration in vivo, Yap/Taz mutant mice were exposed to SpT4-induced lung injury. Yap/Taz mutant mice showed delayed recovery from bacterial pneumonia, as they regained body weight more slowly compared with the SPC-CreERT2 control mice (Supplemental Figure 4A). Yap/Taz mutant mice had higher levels of total protein in their bronchoalveolar lavage fluid (BALF) at 7 dpi and 14 dpi compared with the control mice (Supplemental Figure 4B), indicating more damage to lung tissue integrity in Yap/Taz mutant mice. SPC-CreERT2 control mice started to regenerate AECIs (T1a+) and AECIIs (SPC+) within a week after SpT4 infection and had full recovery of those cell populations by 14 dpi (Figure 1, D and E and Figure 3, A and B). In contrast, Yap/Taz mutant mice showed persistently low levels of AECIs and AECIIs for more than 2 weeks and only started to regenerate those cell populations by 21 dpi, indicating delayed alveolar epithelial regeneration following SpT4 infection (Figure 3, A and B). Yap/Taz mutant mice developed severe fibrotic lesions within the alveolar region at 14 dpi, as evidenced by increased Ashcroft scores in Trichrome-stained lung sections and collagen burden using hydroxyproline assay on lung tissue lysates compared with SPC-CreERT2 control mice (Figure 3, C–E). Fibrotic lesions regressed over time, as seen by the gradual decreases in Ashcroft score and collagen burden by 56 dpi in Yap/Taz mutant lungs (Figure 3, D and E, and Supplemental Figure 4C). Lungs of single Yap mutant mice developed minor fibrotic lesions at 14 dpi, while lungs of single Taz mutant mice recovered from bacterial pneumonia without acquisition of fibrotic lesions at 14 dpi (Supplemental Figure 4D). We focused on Yap/Taz mutant mice in this study, since single mutant mice had minor or no phenotypes in response to SpT4 infection. Figure 3 Phenotypes of Yap/Taz mutant lungs during bacterial pneumonia. (A) Immunostaining on lung sections with nuclei labeled by DAPI (blue) and antibodies to T1a (red) or pro-SPC (SPC) (green). (B) Lung cells were dissociated and T1a+ cells were quantified as percentage of total CD45– cells by flow cytometry. SPC+ cells were quantified by counting the number of SPC+ cells per field (≥10 randomly selected fields per animal) (n = 3–8 per group). (C) Lung tissue sections were stained with Alcian blue and Nuclear Fast Red. Lung fibrotic lesions were quantified by measuring (D) Aschcroft score and (E) hydroxyproline assay (n = 3–4 per group). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001, 2-way ANOVA. Scale bars: 50 μm (A, C [bottom panel]); 500 μm (C [top panel]). To determine the causes of impaired alveolar epithelial regeneration observed in Yap/Taz mutant mice, we profiled lung bacterial loads by measuring CFU and found no differences in bacteria number in lung homogenates and blood serum from Yap/Taz mutant mice compared with SPC-CreERT2 controls after SpT4 infection (Supplemental Figure 4, E and F). We next evaluated the effects of the Yap/Taz deletion on the process of alveolar epithelial damage. TUNEL staining of lung sections showed no differences in the number of apoptotic cells between Yap/Taz mutant and control lungs (Supplemental Figure 4, G–H). These results indicated that bacterial load and cell apoptosis were unlikely to explain the effects of the Yap/Taz deletion on alveolar epithelial regeneration and the development of lung fibrotic lesions during bacterial pneumonia. To determine how Yap/Taz might contribute to alveolar epithelial regeneration, we examined their effects on AECII proliferation and AECII-to-AECI differentiation (Figure 4A). The proliferation index of AECIIs in Yap/Taz mutant mice at 7 dpi was significantly lower than that in SPC-CreERT2, Rosa26-mTmG control mice, as shown by the percentage of lineage-labeled AECIIs (GFP+) that were either EdU+ or Ki67+ (Figure 4, B and C). At 14 dpi, 24.1% ± 2.1% of total lineage-labeled GFP+ cells in control lungs were T1a+ (GFP+T1a+). In contrast, GFP+T1a+ cells were reduced by 72.6% (6.6% ± 2.1%) in Yap/Taz mutant lungs (Figure 4, D–F), indicating decreased AECII-to-AECI differentiation in Yap/Taz mutant lungs. The AECII-to-AECI differentiation defect was persistent in Yap/Taz mutant lungs, as the levels of AECIs derived from lineage-labeled AECIIs (GFP+T1a+) were still significantly lower in Yap/Taz mutant lungs at 56 dpi than in SPC-CreERT2, Rosa26-mTmG control lungs (Figure 4F). Collectively, these data indicated that loss of Yap/Taz in AECIIs diminished alveolar epithelial regeneration derived from preexisting SPC-expressing AECIIs and led to prolonged fibrotic lesions in the lung during bacterial pneumonia. Figure 4 AECII proliferation and differentiation in Yap/Taz mutant lungs during bacterial pneumonia. (A) Schematic of experimental design for studies shown in B–F. (B) Lung tissue sections from SPC-CreERT2, Rosa26-mTmG mouse were immunostained with DAPI (blue) and antibody against GFP (lineage-labeled AECIIs) (green), and colabeling with Click-iT EdU Alexa Fluor (red) and confocal images were taken. The percentages of GFP+EdU+ cells of total GFP+ cells per field were graphed (bottom panel). (C) Confocal image of lung section from SPC-CreERT2, Rosa26-mTmG mouse at 7 dpi with nuclei labeled by DAPI (blue) and antibodies against GFP (green) and Ki67 (red). Percentages of GFP+Ki67+ cells of total GFP+ cells per field were graphed (bottom panel). (D) Confocal images of lung section at 14 dpi with nuclei labeled by DAPI (blue) and antibodies against GFP (green) and T1a (red). Asterisks indicate regions double-positive for GFP and T1a. (E) Lung cells were dissociated and flow cytometry was performed by gating on GFP+T1a+. The numbers in the top left gates represent all GFP+ cells of total live CD45– cells. The numbers in the top right gates represent GFP+T1a+ cells of total T1a+ cells. The numbers in the bottom right gates represent all T1a+ cells of total live CD45– cells. (F) Quantification of GFP+T1a+ cells as the percentage of GFP+T1a+ of total T1a+ cells by flow cytometry. n = 4–5 per group (B and C); n = 4–8 per group (F). **P < 0.01; ***P < 0.001; ****P < 0.0001, 2-way ANOVA. Scale bars: 25 μm (B and C); 20 μm (D). Yap/Taz regulate AECII inflammatory responses through targeting IκBa. To understand the underlying mechanisms for these observations, we performed microarray analysis on lineage-labeled AECIIs (GFP+) isolated from Yap/Taz mutant and SPC-CreERT2 control lungs at 8 dpi. After normalization of data sets, we used ToppGene to identify pathways that were induced or repressed (FDR, <5% and >2-fold expression) in GFP+ AECIIs of Yap/Taz mutant mice. While gene ontologies associated with negative regulation of the apoptotic process, positive regulation of cell proliferation, and differentiation were repressed in Yap/Taz mutant mice, gene ontologies associated with negative regulation of cell differentiation, inflammation response, leukocyte chemotaxis, and lipid metabolic process were all induced in Yap/Taz mutant mice (Supplemental Figure 5A and Supplemental Tables 1 and 2). qRT-PCR confirmed decreased expression of Yap, Taz, tand Yap/Taz-dependent target genes, including Ctgf, Cyr61, Birc5, and Birc2, in GFP+ AECIIs of Yap/Taz mutant mice at 7 dpi compared with those of SPC-CreERT2 control mice (Figure 5A). The expression of several markers associated with positive regulation of cell proliferation and differentiation, including Fgf1, Fgfr3, Tgfb2, Wnt3a, Bmp4, and Egfr, was significantly reduced in GFP+ AECIIs of Yap/Taz mutant mice (Figure 5A). These data suggested that loss of Yap/Taz in AECIIs led to decreased proliferation and differentiation of AECIIs in response to SpT4-induced lung injury. Notably, while expression of Nfkb1 itself was unchanged, expression of Nfkb1 repressor genes Ikba (Nfkbia) and Ikbb (Nfkbib) was significantly decreased in Yap/Taz mutant GFP+ AECIIs (Figure 5A). In contrast, the expression of NF-kB–dependent target genes, including Cxcl3 and Ccl21a, was significantly increased in Yap/Taz mutant GFP+ AECIIs (Figure 5A). We detected a significant increase in IL-1b protein, a cytokine known to expand differentiated T cells (18), in BALF from Yap/Taz mutant mice at both 7 dpi and 14 dpi compared with that from SPC-CreERT2 control mice (Figure 5B). Cytokine array on lung tissue lysates showed the higher levels of cytokines, including IL-1b as well as CXCL9, a T cell chemotactic cytokine (19), in Yap/Taz mutant mice at 7 dpi and 14 dpi compared with SPC-CreERT2 control lungs (Figure 5C and Supplemental Figure 5, B and C). Consistent with the elevated expression of genes and cytokines associated with T cell inflammatory responses in Yap/Taz mutant AECIIs, we observed a significant increase in the number of CD3+ T cells in Yap/Taz mutant lungs at 7 dpi and 14 dpi (Figure 5D). Immunostaining on sectioned lung tissues confirmed the increased number of CD3+ T cells located in lung alveoli of Yap/Taz mutant mice at 14 dpi (Figure 5, E and F). Interestingly, CD3+ T cells were enriched in areas where AECIIs failed to differentiate into AECIs in Yap/Taz mutant lungs (Figure 5F). In addition, the number of macrophages (CD11c+CD64+) in Yap/Taz mutant lungs was lower at 7 dpi, but higher at 14 dpi, than those in SPC-CreERT2 control lungs (Supplemental Figure 5D). In contrast, the number of neutrophils (Ly6G+) was not significantly changed in Yap/Taz mutant lungs compared with control lungs (Supplemental Figure 5D). The effect of Yap/Taz mutant AECIIs on lung inflammation waned over time, as seen by the equivalent levels of T cells (CD3+) and macrophages (CD11c+CD64+) at 56 dpi between Yap/Taz mutant and SPC-CreERT2 controls (Figure 5D and Supplemental Figure 5D). These results indicated that loss of Yap/Taz in AECIIs led to prolonged inflammatory responses in lung alveoli during bacterial pneumonia. Figure 5 Inflammatory responses in Yap/Taz mutant lungs. (A) GFP+ AECIIs at 7 dpi were sorted by FACS and analyzed by qRT-PCR (n = 4 per group). (B) BALF was analyzed for IL-1b by ELISA assay (n = 4–6 per group). (C) Cytokine assay showed protein levels of IL-1b and CXCL9 in mouse lung lysates (n = 1 per group). (D) Flow cytometry of dissociated lung cells was performed by gating on CD3+CD45+ cells, and quantification of total number of CD3+CD45+ cells in the lung was graphed (n = 3–10 per group). (E) Confocal images of lung sections at 14 dpi with nuclei labeled by DAPI (blue) and antibodies to GFP (green) and CD3 (red). Scale bars: 20 μm. (F) Quantification of the number of CD3+ cells as the ratio of CD3+ cell number versus GFP+ area per field using ImageJ (n = 3–5 per group). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001, Student’s t test (A) and 2-way ANOVA (B, D, F). Because Yap/Taz mutant AECIIs had reduced expression of Ikba and Ikbb, we hypothesized that an IκB/NF-κB pathway drives the inflammatory responses in Yap/Taz mutant lungs during bacterial pneumonia. Given recent identification of the Yap/IκB signaling axis as a potential means of generating inflammatory response in Drosophila post-Gram+ bacterial infection (20), we investigated whether Yap/Taz controls IκB levels in the mouse lung. Examination of the Ikba genomic locus revealed 5 potential Tead-binding motifs (TBM) within a 10 kb region upstream of the transcriptional start site (Figure 6A), while there was no potential TBM in the 10 kb region upstream of the Ikbb genomic locus. For the subsequent experiments, we focused on IκBa because it was shown to be the most efficient IκB molecule to control NF-κB transcriptional activity in previous studies (21–23). We first performed ChIP assays on chromatin obtained from the murine lung epithelial cell line MLE-15 using either an anti-Tead or anti-Yap antibody to determine whether Tead and Yap were associated with any of the 5 TBMs. ChIP analysis showed that Yap and Tead were associated with the TBM1 site of Ikba (Figure 6B). To determine whether the TBM1 site was responsive to Tead transactivation, MLE-15 cells were transfected with the pGL3 luciferase reporter containing TBM1–3 region along with an expression plasmid encoding murine Tead2 (Figure 6C). This study showed that expression of Tead2 transactivated the pGL3-IκBa-TBM1–3.luc reporter. Coexpression of Tead2 with Yap or Taz further enhanced the transactivation of the pGL3-IκBa-TBM1-3.luc reporter (Figure 6C). In contrast, expression of Tead2 did not activate the pGL3-IκBa-TBM1-3 mutant.luc reporter in which TBM1 was mutated (Figure 6C). Figure 6 Regulation of Yap/Taz on IκBa expression and NF-κB transcriptional activity in AECIIs. (A) Schematic of IκBa genomic locus showing potential Tead binding sites (TBM) in the 10 kb upstream of the IκBa transcription start site. (B) Chromatin from MLE-15 cells was immunoprecipitated with either Tead or Yap antibody, and qRT-PCR results were graphed (n = 3). (C) MLE-15 cells were transfected with pGL3 vector containing a IκBa-TBM1-3 or a IκBa-TBM1-3 mutation, in which TBM1 was mutated, along with the expression plasmid encoding either murine Tead2 or Yap or Taz. Twenty-four hours after transfection, cells were processed for luciferase activity measurement (n = 3–6 per group). (D) Adult WT mouse AECIIs were purified, cultured, and infected with either Yap shRNA lentivirus or scramble shRNA lentivirus or were given no treatment. Forty-eight hours after lentiviral infection, cells were processed for NF-κB transcription activity measurement. (E) AECIIs were purified from mouse lungs at 0 dpi and 7 dpi, and their NF-κB transcriptional activity was graphed. (F) MLE-15 cells were infected with either Yap shRNA lentivirus or scramble shRNA lentivirus or were given no treatment. Forty-eight hours after lentiviral infection, cells were transfected with NF-κB luciferase vector containing NF-κB response elements, along with an expression plasmid encoding either murine Yap or Taz. Twenty-four hours after transfection, cells were processed for luciferase activity measurement (n = 3). (G) Schematic model of interaction of Yap/Taz and IκBa/NF-κB in AECIIs. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001, 1-way ANOVA (B, C, D, F) and 2-way ANOVA (E). IκBa has previously been shown to be required for inhibiting NF-κB–mediated inflammatory response in the lung (24), raising the possibility that the regulation of IκBa expression by Yap/Taz may affect NF-κB transcriptional activity. To test this hypothesis, AECIIs from adult mouse lungs were cultured for 48 hours in the presence of a Yap shRNA lentivirus that inhibited Yap expression (25). Gene expression analysis showed that expression of Yap shRNA reduced the expression of Ikba in AECIIs compared with scrambled shRNA lentivirus–treated AECIIs (Supplemental Figure 5E). Analysis of NF-κB transcriptional activity showed that Yap shRNA–treated AECIIs had higher levels of NF-κB transcriptional activity compared with scrambled shRNA-treated or nontreated control AECIIs (Figure 6D). Moreover, AECIIs purified from Yap/Taz mutant mice at 7 dpi showed higher levels of NF-κB transcriptional activities than those from SPC-CreERT2 control mice (Figure 6E). In contrast, overexpression of Yap or Taz in AECIIs using YAP-5SA or TAZ-S89A adeno-associated virus type 6 (AAV6) led to increased Ikba expression and decreased NF-κB transcriptional activity compared with AAV6 GFP–treated AECIIs (Supplemental Figure 5, E and F). To determine whether Yap/Taz-induced IκBa expression affected NF-κB signaling, MLE-15 cells were transfected with the NF-κB response luciferase reporter. This study showed that expression of Yap shRNA led to increased transactivation of the NF-κB response luciferase reporter. In contrast, overexpression of Yap or Taz reduced the transactivation of the NF-κB response luciferase reporter (Figure 6F). These results indicated that Yap/Taz regulated NF-κB response through targeting IκBa in AECIIs (Figure 6G). IκBa overexpression promotes inflammatory resolution and alveolar epithelial regeneration in Yap/Taz mutant lung. To determine whether loss of IκBa expression was responsible for the prolonged inflammatory response and delayed alveolar epithelial regeneration observed in Yap/Taz mutant mice, we restored IκBa expression in Yap/Taz mutant lungs using the AAV6 vector for the expression of murine IκBa. We first evaluated whether intratracheal instillation of AAV6 led to efficient gene transfer and expression in AECIIs. Mouse lungs treated with AAV6-GFP at a dose of 2.5 × 1010 vp/animal exhibited abundant GFP staining in the lung alveolar region and much lower staining in the airway epithelium at 48 hours after instillation (Supplemental Figure 6A). In lung alveoli, 97.3% ± 1.1% of GFP+ cells were SPC+ and less than 2.5% of GFP+ cells were CD3+ (Supplemental Figure 6B). These results indicated that intratracheal instillation of AAV6 resulted in preferential transduction of SPC+ AECIIs in mouse lungs. Next, Yap/Taz mutant mice were administrated with AAV6-IκBa at a dose of 2.5 × 1010 vp/animal through intratracheal instillation at 7 dpi. Twenty-four hours later, AECIIs were purified and performed for gene expression analysis by qRT-PCR. IκBa overexpression in Yap/Taz mutant lungs restored IkBa transcription, while IkBa expression in inflammatory cells (CD45+) was unchanged (Supplemental Figure 6C). Restoration of IkBa expression in Yap/Taz mutant lungs was sufficient to inhibit NF-κB–mediated gene expression in AECIIs, since AAV6-IκBa–treated AECIIs exhibited decreased expression of NF-κB–dependent genes (Cxcl3, Ccl21a) compared with PBS treatment (Supplemental Figure 6C). We observed a significant decrease in IL-1b protein in BALF from AAV6-IκBa–treated Yap/Taz mutant mice at 14 dpi (Figure 7, A and B). Cytokine array showed that AAV6-IκBa–treated Yap/Taz mutant lungs at 14 dpi produced lower levels of cytokines, including cytokines associated with T cell accumulation (IL-1b, CXCL9) and antiinflammatory resolution responses (TIMP-1, TREM-1) (Figure 7C and Supplemental Figure 7, A and B). PBS-treated Yap/Taz mutant lungs showed results similar to those of AAV6-null–treated Yap/Taz mutant lungs and were used as the controls in this study (Supplemental Figure 7, A–C). The impairment in inflammatory resolution in Yap/Taz mutant lung was partially recovered by ectopic IκBa overexpression, as AAV6-IκBa treatment on Yap/Taz mutant mice led to a significant reduction in the number of CD3+ T cells in the lung at 14 dpi (Figure 7D). Inflammatory responses were not completely absent in AAV6-IκBa–treated Yap/Taz mutant lungs, however, as the numbers of neutrophils (Ly6G+) and macrophages (CD11c+CD64+) were equivalent between AAV6-IκBa–, PBS-, and AAV6-null–treated Yap/Taz mutant lungs (Supplemental Figure 7, D and E). Figure 7 Inflammatory resolution and alveolar epithelial recovery in Yap/Taz mutant lungs with AAV6-IκBa treatment. (A) Schematic of experimental design for B–K. (B) BALF was analyzed for IL-1b by ELISA assay (n = 4–5 per group). (C) Cytokine assay showed protein levels of IL-1b and CXCL9 in mouse lung lysates (n = 1 per group). (D) Flow cytometry of dissociated lung cells was performed by gating on CD3+CD45+ cells, and quantification of total number of CD3+CD45+ cells in the lung at 14 dpi were graphed (n = 5–6 per group). (E) Immunostaining on lung sections at 14 dpi with nuclei labeled by DAPI (blue) and antibody to T1a (red). White dashed lines indicate T1a– region in the lung. (F) Flow cytometry of dissociated lung cells was performed by gating on T1a+CD45– cells. (G) Quantification of the percentage of T1a+CD45– cells of total CD45– cells in the lung at 14 dpi (n = 4–5 per group). (H) Quantification of total protein in BALF at 14 dpi (n = 5–6 per group). (I) Lung sections at 14 dpi were stained with Alcian blue and nuclear fast red. (J) Quantification of lung fibrotic regions at 14 dpi using Ashcroft scoring method (n = 4–6 per group). (K) Quantification of collagen level on lung tissue lysates at 14 dpi using hydroxyproline assay (n = 4 per group). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001, 1-way ANOVA. Scale bars: 50 μm. We next assessed whether partial inflammatory resolution was sufficient to accelerate alveolar epithelial regeneration and regression of fibrotic lesions in Yap/Taz mutants. Analyses of lung sections by immunostaining and flow cytometry showed AAV6-IκBa–treated Yap/Taz mutant lungs exhibited significant improvement in the recovery of AECIs at 14 dpi compared with PBS- or AAV6-null–treated Yap/Taz mutant lungs (Figure 7, E–G, and Supplemental Figure 7C). Moreover, AAV6-IκBa treatment led to improved lung tissue integrity in Yap/Taz mutant mice, as gauged by the total protein measurement in their BALF (Figure 7H and Supplemental Figure 7C). The regenerated AECIs in AAV6-IκBa–treated Yap/Taz mutant lungs were not associated with the differentiation of preexisting SPC+ AECIIs into AECIs, since AECII-to-AECI differentiation (GFP+T1a+) in AAV6-IκBa–treated Yap/Taz mutant lungs was equivalent to that in PBS-treated Yap/Taz mutant lungs (Supplemental Figure 8, A–C), suggesting the recovered AECIs were contributed by non–lineage-labeled AECIIs. We measured Ashcroft scores in Trichrome-stained lung sections and collagen burden in lung tissue lysates at 14 dpi and found a significant decrease in lung fibrotic lesions in AAV6-IκBa–treated Yap/Taz mutant mice compared with PBS- or AAV6-null–treated mice (Figure 7, I–K, and Supplemental Figure 7C). Together, these findings indicated that restoration of IκBa expression in Yap/Taz mutant lung promoted inflammatory resolution and accelerated alveolar epithelial regeneration and regression of lung fibrotic lesions during bacterial pneumonia. Discussion Bacterial pneumonia has long been associated with lung damage, but how the host recovers from lung damage and the potential role of lung regeneration versus fibrosis have not been fully understood. Here, we investigated the recovery from bacterial pneumonia in mice infected by S. pneumoniae. We found that mice had massive tissue damage exclusively in lung alveoli during bacterial pneumonia. Mice recovering from pneumococcal infection lacked detectable lung fibrosis, suggesting that considerable regeneration of lung tissue must be acting during this recovery. Numerous studies have demonstrated alveolar epithelial regeneration after injury involving SPC+ AECIIs (3). We showed that SPC+ AECIIs increased proliferation within 7 dpi. We also showed that lineage-labeled SPC+ AECIIs differentiated to AECIs at sites of damaged lung alveoli during 7–14 dpi, coinciding with the time frame when inflammation was resolved in the lung. AECIIs synthesize and secrete surfactant and proliferate and differentiate into AECIs after injury in order to maintain the integrity of the alveolar wall. AECIIs also secrete cytokines and chemokines under appropriate stimulation, showing their role in modulating immunologic activity in the alveolar space. Given the important contributions of AECIIs in lung repair and regeneration, it is critical to determine the molecular mechanisms underlying AECII activities in intact tissues. We showed in this study that Yap and Taz expression and nuclear activity increased in AECIIs upon pneumococcal infection. Yap and Taz regulated AECII activities, including their proliferation, differentiation into AECIs, and inflammatory responses during both homeostasis and lung regeneration following bacterial pneumonia–induced lung injury. Yap and its closely related paralogue Taz are the transcriptional coactivators that are the main downstream mediators of the Hippo pathway. When Hippo signaling is inhibited, Yap and Taz (Yap/Taz) can accumulate in the nucleus where they interact with transcription factors such as the TEA domain family members (TEAD) and activate gene expression associated with cell survival, proliferation, and differentiation (16). Recent studies have shown the important role of the Hippo/Yap pathway in controlling lung epithelial progenitor cell differentiation during embryogenesis and basal stem cell maintenance in the upper airway of adult lung (26–28). Furthermore, studies of pneumonectomy-induced alveolar regeneration show that Yap is essential for AECII proliferation and differentiation in response to mechanical tension in the lung (29). We measured both AECII proliferation and AECII-to-AECI differentiation in the bacterial pneumonia mouse model. Our data are consistent with previous studies demonstrating the importance of Yap/Taz for the proliferation and differentiation of lung epithelial progenitor cells. This study demonstrates what we believe is a previously unknown function for Yap/Taz in negative regulation of AECII-derived inflammation. We found that Gram-positive pathogen Streptococcus stimulation of AECIIs to effect inflammatory cytokine/chemokine production required the activation of Yap/Taz-IκBa, a process that had been previously reported in Drosophila fat cells, both in vitro and in vivo (20). A potential mechanism for Gram-positive pathogen-dependent activation of Yap/Taz-IκB may be the TLR-mediated antimicrobial response (20). The inhibitory IκB proteins have been discovered as fundamental regulators of the inducible transcription factor NF-κB (30). IκBa preferentially binds to heterodimers containing p50, p65, and c-Rel (31–34). Several comparative studies have demonstrated that IκBa is the most efficient IκB molecule to remove NF-κB from target gene promoters and to terminate NF-κB–mediated gene induction (35–38). Generally, various stimuli such as TLR ligands triggers rapid IκBa degradation, resulting in the release of NF-κB dimers and subsequent transcriptional activation (30, 39, 40). We found IκBa was transcriptionally regulated by Yap/Taz-Tead in AECIIs. Thus, Yap/Taz-dependent functions are associated with the termination of NF-κB–dependent transcription of inflammatory genes through the induction of IκBa expression. Loss of Yap/Taz removed a “brake” on AECII activation and increased expression of inflammatory genes, leading to elevated local inflammation and enhanced accumulation and persistence of inflammatory cells in the lung (Supplemental Figure 9). Higher inflammatory responses in pneumococci-infected lungs have been previously reported (12, 13, 41), but prior work did not examine immune responses in vivo that could affect lung regeneration. Another study found increased AECII proliferation in mice following pneumonectomy, which was correlated with increased monocyte and macrophage recruitment and an increased inflammatory state in the lung (11). They further showed that loss of recruited monocytes/macrophages impaired lung regeneration (11). However, the impact of prolonged inflammation on lung regeneration has not been clear. There is a growing understanding that overly exuberant or persistent inflammatory responses can be found in fibrotic lungs (42–44). Our observations of increased IL-1b, Cxcl3, and Ccl21a expression in the fibrotic lungs of Yap/Taz mutants are consistent with a more inflammatory milieu during bacterial pneumonia. We found that SPC+ AECIIs required Yap/Taz to active IκBa, which inhibited NF-κB–mediated inflammation. Loss of Yap/Taz in SPC+ AECIIs led to persistent lung inflammation and fibrotic lesions in lung alveoli during bacterial pneumonia, indicative of impaired alveolar regeneration. We found no Yap/Taz-dependent defects in clearing pneumococcal load in vivo, leading us to the conclusion that it is a role for Yap/Taz in the resolution of lung inflammation through regulation of NF-κB nuclear activity that underlies alveolar regeneration. During development, epithelial NF-κB signaling is known to regulate lung inflammation and alveolar development in both humans and mice (45–47). In adulthood, correlations between persistent lung inflammation and decreased lung regeneration have been observed in both various animal injury models and human patients (8, 10), but it has not been clear whether higher inflammatory burdens in the lung precede loss of lung regeneration or are an effect of respiratory disease. In this work, we found that loss of Yap/Taz expression in SPC+ AECIIs, by promoting constitutive nuclear activity of NF-κB, led to persistent inflammatory responses in lung alveoli. We also found that restoration of the Yap/Taz target IκBa, the inhibitor of NF-κB, promoted resolution of lung inflammation and alveolar epithelial recovery from bacterial pneumonia in Yap/Taz mutant mice. Recent studies have reported alveolar regenerative capacity following injury, which is mediated by several stem/progenitor cell populations, including basal cells, club cells, AECIIs and lineage-negative progenitors or distal airway stem cells, and AECIs (10, 48–54). Our work showed that IκBa treatment did not improve AECII-to-AECI differentiation in Yap/Taz mutant lungs, suggesting that regenerated AECIs were contributed by non-SPC+AECIIs. These results emphasize the importance of cellular niche in considering how inflammatory signaling influences adult lung regeneration. The resolution of inflammation in the lung is a highly controlled and coordinated process that involves the suppression of inflammatory gene expression and inflammatory cell clearance. We highlight the role of the endogenous antiinflammatory mechanism of Yap/Taz that limits the excessive and prolonged production of inflammatory mediators in AECIIs. In general, the onset of inflammation is associated with secretion of cytokines/chemokines to attract circulating immune cells to the site of injury to counteract the infected agent in the affected lung tissue. The activated AECIIs directed the inflammatory processes toward resolution after the tasks of immune cells are successfully accomplished. However, in pathologic chronic inflammation, the inflammatory processes are not resolved, and active inflammation continues in a dysregulated fashion. The persistent lung inflammatory processes in Yap/Taz mutants suggest that inappropriate Yap/Taz and IκBa/NF-κB signaling pathways may also be the major contributors that cause failed regeneration in pathologic chronic lung disease. Methods Mice. All mice were bred and raised in the pathogen-free mouse facility at Temple University. C57BL/6 mice (Jackson Laboratories) and SPC-CreERT2 mice (52) and Yapfl, Tazfl (55), and Rosa26-mTmG mice (Jackson Laboratories) were used. Generation and genotyping of the SPC-CreERT2, Rosa26-mTmG, Yapfl, and Tazfl lines have been previously described (52, 55). SPC-CreERT2 mice, Yapfl/fl mice, Tazfl/fl mice and Rosa26-mTmG mice were kept on a mixed C57BL/6:129SVJ background. All experiments used 6- to 10-week-old male mice. Bacterial infection. The pneumococcal strain used was the clinical isolated strain SpT4 (56). SpT4 was stored at –80°C and grown in tryptic soy agar plus catalase (57 μgP/ml) under microaerophilic conditions for 14–16 hours at 37°C with 5% CO2, then subcultured and grown to an optical density of 0.5. The broth was centrifuged, and the bacteria were washed in sterile PBS and resuspended in 4 ml of sterile PBS immediately prior to infection. Mice were anesthetized using ketamine/xylazine mixture and infected i.n. with a dose of approximately 5 × 106 CFU in 30 μl of sterile PBS. Bacterial loads. Pneumococcal loads were determined by homogenization of lung tissue in sterile PBS and blood serum at 1, 4, 7, and 14 days after infection of the animals. Tissue homogenate (100 μl) or blood serum (50 μl) and further dilutions were plated on tryptic soy agar plates plus catalase (57 μgP/ml) for culture overnight at 37°C with 5% CO2, and the number of CFU was counted. Quantification of lung damage. Lung damage was assessed in BALF. Lungs were lavaged with 1 ml of PBS. BALF was recovered and centrifuged at 1,650 g for 3 minutes at 4°C and supernatant collected. Protein levels were quantified by Pierce BCA protein assay (Thermo Scientific) per the manufacturer’s instructions. All plates were read on a Safire II plate reader (Tecan). Tissue harvest and immunostaining for histology. Mice were euthanized by overdose of tribromoethanol (Avertin, 300 mg/kg) followed by cervical dislocation. After exposing the heart and lungs, the descending aorta was cut and a 25-gauge needle was inserted into the right ventricle to flush at least 10 ml of PBS until the color of lungs changed to pink/white. A tracheostomy was performed on the exposed trachea and a 23-gauge stub needle tip was placed into the trachea. The lungs were inflated with 4% paraformaldehyde (PFA) at a pressure of 20 cm H2O. The trachea was then tied off, and intact lungs were immersed in 4% PFA for 4 hours at 4°C. Lung lobes were separated and washed with cold PBS overnight. After ethanol dehydration, lungs were embedded in paraffin and sectioned at 6 μm. To perform immunohistochemical staining, slides were deparaffinized and rehydrated. Tissue sections were then incubated in citrate buffer (pH 6.0) for 20 minutes at 95–100°C, followed by permeabilization and blocking with 0.3% Triton X-100 and 5% goat serum or horse serum (for goat primary antibodies) in PBS for 1 hour at room temperature (r.t.). Primary antibodies were diluted in PBS and incubated overnight at 4°C. Secondary antibodies were diluted in PBS and incubated for 1 hour at r.t. Immunostainings for Yap and Taz protein were performed using a TSA Fluorescein Tyramide Amplification System. EdU staining, TUNEL staining, and DAPI nuclear staining were performed according to the manufacturers’ recommendations. Tissue sections were washed 3× with PBS between antibody incubations for 15 minutes each. Slides were mounted with Aqua-Poly/Mount (Polysciences Inc.), and images were captured on a Zeiss LSM 710 confocal microscope and a Nikon eclipse fluorescence microscope. Images were processed with ImageJ/FIJI (NIH). Quantitation of cell numbers was completed using at least 10 randomly selected images per animal. Reagents used in this study are provided in Supplemental Table 3. Lung dissociation for flow cytometry. Alveolar epithelial cells were isolated using previously described protocols (57). Briefly, after lungs were cleared of blood by perfusion with PBS, as described for the tissue harvest procedure above, 3 ml of dispase (25 U/ml, 37°C) was instilled into the lungs through the trachea followed with instillation of 0.5 ml of 1% low melting agarose into the lungs. Agarose was hardened by adding ice on top of the lungs for 2 minutes. Lung lobes were then separated and incubated in 1 ml of dispase for 6 minutes at 37°C. Each lung lobe was minced in DMEM containing penicillin G, streptomycin, l-glutamine, HEPES, and DNase I (120 U/ml), followed by a rotating incubation for 10 minutes at r.t. The cells were then filtered sequentially through 100 and 40 μm strainers and centrifuged. The cells were incubated with red blood cell lysis buffer for 1 minute on ice and washed with PBS containing 5% BSA and 0.5 M EDTA. For immune cell flow cytometry, lungs were dissociated with a collagenase/DNase I solution, as previously described (11, 12). Lung lobes were cut into small pieces by scalpel blade and then incubated in 5 ml of digestion solution containing collagenase type I (3 mg/ml), DNase I (80 U/ml), 10% FBS, and 2-ME (0, 055 mM) for 45 minutes at 37°C. Flow cytometry was performed on a BD LSR II and sorting was performed on a BD INFLUX. Immunostaining for flow cytometry. Dissociated cells were blocked with CD16/CD32 Fc receptor block at 1:100 for 20 minutes. After washing with PBS, cells were incubated with fluorophore-conjugated antibodies for 1 hour. For live cell staining, LIVE/DEAD Fixable Aqua Dead Cell Stain Kit was added along with fluorophore-conjugated antibody mix at a 1:1000 dilution. Cells were washed twice with PBS, placed in FACS buffer containing 1% BSA and 0.05% sodium azide in PBS, and then assessed using a LSR II (BD). Analysis was performed using FlowJo (10.4.2). Gene expression and microarray analysis. qPCR analysis was performed using TRIzol-isolated RNA, which was used to generate cDNA using random hexamer primers and SuperScript III RT. Primer sequences are listed in Supplemental Table 4. For microarray analysis, total RNA was extracted from 6- to 8-week-old SPC-CreERT2, Rosa26-mTmG GFP+ cells at 0 and 8 dpi, and SPC-CreERT2, Yapfl/fl, Tazfl/fl, Rosa26-mTmG GFP+ cells at 8 dpi, converted to cDNA and used on Affymetrix Mouse Gene 1.0 ST arrays. Data were analyzed using Affymetrix Microarray Suite 5.0, Significance Analysis of Microarray (SAM), and the Empirical Bayes Analyses of Microarrays (EBAM). Genes with 1.5-fold or greater changes over that of the experimental mean at P < 0.01 (ANOVA) were considered significant. All original microarray data were deposited in the NCBI’s Gene Expression Omnibus database (GEO GSE86027). Hierarchical clustering was performed using Gene Cluster 3.0. Gene Ontology (GO) associations and related P values were determined using the ToppGene Suite (58). (https://toppgene.cchmc.org) Western blot. For extraction of proteins from AECIIs, lungs were dissociated with a dispase solution, as described in the procedure of lung dissociation for flow cytometry above. AECIIs were further purified using magnetic beads and column isolation, as previously described (58). Briefly, cells were incubated with CD45 Microbeads for 15 minutes at 4°C and washed with cold column buffer (PBS containing 5% BSA and 0.5M EDTA). Cells were resuspended in cold column buffer and applied to MACS LS columns on a MACS separator (Miltenyi Biotec). The flow-through cells were blocked with FcR blocking reagent for 10 minutes at 4°C, followed by an incubation with binotinylated EpCAM antibody for 30 minutes at 4°C. Cells were washed and incubated with Streptavidin Microbeads for 20 minutes at 4°C. Cells were resuspended in cold column buffer and applied to MACS LS columns. The columns were then removed from the magnetic separator. The magnetically labeled AECIIs were collected by forcing 5 ml of cold column buffer into the column. After centrifugation, AECIIs were lysed using RIPA buffer containing protease and phosphatase inhibitors. For extraction of proteins from tissues, lungs were homogenized in RIPA buffer using a tissue grinder. Protein concentrations were determined using the BCA Protein Assay Reagent Kit (Bio-Rad). Protein extracts were analyzed on polyacrylamide gels (10% NuPAGE Bis-Tris Gel; Invitrogen) and transferred to nitrocellulose membrane (Bio-Rad). The blots were blocked in 5% BSA in TBS for 1 hour at r.t., followed by incubation with primary antibody diluted in block overnight at 4°C. The blots were washed 3× with TBS containing 0.1% Tween 20 (TBST) for 15 minutes and incubated with near-infrared fluorophore-conjugated secondary antibodies diluted in TBST for 1 hour at r.t. Signals were detected using the Odyssey imaging system (LI-COR Biosciences). Cell lines, transfections, and treatments. Mouse MLE-15 cells (abm Inc., catalog T0495) were grown in DMEM/F12 containing 2% FBS, 10 mM HEPES, and 10 nM hydrocortisone. HEK293T cells were purchased from ATCC and grown in DMEM containing 10% FBS. DNA transfections were done with X-tremeGENE HP DNA Transfection Reagent (Roche) in antibiotic-free medium according to the manufacturer’s instructions. Yap shRNA and scrambled shRNA were used as previously described (25). Lentiviral particles were prepared by transiently transfecting HEK293T cells with lentiviral vectors together with packaging vectors (pMD2-VSVG and psPAX2). Plasmids. Murine Yap was described previously (59) and subcloned into the pCMV-Tag3B vector. Murine Taz was expressed in the pCMV-Sport6 vector (Addgene plasmid 27318). YAP-5SA and TAZ-S89A plasmids were purchased from Addgene (plasmid 27371 and 32840). Murine Tead2 cDNA was purchased from OriGene (catalog MR223302) and subcloned into the pcDNA3.1(+) vector. NF-κB response luciferase reporter was purchased from Promega (catalog E8491). A 1900 bp region of the murine IκBa, gene enhancer (–8600 to approximately –6700 kb relative to the ATG site) was cloned into a pGL3 luciferase reporter vector (Promega, GenBank, catalog U47298). Point mutations within the IκBa, enhancer–luciferase reporter plasmid were generated using the QuikChange Site-Directed Mutagenesis Kit (Stratagene). Sequences of cloning primers are p

|
Scooped by
Gilbert C FAURE
March 7, 2019 3:01 AM
|
Streptococcus pneumoniae is a bacterial pathogen that causes various diseases of public health concern worldwide. Current pneumococcal vaccines target the capsular polysaccharide surrounding the cells.

|
Scooped by
Gilbert C FAURE
February 10, 2019 12:46 PM
|
Vaccines have saved millions of lives, but nobody likes getting a shot. That's why scientists are trying to develop oral vaccines for infectious diseases. But to be effective, the vaccine must survive digestion and reach ...
|

|
Scooped by
Gilbert C FAURE
August 9, 2020 6:46 AM
|
Mucosal surfaces provide a remarkably effective barrier against potentially dangerous pathogens. Therefore, enhancing mucosal immunity through vaccines—strengthening that first line of defense—hold...

|
Scooped by
Gilbert C FAURE
July 15, 2020 2:32 AM
|
Some experts say a vaccine puffed in the nose would be better at protecting people from infection. But nasal vaccines won’t be ready right away.

|
Scooped by
Gilbert C FAURE
May 29, 2020 9:08 AM
|
As we continue to struggle to contain the pandemic of our age, scientists are exploring vaccines that do not prevent initial infection but may prevent or limit viral replication and delay disease p...

|
Scooped by
Gilbert C FAURE
March 8, 2020 4:59 AM
|
Bovine Coronavirus Related terms: View all Topics Coronaviridae In Fenner's Veterinary Virology (Fifth Edition), 2017 CORONAVIRUSES of Cattle and Horses BOVINE CORONAVIRUS Bovine coronavirus infections are associated with three distinct clinical syndromes in cattle: calf diarrhea, winter dysentery (hemorrhagic diarrhea) in adult cattle, and respiratory infections in cattle of various ages, including the bovine respiratory disease complex (shipping fever) in feedlot cattle. Coronaviruses were first reported as a cause of diarrhea in calves in the United States in 1973, and since then they have been recognized worldwide in association with the three clinical syndromes. The economic impact of respiratory disease and calf diarrhea is considerable. Although many coronaviruses have restricted host ranges, betacoronaviruses such as bovine and SARS coronaviruses (Table 24.1) can infect other animal species, including wildlife. Bovine coronavirus is closely related to the human coronavirus OC43 that causes the common cold; indeed, OC43 has been proposed to represent prior zoonotic transmission of bovine coronavirus. Bovine coronavirus has also been shown to infect dogs subclinically and to infect turkey poults, leading to fecal virus shedding, diarrhea, seroconversion, and transmission to contact controls. Genetically and/or antigenically related bovine coronavirus variants have been isolated from dogs with respiratory disease, humans with diarrhea, and captive or wild ruminants with intestinal disease similar to winter dysentery of cattle. The latter include Sambar deer (Cerous unicolor), waterbuck (Kobus ellipsiprymnus), giraffe (Giraffa camelopardalis), and white-tailed deer (Odocoileus virgineanus). Bovine coronavirus has also been linked to enteric disease in South American camelids. Interestingly, the human enteric coronavirus and wild ruminant coronaviruses both infected and caused diarrhea in experimentally exposed gnotobiotic calves, and the inoculated calves were subsequently immune to infection with bovine coronavirus. Despite the different disease syndromes and apparent interspecies transmission of bovine coronavirus and its variants, only a single serotype of bovine coronavirus is recognized, and there is little sequence diversity between the wild ruminant coronaviruses and coronaviruses associated with the different disease syndromes in cattle. Furthermore, there are few common sequence differences to explain differences in host or tissue tropism. The host cell receptor for bovine coronavirus is sialic acid, which reflects the wide tropism of this virus and explains the presence of a HE gene in the virus. Clinical Features and Epidemiology Coronavirus-induced diarrhea commonly occurs in calves under 3 weeks of age after the decline of passively acquired antibodies, but disease can occur in calves up to 3 months of age. The severity of diarrhea and dehydration depends on the infecting dose as well as the age and immune status of the calf. Coinfections with other enteric pathogens such as rotavirus, torovirus, cryptosporidia, and enterotoxigenic or enteropathogenic E. coli are common; their additive or synergistic effects increase the severity of diarrhea. Calf coronavirus diarrhea is often seasonal, being more common in winter in part because of the increased stability of the virus in the cold. Bovine coronavirus has also been implicated as a cause of winter dysentery, a sporadic, acute enteric disease of adult cattle worldwide that is especially prevalent during winter months, as the name implies. Winter dysentery is characterized by explosive, often bloody diarrhea, accompanied by decreased milk production, depression, anorexia, and frequent respiratory signs. Morbidity rates range from 20% to 100% in affected herds, but mortality rates are usually low (1–2%). A similar winter dysentery syndrome associated with bovine coronaviruses variants occurs in captive and wild ruminants. This finding suggests that certain wild ruminants (deer, elk, caribou, etc.) that share common grazing areas with cattle could be a reservoir for coronavirus strains transmissible to cattle, or vice versa. Bovine coronavirus also causes mild respiratory disease (coughing, rhinitis) or pneumonia in 2–6-month-old calves. An epidemiologic study of calves from birth to 20 weeks of age confirmed both fecal and nasal shedding of coronavirus, with diarrhea prominent upon initial infection. The calves subsequently shed virus intermittently via the respiratory route, with or without signs of disease, suggesting that long-term mucosal immunity in the upper respiratory tract is ineffective in mediating virus clearance. As a consequence, coronavirus may recycle among cattle of all ages and regardless of their immune status, with sporadic nasal or fecal shedding from individual animals. Alternatively, new virus strains may be introduced when cattle from different sources are comingled, or from cohabiting wild ruminants. Since 1993, bovine coronavirus has been incriminated as a precipitating cause of the bovine respiratory disease (shipping fever) complex. Both respiratory and enteric shedding of bovine coronavirus are common in affected feedlot cattle, peaking shortly after arrival at feedlots. Since its discovery, bovine coronavirus repeatedly has been identified in the lungs of feedlot cattle that died with bovine respiratory disease complex. Most feedlot cattle also seroconvert to bovine coronavirus within 3 weeks of arrival. Importantly, studies suggest that cattle arriving at feedlots with high serum titers of bovine coronavirus antibody were less likely to shed virus or to develop shipping fever. This observation suggests a role for serum antibodies in protection, or as an indicator of recent infection and active immunity. Pathogenesis and Pathology Concurrent fecal and nasal virus shedding persists for up to 10 days after coronavirus infection of calves. Coronavirus antigen is commonly detected in epithelial cells of both the upper respiratory and intestinal tracts, and occasionally in the lung. The pathogenesis of coronavirus enteritis in calves is similar to that caused by rotavirus, with the notable exception of extensive involvement of the large intestine by coronavirus. Disease occurs most commonly in calves at about 1–3 weeks of age, when virus exposure increases and antibody titers in the dam’s milk begin to wane. The pathogenesis and consequences of enteric coronavirus infection of calves are similar to those described for transmissible gastroenteritis in piglets. The destruction of the mature absorptive cells lining the intestinal villi and mucosal surface in the large intestine leads to maldigestion and malabsorption, with rapid loss of water and electrolytes. The resultant hypoglycemia, acidosis, and hypovolemia can progress to circulatory failure and death, especially in very young animals. The pathogenesis and lesions of winter dysentery of dairy and beef cattle resemble those of calf diarrhea, but often with marked intestinal hemorrhage and extensive necrosis of cells within the crypts of the large intestinal mucosa. Nasal and fecal shedding is more transient (up to 4–5 days). The anorexia and depression seen in dairy cattle with winter dysentery may explain the precipitous and sometimes prolonged decrease in milk production. The cause of the acute and often voluminous bloody diarrhea in some cattle is unexplained. Both nasal and fecal shedding of bovine coronavirus can occur soon after cattle are transported to feedlots. Coronavirus infection is probably important in predisposing cattle entering feedlots to secondary bacterial infection that results in the characteristic shipping fever pneumonia—a severe, often fatal fibrinous bronchopneumonia caused by Mannheimia haemolytica biotype A, serotype 1 infection. Bovine coronavirus antigen also has been detected in epithelial cells of the upper (trachea, bronchi) and lower (terminal bronchioles and alveoli) respiratory tract of some affected cattle, but the precise role of coronavirus in precipitating the bovine respiratory disease complex awaits definitive characterization. Diagnosis Initially, the diagnosis of enteric bovine coronavirus infections was based on the detection of virus by electron microscopy. Cell culture isolation became a viable option when it was discovered that the virus could be grown when trypsin was added to the medium—virus replication is recognized by hemadsorption or cytopathogenic effects, and the presence of coronavirus is confirmed by diagnostic tests. An array of assays is now available for detection of bovine (or variant) coronaviruses in cell culture or diagnostic specimens such as feces or nasal swabs, including ELISAs that incorporate monoclonal antibodies for antigen capture, immune electron microscopy using hyperimmune antiserum, and RT-PCR using bovine coronavirus or pan-coronavirus-specific primers to detect viral RNA. The use of RT-PCR for detection of bovine coronavirus has significantly increased the detection of this agent, particularly in respiratory samples, and has also substantially increased the recognized period of virus shedding by infected animals. Postmortem diagnosis is performed on acute fresh or fixed respiratory or intestinal tissues using hyperimmune antisera or monoclonal antibodies for immunofluorescence or immunohistochemical tissue staining. Immunity, Prevention, and Control Passive Immunity to Enteric Bovine Coronavirus Infections in Calves Because coronavirus diarrhea occurs in young calves during the nursing period, maternal vaccination is required to provide immediate passive (lactogenic) immunity. Passive immunity to enteric viral infections in calves correlates with high levels of IgG1 antibodies in colostrum and milk. In ruminants, IgG1 antibodies are dominant in colostrum and milk and are selectively transported from serum. Most adult cattle are seropositive for antibodies to bovine coronavirus. Therefore, parenteral vaccination of mothers with adjuvanted inactivated bovine coronavirus vaccines effectively boosts IgG1 antibody titers in serum and mammary secretions, to provide enhanced passive immunity to calves. Immunity to Respiratory Bovine Coronavirus Infections The correlates of immunity to respiratory coronavirus infections in cattle are not clearly defined. The serum antibody titer to bovine coronavirus may be a marker for respiratory protection, as coronavirus-specific antibody titers and isotype (IgG1, IgG2, IgA) were correlated with protection of calves and feedlot cattle against subsequent occurrence of respiratory disease, pneumonia, or coronavirus shedding. However, it can be difficult to distinguish whether serum antibodies are correlates of protection, or whether they merely reflect prior enteric or respiratory coronavirus infection. Intranasal vaccination using live-attenuated enteric coronavirus vaccine has been proposed to reduce the risk of bovine respiratory disease complex (so-called “shipping fever”) in cattle entering feedlots. Respiratory System Jeff L. Caswell, Kurt J. Williams, in Jubb, Kennedy & Palmer's Pathology of Domestic Animals: Volume 2 (Sixth Edition), 2016 Bovine coronavirus. Bovine coronavirus (BCoV) is an important cause of enteric disease in young calves, and the same strains occasionally induce respiratory disease in calves 2-16 weeks of age. BCoV is a less frequent but nonetheless important cause of respiratory disease in calves, and deserves more attention than it has received in the past. Isolates from occurrences of respiratory and diarrheic diseases have similar genotypes, and a single outbreak may include both forms of disease. Respiratory signs include fever, serous nasal discharge, sneezing, and coughing. The virus replicates primarily in the nasal and tracheal epithelium, and occasionally in the lung. Many calves shed BCoV in their nasal secretions and/or seroconvert to BCoV in the first month after arrival in feedlots, but the association between seroconversion or virus shedding and increased risk of respiratory disease has been variable in the studies reported. Bronchiolar necrosis is the typical lesion seen in BCoV-infected calves (eFig. 5-83). Bronchiolar syncytia have been described in feedlot calves with concurrent BCoV infection and bacterial pneumonia, but the contribution of other viruses, such as bovine respiratory syncytial virus, to these lesions is uncertain. BCoV may be demonstrated using immunohistochemistry or RT-PCR, or isolated using specific rectal tumor cell lines. Further reading Bidokhti MR, et al. Tracing the transmission of bovine coronavirus infections in cattle herds based on S gene diversity. Vet J 2012;193:386-390. Fulton RW, et al. Bovine coronavirus (BCV) infections in transported commingled beef cattle and sole-source ranch calves. Can J Vet Res 2011;75:191-199. Park SJ, et al. Dual enteric and respiratory tropisms of winter dysentery bovine coronavirus in calves. Arch Virol 2007;152:1885-1900. Storz J, et al. Isolation of respiratory bovine coronavirus, other cytocidal viruses, and Pasteurella spp from cattle involved in two natural outbreaks of shipping fever. J Am Vet Med Assoc 2000;216:1599-1604. Storz J, et al. Coronavirus and Pasteurella infections in bovine shipping fever pneumonia and Evans' criteria for causation. J Clin Microbiol 2000;38:3291-3298. Diseases of the Respiratory System In Veterinary Medicine (Eleventh Edition), 2017 Bovine Coronavirus Bovine coronavirus (BoCV) is one of the more newly identified viral respiratory pathogens of cattle, being first described in 1993. As a consequence, the clinical significance of BoCV in bovine respiratory disease, and enzootic pneumonia in particular, is still being determined. The current evidence indicates that BoCV plays a primary and important role in enzootic pneumonia. BoCV was the most commonly identified viral pathogen identified in nasal swabs from calves with respiratory disease in Ireland, being present in 23% of calves.3 BoCV was identified throughout the year, but at a much lower rate in summer.3 BoCV was the only virus detected in approximately 75% of respiratory disease outbreaks in two to 3-month calves in Italy.4 Biology and Diseases of Ruminants (Sheep, Goats, and Cattle) Wendy J. Underwood DVM, MS, DACVIM, ... Adam Schoell DVM, DACLAM, in Laboratory Animal Medicine (Third Edition), 2015 Coronavirus Bovine coronavirus, of the family Coronaviridae, produces a more severe, long-lasting disease compared to rotavirus. Clinical signs in lambs and calves are similar to above, although the incubation period tends to be shorter (20–36 h). In addition, mild respiratory disease may be noted (Janke, 1989). Coronavirus infections may be complicated by parasite infestation (e.g., Cryptosporidia, Eimeria) or bacterial infections (e.g., E. coli, Salmonella). Treatment is aimed at correcting dehydration, electrolyte imbalances, and acidosis. Strict hygiene and effective passive transfer by developing good colostrum-management protocols are critical. Bovine vaccines are available both for delivery to pre-partum dams and for the neonate. Rotaviruses, coronavirus, and adenoviruses affect neonatal goats; however, little has been documented on the pathology and significance of these agents in this age group. Unlike calves, it appears that bacteria play a more important role in neonatal kid diarrheal diseases than in neonatal calf diarrheas. Parvovirus and BVDV also may cause diarrhea in neonatal calves. Viral Diseases of the Bovine Respiratory Tract Robert W. Fulton, in Food Animal Practice (Fifth Edition), 2009 BOVINE CORONAVIRUS Bovine coronaviruses (BCVs) were initially associated with neonatal calf diarrhea.15 Then BCVs were identified with “winter dysentery” in adult dairy cattle.7,15 Later BCVs were detected in respiratory secretions of infected calves with subsequent isolation from cattle with BRD signs. This isolation of BCVs from calves with “shipping fever” pneumonias led to the assumption that BCVs were a major etiology for BRD. In some studies other agents such as BRSV, BVDV, and PI-3V along with bacteria were also found in these severely ill cattle. No doubt BCVs are found in conjunction with other respiratory tract infections, yet their sole or primary BRD role has not been clearly established. Including BCV along with other bovine respiratory tract viruses contributing to BRD is best. Clearly, experimental reproduction of detectable and severe respiratory tract disease such as pneumonia would better make the case for BCV as a significant primary pathogen in respiratory tract disease in cattle. Etiology/Epidemiology BCVs are RNA viruses of the viral family Coronaviridae.6,15 They are enveloped viruses, thus sensitive to disinfectants and the environment. It is not unexpected that cattle would have a coronavirus with tropism for the respiratory tract. Coronaviruses infect the respiratory tract of other species including humans, pigs, turkeys, and chickens. BCV infections in cattle are worldwide. Initially implicated in neonatal calf diarrhea, BCVs were also reported with etiology in “winter dysentery” of adult cattle. Subsequently BCVs have been isolated from the nasal samples of cattle undergoing respiratory tract disease.62-66 Thus this virus has a purported role in both respiratory tract disease and enteric diseases. Only one serotype is recognized, but likely there is some antigenic variability.15 The dilemma for working with BCV experimentally and diagnostic laboratories attempting to isolate the virus is that BCV replicates poorly or is quite difficult to isolate in standard cell cultures. A specialized cell line, a human rectal adenocarcinoma line, is permissive for BCV and has been used for virus isolation from feces and nasal swabs by selected laboratories. The BCV is considered relatively common in enteric infections in both beef and dairy operations. The virus has been isolated from cells with disease including calf pneumonias, as well as beef cattle entering feedlots in various U.S. regions. The BCV was isolated from both healthy and sick cattle in these BRD episodes. And BCV was detected by seroconversions during the first month in feedlots in transported cattle. Clinical Disease The association of BCV with BRD has been primarily by the isolation of virus from nasal swabs of cattle with BRD signs and seroconversions to BCV. The virus has been found in healthy calves as well. Likewise, antibody testing has detected seroconversions in cattle in BRD cases. The clinical signs in the BRD cases are not unlike other BRD cases with viral etiologies present such as BVDV; PI-3V; BRSV; and other viruses with fever, nasal and ocular discharges, anorexia, and coughing. Typically these BCV isolations and seroconversions occur soon after arrival to the feedlot. As expected there is often involvement of secondary bacteria such as M. haemolytica and/or P. multocida. Attempts have been made to demonstrate the pathogenicity of BCV for the bovine respiratory tract. After experimental challenge in young calves, the virus could be found in feces of diarrheic calves and nasal swabs for up to 5 days.15 Respiratory disease signs occurred in only a few calves. Lesions of emphysema and interstitial pneumonia were evident in only a few calves.67,68 For other studies, there are mixed reports of BCV detected in lung tissues of cattle with BRD, one report with no BCV detection in lungs of cattle with BRD,6 and another detecting BCV antigen by immunofluorescence in respiratory tissues.69 Diagnosis The virus can be isolated in cell culture provided that a unique cell culture is available to the diagnostic laboratory, the human rectal adenocarcinoma line (HRT-18).15 The nasal swabs collected appear to be the choice of collections from live cattle for testing. An antigen capture ELISA originally used for detecting BCV antigen in fecal samples is also used by some diagnostic laboratories for BCV detection in respiratory disease samples. Also, BCV immunofluorescence is available to detect BCV antigen. Selected diagnostic laboratories and research units have used PCR to detect BCV in diagnostic samples. Use of electron microscopy could detect BCV in respiratory samples similar to the use of EM for fecal samples. Selected research laboratories have used ELISA tests for BCV antibodies, and in some selected studies they found seroconversions when paired samples were available. Clinicians should consult with their respective diagnostic laboratory for their testing for BCV. Prevention and Control Although there are licensed BCV vaccines for enteric disease protection, there are no licensed BCV vaccines in the United States to control respiratory tract disease in cattle. Treatment focuses on the use of antimicrobials to control the bacterial secondary infections. As in prevention of the neonatal enteric disease, it is assumed that adequate colostrum is available to provide protection in the young calf. Diseases of the Alimentary Tract–Ruminant In Veterinary Medicine (Eleventh Edition), 2017 Synopsis Etiology Bovine coronavirus. Epidemiology Northern climates. Adult lactating dairy cows, usually during winter months when housed. Immunity develops and lasts variable periods. High morbidity with outbreaks; low mortality. Transmitted by fecal–oral route. Signs Sudden onset of diarrhea affecting almost entire herd within several days. Mild fever, decline in milk production, inappetence. Recover in few days. Some coughing. Clinical pathology None routinely. Lesions Crypt atrophy on intestinal mucosa; enterocolitis. Diagnostic confirmation Detection of virus in feces. Serology. Treatment None required. Control No specific control measures available. Hygiene. Minimize overcrowding in dairy housing. Alimentary System Francisco A. Uzal, ... Jesse M. Hostetter, in Jubb, Kennedy & Palmer's Pathology of Domestic Animals: Volume 2 (Sixth Edition), 2016 Bovine coronavirus. In neonatal calves, Bovine coronavirus (BCoV) infection is a common cause of diarrhea, either alone or in combination with other agents, particularly Rotavirus and Cryptosporidium. The disease may be severe in combination with BVDV infection. BCoV is capable of infecting absorptive epithelium in the full length of the small intestine, and in the large bowel. Viral antigen is also found in macrophages in the lamina propria of villi and in mesenteric lymph nodes. In field infections, microscopic lesions are found most consistently in the lower small intestine and colon. Calves with BCoV infection usually develop mild depression, but continue to drink milk despite developing profuse diarrhea. With progressive dehydration, acidosis, and hyperkalemia, the animals become weak and lethargic; death can ensue as a result of hypovolemia, hypoglycemia, and potassium cardiotoxicosis. Diarrhea in survivors resolves in 5-6 days. At autopsy, affected animals have the nonspecific lesions of undifferentiated neonatal calf diarrhea. Rarely, mild fibrinonecrotic typhlocolitis is recognized in calves with coronaviral infection. Mesenteric lymph nodes may be somewhat enlarged and wet. Virus replication is cytocidal and initially occurs throughout the length of the villi in all levels of the small intestine, eventually spreading throughout the large intestine up to the end of the large colon and rectum, causing a malabsorptive diarrhea. Large concentration of BCoV can be typically found in the spiral colon. Infected epithelial cells die, slough off, and are replaced by immature cells. The microscopic lesions of coronaviral infection in calves vary with the severity and duration of the infection; villus atrophy in combination with mild colitis is typical (Fig. 1-114). In the calf small intestine, villus atrophy is rarely as severe as that seen in neonatal swine with TGE. Rather, villi are moderately shortened, or have subtotal atrophy with stumpy, club-shaped, or pointed tips, and villus fusion may be common. In the early phase of the clinical disease, villi are often pointed and covered by cuboidal to squamous epithelium. Exfoliation of epithelium and microerosion may be evident. Later, the epithelium is cuboidal to low columnar, basophilic, with irregular nuclear polarity and an indistinct brush border. Cryptal epithelium is hyperplastic. The lamina propria may contain a moderate infiltrate of mainly mononuclear inflammatory cells, some of which may have pyknotic or karyorrhectic nuclei. In the early stages of infection, necrosis of cells in mesenteric lymph nodes is associated with viral replication. Peyer's patches in animals examined after 4-5 days of clinical illness often appear involuted, and are dominated by histiocytic cells. Whether this is the result of viral activity or the effect of endogenous glucocorticoids is unclear. In the colon during the early phase of infection, surface epithelium may be exfoliating, flattened, and squamous or eroded in patchy areas. Some colonic glands may be dilated, lined by flattened epithelium and contain exfoliated cells and necrotic debris. A moderate mixed inflammatory reaction is present in the lamina propria, and neutrophils may be in damaged glands or effusing into the lumen through superficial microerosions. Later in infection, some dilated debris-filled colonic glands will remain, but other glands will be lined by hyperplastic epithelium, and the surface epithelium will be restored to a cuboidal or low columnar cell type. Goblet cells are usually relatively uncommon. Colonic lesions may be recognizable in tissues from animals submitted dead, even though postmortem change has obscured changes in the small intestine. Live calves in the early stages of clinical disease are the best subjects for confirmation of an etiologic diagnosis. In calves becoming ill <7 days of age, enterotoxigenic Escherichia coli is the main alternative diagnosis. Rotavirus, Cryptosporidium, and combined infections must be considered in calves 5-15 days of age. Infectious bovine rhinotracheitis, salmonellosis, and bovine viral diarrhea must also be considered. Both salmonellosis and bovine viral diarrhea may be associated with depletion of Peyer's patches and colitis that can be confused with that of coronaviral infection; neither is common in the strictly neonatal age group (<7-14 days of age). Respiratory tract infection also occurs in calves and feeders infected with BCoV. The virus replicates in the epithelium of the nasal turbinates and tracheobronchial tree, and respiratory infection may precede, be concurrent with, or follow enteric infection. Calf pneumonia caused by BRCoV can be observed in calves 6-9 months of age. Affected animals may develop fever, serous to mucopurulent nasal discharge, coughing, tachypnea, and dyspnea. Respiratory infections may play a role in maintaining the virus within a herd, and significant, but poorly characterized, pneumonia has been reported in some experimentally infected calves. In addition, coronaviral infection may predispose to subsequent respiratory bacterial infections or contribute to more severe respiratory disease as part of the shipping fever syndrome. Virus may be identified in tissue or nasal secretions by immunofluorescence or immunohistochemistry. Winter dysentery is a syndrome in adult cattle that has been associated with BCoV in a number of areas around the world. Animals develop blood-tinged diarrhea, nasolacrimal discharge or cough, anorexia, and drop in milk production. Mortality is rare, but may occur. The disease is characterized by a high morbidity rate ranging from 50-100%, but usually low mortality rate, typically <2%. Winter dysentery outbreaks are predominantly seen in young postpartum dairy cows, which then experience a drop of 25-95% in milk production. Occasional cases are also observed in adult dairy and beef cattle. Despite its name, cases of winter dysentery can be observed, albeit infrequently, during the warmer season. The pathophysiologic characteristics of winter dysentery are mostly attributed to lesions of the colonic mucosa. Grossly, the colon of affected animals has linear congestion and hemorrhage along the crests of mucosal folds and there may be a large amount of blood mixed with colonic contents (Fig. 1-115). The histologic lesions are similar to those seen in calves with classical BCoV diarrhea, although they are mostly restricted to the colon with only occasional lesions seen in the terminal small intestine. Large amount of BCoV can be detected in colonic epithelium by immunohistochemistry. Coronaviruses are commonly demonstrated in the feces of cattle with winter dysentery; seroconversions occur, and seroprevalence increases in affected herds. Coronavirus antigen is found in the colonic glands of affected animals, in which there is necrosis and exfoliation of epithelial cells. Certain management practices, notably housing animals in stanchions and use of equipment that handles both manure and feed, have been associated with the development of winter dysentery. Further reading Blanchard PC. Diagnostics of dairy and beef cattle diarrhea. Vet Clin North Am Food Anim Pract 2012;28:443-464. Boileau MJ, et al. Bovine coronavirus associated syndromes. Vet Clin North Am Food Anim Pract 2010;26:123-146. Cho KO, et al. Detection and isolation of coronavirus from feces of three herds of feedlot cattle during outbreaks of winter dysentery-like disease. J Am Vet Med Assoc 2000;217:1191-1194. Heckert RA, et al. Epidemiologic factors and isotype-specific antibody responses in serum and mucosal secretions of dairy calves with bovine coronavirus respiratory tract and enteric tract infections. Am J Vet Res 1991;52:845-851. Kanno T. Bovine coronavirus infection: pathology and interspecies transmission. J Disast Res 2012;7:293-302. Kapil S, et al. Experimental infection with a virulent pneumoenteric isolate of bovine coronavirus. J Vet Diagn Invest 1991;3:88-89. Natsuaki S, et al. Fatal winter dysentery with severe anemia in an adult cow. J Vet Med Sci 2007;69:957-960. Park SJ, et al. Dual enteric and respiratory tropisms of winter dysentery bovine coronavirus in calves. Arch Virol 2007;152:1885-1900. Saif LJ, et al. Winter dysentery in dairy herds: electron microscopic and serological evidence for an association with coronavirus infection. Vet Rec 1991;128:447-449. Smith DR, et al. Epidemiologic herd-level assessment of causative agents and risk factors for winter dysentery in dairy cattle. Am J Vet Res 1998;59:994-1001. Traven M, et al. Experimental reproduction of winter dysentery in lactating cows using BCV—comparison with BCV infection in milk-fed calves. Vet Microbiol 2001;81:127-151. Zhang Z, et al. Application of immunohistochemistry and in situ hybridization for detection of bovine coronavirus in paraffin-embedded, formalin-fixed tissues. J Clin Microbiol 1997;35:2964-2965. Infectious Diseases of the Gastrointestinal Tract Simon F. Peek, ... Kevin J. Cummings, in Rebhun's Diseases of Dairy Cattle (Third Edition), 2018 Etiology Based on seroprevalence studies, the bovine coronavirus (BCoV) responsible for calf diarrhea is quite prevalent in U.S. cattle herds, as is rotavirus. There is much debate among researchers at this point as to whether BCoV isolates obtained from calf and adult diarrhea cases are the same virus or distinct from those that have been incriminated in respiratory disease outbreaks in feedlot and dairy calves. Whether or not there are antigenic or genomic differences in BCoV strains that mediate different organ tropism is similarly unclear. Winter dysentery in adult cattle has been associated with BCoV, and the same strain that causes diarrhea in calves has been used to experimentally create winter dysentery in adult cattle. Therefore, the upper age limit of susceptibility to infection by this agent is apparently longer than traditionally thought. Although not as common as rotavirus as a cause of viral enteritis in dairy calves, coronavirus has been identified in neonatal calf diarrhea outbreaks, especially in the winter months and with mixed infections. A number of studies indicate that clinical disease associated with BCoV in calves is more severe than rotavirus, with higher mortality rates. Affected calves tend to be slightly older than calves infected with pure ETEC or pure rotavirus. They average 7 to 10 days of age at onset, with some observed as late as 3 weeks of age. The virus causes a severe enterocolitis characterized by villous enterocyte destruction in the small intestine and destruction of both ridges and crypts in the large intestine. Maldigestion, malabsorption, and inflammation all contribute to the pathophysiology of coronavirus diarrhea in calves. The virus is cytolytic, and affected villous enterocytes in the small intestine are replaced by cuboidal cells from the crypts, but the colonic lesions leave denuded mucosa in affected areas of the colon. The severity of this damage helps explain why coronavirus enteritis, unlike rotavirus, may cause some flecks of blood to appear in the stool and can kill calves even in a germ-free isolation facility. Thus, in the natural setting, coronavirus enteritis creates a severe clinical diarrhea and can also be associated with > 50% mortality when combined with other viral, bacterial and C. parvum infections.

|
Scooped by
Gilbert C FAURE
December 9, 2019 1:32 PM
|
Insufficient T-cell responses contribute to the increased burden of viral respiratory disease in infancy. Neonatal dendritic cells (DCs) often provide defective activation of pathogen-specific T cells through mechanisms that are incompletely understood, which hinders vaccine design for this vulnerable age group. Enhancing our characterization of neonatal DC sub-specialization and function is therefore critical to developing their potential for immunomodulation of T-cell responses. In this study, we engineered respiratory syncytial virus (RSV) to express a model protein, ovalbumin, to track antigen-presenting DCs in vivo. We found that murine neonatal conventional DC1s (cDC1s) efficiently migrated and presented RSV-derived antigen, challenging the paradigm that neonatal DCs are globally immature. In a key observation, however, we discovered that during infection neonatal cDC1s presenting viral antigen were unable to upregulate costimulatory molecules in response to type I interferons (IFN-I), contributing to poor antiviral T-cell responses. Importantly, we showed that the deficient response to IFN-I was also exhibited by human neonatal cDC1s, independent of infection. These findings reveal a functionally distinct response to IFN-I by neonatal cDC1s that may leave young infants susceptible to viral infections, and provide a new target for exploration, in light of failed efforts to design neonatal RSV vaccines.

|
Scooped by
Gilbert C FAURE
November 26, 2019 12:35 PM
|
It's November, a time when breeders concentrate on getting their broodmares in top health for foaling and breeding season and horsemen ship yearlings to trainers for breaking and introduction to race training.

|
Scooped by
Gilbert C FAURE
November 2, 2019 4:08 PM
|
The generation of tissue-resident memory T cells (TRM) is an essential aspect of immunity at mucosal surfaces, and it has been suggested that preferential generation of TRM is one of the principal advantages of mucosally administered vaccines.

|
Scooped by
Gilbert C FAURE
September 18, 2019 2:26 PM
|
Respiratory syncytial virus (RSV) is the most prevalent viral etiological agent of acute respiratory tract infection. Although RSV affects people of all ages, the disease is more severe in infants and causes significant morbidity and hospitalization in young children and in the elderly. Host factors, including an immature immune system in infants, low lymphocyte levels in patients under 5 years old, and low levels of RSV-specific neutralizing antibodies in the blood of adults over 65 years of age, can explain the high susceptibility to RSV infection in these populations. Other host factors that correlate with severe RSV disease include high concentrations of proinflammatory cytokines such as interleukins (IL)-6, IL-8, tumor necrosis factor (TNF)-α, and thymic stromal lymphopoitein (TSLP), which are produced in the respiratory tract of RSV-infected individuals, accompanied by a strong neutrophil response. In addition, data from studies of RSV infections in humans and in animal models revealed that this virus suppresses adaptive immune responses that could eliminate it from the respiratory tract. Here, we examine host factors that contribute to RSV pathogenesis based on an exhaustive review of in vitro infection in humans and in animal models to provide insights into the design of vaccines and therapeutic tools that could prevent diseases caused by RSV.

|
Scooped by
Gilbert C FAURE
August 18, 2019 2:11 PM
|
KEY POINTS Trout have limited IgM and IgT repertoire diversity in NALT. Intranasal vaccination in trout triggers systemic and mucosal Ig response. IgM and IgT respond to i.p. and intranasal bacterin vaccination. Abstract Bony fish represent the most basal vertebrate branch with a dedicated mucosal immune system, which comprises immunologically heterogeneous microenvironments armed with innate and adaptive components. In rainbow trout (Oncorhynchus mykiss), a nasopharynx-associated lymphoid tissue (NALT) was recently described as a diffuse network of myeloid and lymphoid cells located in the olfactory organ of fish. Several studies have demonstrated high levels of protection conferred by nasal vaccines against viral and bacterial pathogens; however, the mechanisms underlying the observed protection are not well understood. We applied 5′RACE and a deep sequencing–based approach to investigate the clonal structure of the systemic and mucosal rainbow trout B cell repertoire. The analysis of Ig repertoire in control trout suggests different structures of IgM and IgT spleen and NALT repertoires, with restricted repertoire diversity in NALT. Nasal and injection vaccination with a bacterial vaccine revealed unique dynamics of IgM and IgT repertoires at systemic and mucosal sites and the remarkable ability of nasal vaccines to induce spleen Ig responses. Our findings provide an important immunological basis for the effectiveness of nasal vaccination in fish and other vertebrate animals and will help the design of future nasal vaccination strategies. Footnotes This work was supported by USDA AFRI Grant 2DN70-2RDN7 (to I.S.). S.M. also received funding from the People Programme (Marie Curie Actions) of the European Union’s Seventh Framework Programme (FP7/2007-2013) under REA Grant Agreement 600391. The dataset presented in this article has been submitted to the National Center for Biotechnology Information (https://www.ncbi.nlm.nih.gov/sra/) under accession number PRJNA551127. The online version of this article contains supplemental material. Received February 7, 2019. Accepted July 9, 2019. Copyright © 2019 by The American Association of Immunologists, Inc.

|
Scooped by
Gilbert C FAURE
May 29, 2019 2:42 PM
|
Vaxart, Inc.(Nasdaq: VXRT), a clinical-stage biotechnology company developing oral recombinant vaccines that are administered by tablet rather than by inj...

|
Scooped by
Gilbert C FAURE
March 19, 2019 3:39 PM
|
Recently, the risk of viral infection has dramatically increased owing to changes in human ecology such as global warming and an increased geographical movement of people and goods. Howev-er, the efficacy of vaccines and remedies for infectious diseases ...

|
Scooped by
Gilbert C FAURE
February 28, 2019 11:52 AM
|
The effect of the microbiome on the response to vaccines was the topic of the last talk of the "Microbiome Club", an initiative of the IMIM and Hospital de Mar.
|
 Your new post is loading...
Your new post is loading...
 Your new post is loading...
Your new post is loading...