Your new post is loading...
 Your new post is loading...

|
Scooped by
I2BC Paris-Saclay
Today, 5:04 AM
|
Iron Sulfides Produced by Thermococcales: An Iron Detoxification Mechanism
Thermococcales, hyperthermophilic archaea from hydrothermal vents, promote iron sulfide precipitation, enabling survival in iron-rich environments. Some cells become encrusted in pyrite and do not survive mineralization, while surviving cells activate metal detoxification genes. Thermococcales, sulfur-reducing archaea inhabiting the hottest parts of hydrothermal vents, have evolved to thrive in environments rich in iron and sulfide species. In this study, using experimental analogues of sulfur-rich hydrothermal chimneys, we confirm previous suggestions that the precipitation of iron sulfide minerals promoted by Thermococcales contributes to a population-wide adaptation to reactive species induced by the presence of high levels of iron. In parallel with mineral phases identification, cellular metabolic activity was monitored during mineralization, revealing a mechanism in which a subpopulation of cells does not survive mineralization and becomes encrusted in pyrite, while the remaining living cells exhibit a gene expression profile focused on DNA repair and metal excess associated detoxification. Compared to abiotic conditions, Thermococcales induce a faster precipitation of dissolved iron, immobilising excess metal. Our results clarify the role of mineralizing cells in this survival mechanism, suggesting that this biomineralization process allows resilience to extreme chemical stress. Upon drastic levels of toxic dissolved iron, thanks to a population of mineralizing cells, the surviving Thermococcales are thus more likely to endure those still harsh environments. This complex mechanism is likely a key factor in the adaptation of microorganisms to the hottest environments of hydrothermal vents. More information : https://enviromicro-journals.onlinelibrary.wiley.com/doi/10.1111/1462-2920.70242 Contact : Aurore Gorlas aurore.gorlas@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
Today, 4:52 AM
|
The tRNA moieties of both aminoacyl-tRNA substrates of a cyclodipeptide synthase share a common binding site, as revealed by RNA microhelices mimicking tRNA acceptor arms
Cyclodipeptide synthases (CDPSs) utilize two aminoacyl-tRNAs as substrates to produce diverse natural products. Here, we demonstrate that CDPSs efficiently recognize aminoacylated microhelices (miHxs) that mimic the tRNA acceptor arm. Structural and enzymological analyses using unacylated, misacylated, and engineered miHxs reveal a shared RNA-binding mode for both substrates. These findings establish miHxs as versatile tools to investigate CDPS function and, more broadly, other aminoacyl-tRNA–dependent enzymes. Two teams from the I2BC, in collaboration with the ICSN, combined enzymological and structural approaches to investigate cyclodipeptide synthases (CDPSs), enzymes involved in natural product biosynthesis. CDPSs sequentially use two aminoacyl-tRNAs (AA-tRNAs) to catalyse cyclodipeptide formation. We previously showed that microhelices (miHxs), mimicking the tRNA acceptor arm, are as efficient as full-length AA-tRNAs when aminoacylated by flexizymes.
Here, we generated a diverse set of miHxs (acylated, unacylated, misacylated, mutated, or shortened) and analysed their interactions with CDPSs. We focused on the Nocardia brasiliensis CDPS (Nbra-CDPS), which synthesizes cyclo(L-Ala–L-Glu) from Ala-tRNAAla and Glu-tRNAGlu. Crystal structures of Nbra-CDPS bound to analogues of its first substrate, including unacylated and acylated miHxAla, were determined. Cryo-EM analysis confirmed that miHxs mimic the acceptor stem of full-length tRNAs.
We also solved the structure of Nbra-CDPS bound to unacylated miHxGlu, an analogue of the second substrate, and found that it superimposes well with miHxAla despite sequence differences. Together with results obtained using misacylated substrates, these data reveal a shared RNA-binding mode for both substrates. Our findings establish miHxs as powerful tools to dissect CDPS function and to study other AA-tRNA–dependent enzymes. More information : https://academic.oup.com/nar/article-abstract/doi/10.1093/nar/gkag307/8625897?utm_source=authortollfreelink&utm_campaign=nar&utm_medium=email Contact : Muriel Gondry muriel.gondry@i2bc.paris-saclay.fr
Les 3èmes journées de rencontre du GDR Imabio se tiendront à l’ENS Paris-Saclay du 9 au 12 juin 2026. - Registration is free, but seats are limited (approximately 150 for conferences, 50 for workshops).
- Abstract submission deadline: 26th April 2026
- Registration deadline: 11th May 2026
- Inscription et programme
Via Life Sciences UPSaclay

|
Scooped by
I2BC Paris-Saclay
March 25, 5:30 AM
|
Inspiring the Next Generation of Scientists at Collège A. Fournier, Orsay
Researchers from the I2BC recently visited Collège A. Fournier in Orsay to introduce students to various scientific career paths. Magali Noiray presented the institute’s state-of-the-art facilities, while Olivier Namy showcased the exciting work of its research teams. Science is for everyone, and the journey can start at any age. Let’s inspire the next generation of innovators! More information about I2BC: https://www.i2bc.paris-saclay.fr/ A great thanks to Magali & Olivier for their time and commitment!
L’autophagie est un mécanisme de dégradation des composants cellulaires, indispensable à la survie de la cellule. Mais ses fonctions vont au-delà du nettoyage ou de la survie en conditions de stress : elle est impliquée dans l’immunité, les maladies neurodégénératives, le vieillissement, le cancer… Avec - Audrey Esclatine, professeure à l’Université Paris-Saclay, co-responsable de l’équipe Autophagy and antiviral immunity de l’I2BC (CNRS/CEA/UPSaclay, Gif-sur-Yvette), présidente du Club Francophone de l'Autophagie
- Pierre-Emmanuel Joubert, maître de conférences à Sorbonne Université en immunologie
- Flavie Strappazzon, chargée de recherche CNRS à l’Institut NeuroMyoGène
L'autophagie ("se manger soi-même") est un processus d'autodigestion essentiel au maintien de l'homéostasie cellulaire. Il s'agit d'une dégradation de constituants cellulaires par la voie lysosomale en composants recyclés. Réécouter le podcast de l'émission du 3 mars 2026 -> Contact : audrey.esclatine@i2bc.paris-saclay.fr
Via Life Sciences UPSaclay

|
Scooped by
I2BC Paris-Saclay
March 9, 8:48 AM
|
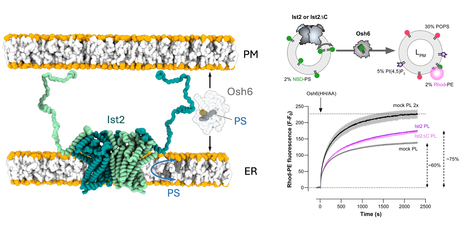
Phosphatidylserine dynamics between the endoplasmic reticulum and the plasma membrane in Saccharomyces cerevisiae.
A functional partnership between a lipid scramblase and a lipid transfer protein in the regulation of phosphatidylserine homeostasis. Phosphatidylserine is synthesised in the endoplasmic reticulum (ER), but this negatively charged lipid is highly concentrated in the plasma membrane (PM). There, it plays numerous roles in processes such as cell signalling and cell fusion, and it can also mediate apoptosis and synaptic pruning when exposed in the outer leaflet of the PM. In the yeast Saccharomyces cerevisiae, selective PS transport between the ER and the PM by the lipid transfer protein Osh6 is required for PS enrichment in the plasma membrane. Osh6 operates at membrane contact sites (MCS), where it interacts with the Ist2 protein embedded in the ER, tethering the ER to the PM via its long, disordered C-terminal tail. In two studies conducted in collaboration with the CRBM (CNRS/University of Montpellier), the IPMC (CNRS/University of Nice Côte d'Azur), and the IBCP (CNRS/University of Lyon), we show that Ist2 is a membrane transporter that catalyses rapid lipid exchange across the two leaflets of the ER – acting as a lipid ‘scramblase’ – and that this scrambling activity sustains Osh6-mediated PS transfer between the ER and the PM. First, we demonstrated that Ist2 catalyses the scrambling of different lipids in vitro after reconstituting purified Ist2 in proteoliposomes. Molecular dynamics simulations were then used to identify a cavity through which the lipid headgroup passes during transport from one leaflet to another. Further cellular studies revealed a close relationship between the COPII complex and the Ist2 protein. This was manifested by impaired yeast growth and vacuolar trafficking, as well as disruption of ER exit sites. We also found that Ist2 deletion stimulates the formation of ER-derived lipid droplets and changes their composition. Finally, using artificial, reconstituted ER-PM contact sites, we demonstrated that Ist2-mediated lipid scrambling sustains Osh6-mediated lipid transfer. Together, our studies identify Ist2 as a lipid scramblase and establish that lipid scrambling in the ER by Ist2 controls various cellular functions, such as vesicular transport and lipid droplet homeostasis. This highlights the importance of lipid dynamics for ER function. Furthermore, we reveal a functional partnership between Ist2-mediated lipid scrambling and Osh6-mediated lipid transfer at MCS. More information : https://doi-org.insb.bib.cnrs.fr/10.1083/jcb.202502112 https://www-science-org.insb.bib.cnrs.fr/doi/10.1126/sciadv.adz2217 Contact : Guillaume Lenoir guillaume.lenoir@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
February 16, 10:02 AM
|
Snapshots of cotranslational N-myristoylation reveal NMT as a ribosome-associated chaperone.
NMT in a new light: associated with the ribosome via the NAC complex, this enzyme not only adds lipid tags to the N-termini of nascent proteins but also acts as a chaperone, cooperating with MetAPs to ensure proper folding and delivery. N-myristoylation is an essential cotranslational lipid modification catalyzed by N-myristoyltransferases (NMTs). Structural and cellular analyses reveal that NMT1 associates with the ribosomal tunnel exit via the nascent polypeptide–associated complex (NAC) and acts sequentially after MetAP-mediated initiator methionine removal, in contrast to previously described simultaneous cotranslational modification assemblies. Unexpectedly, NMT1 also exhibits chaperone-like activity, expanding its functional repertoire in cotranslational protein biogenesis. More information : https://www-nature-com.insb.bib.cnrs.fr/articles/s41467-025-67962-4 Contact : Thierry Meinnel and Carmela Giglione thierry.meinnel@i2bc.paris-saclay.fr and carmela.giglione@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
February 9, 3:57 AM
|
Cytometry workshop (March 2026)

|
Scooped by
I2BC Paris-Saclay
January 13, 3:52 PM
|
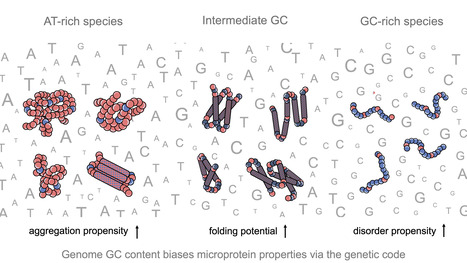
Impact of GC content on de novo gene birth
Noncoding DNA can generate microproteins, some of which evolve into new genes. We show that de novo genes preferentially originate from GC-rich, foldable sequences, revealing how base composition channels the birth of new proteins. Noncoding regions of eukaryotic genomes are widely transcribed and constitute a major source of novel microproteins, some of which eventually become fixed as de novo genes - a process known as de novo gene birth that plays a significant role in species adaptation. However, the structural properties of these nascent proteins and the factors governing their evolutionary fate remain poorly understood. In particular, the role of genome nucleotide composition (GC content) in shaping their biophysical properties has remained unclear. In this study, researchers of the I2BC, analyzed the foldability and sequence properties of millions of putative microproteins encoded by intergenic open reading frames (ORFs) from 3,379 eukaryotic species spanning a broad range of GC contents (18–79%). Results show that GC content strongly influences amino-acid composition and structural tendencies, suggesting distinct cellular impacts if non-genic regions are pervasively expressed. AT-rich species predominantly encode ORFs biased toward hydrophobic, aggregation-prone sequences, whereas GC-rich species tend to encode more hydrophilic, disorder-prone ORFs. ORFs from genomes with intermediate GC content display a more balanced composition and higher folding potential, with many expected to adopt proto-folds. To assess how these properties relate to gene emergence,the authors traced the evolutionary history of several hundred de novo proteins across 22 species using phylostratigraphy, targeted de novo gene searches, and ancestral sequence reconstruction. Researchers find that de novo genes preferentially originate from GC-rich ORFs with intrinsic folding potential. Together, our results reveal that the interplay between GC content and foldability - rooted in the structure of the genetic code - shapes the emergence of novel genes. More information: https://www.nature.com/articles/s41467-025-68022-7 Contact: Anne Lopes anne.lopes@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
January 13, 5:33 AM
|
Flexizyme-Based Strategy for the Synthesis of Stable, Non-Isomerizable Amide-Linked 2’-Aminoacyl-tRNAs and their Shortened Analogs
We developed a flexizyme-based semi-synthetic strategy that provides access to stable 2′- and 3′-amide AA-tRNA analogs, offering robust tools for structural biology and for probing regiospecificity in AA-tRNA-dependent enzymes. The study of the regiospecificity of aminoacyl-tRNA (AA-tRNA)-dependent enzymes and their structural characterization with AA-tRNAs are limited by rapid hydrolysis of the ester bond linking amino acid to tRNA. To overcome this limitation, stable AA-tRNA analogs bearing hydrolysis-resistant linkages, such as amide bonds or ester bioisosteres, have been developed. These analogs are valuable tools for investigating interactions between AA-tRNAs and various enzymes or ribonucleoproteins, including elongation factors, ribosomes, Fem-family transferases, and cyclodipeptide synthases. However, their synthesis remains technically challenging. Recently, flexizymes—engineered ribozymes capable of aminoacylating tRNAs with diverse amino acids or analogs—have enabled the synthesis of 3′-amide-linked AA-NH-tRNAs. Due to their inherent specificity for 3′-OH acylation, flexizymes have not been used to generate 2′-amide-linked analogs, and such regioisomers have remained unexplored. In this study, we demonstrate that while flexizymes cannot directly aminoacylate the 2′ position, they can nevertheless mediate the synthesis of 2′-aminoacyl-NH-tRNAs via a two-step regioisomerization mechanism with excellent yields. This finding provides new insights into the binding mode of AA-tRNAs to flexizymes and expands the chemical space of stable AA-tRNA analogs. Access to both 3′- and 2′-amide regioisomers will enable more precise studies of AA-tRNA recognition and catalysis by various AA-tRNA-dependent systems. More information : https://doi.org/10.1002/chem.202503506 Contact person: Matthieu Fonvielle matthieu.fonvielle@i2bc.paris-saclay.fr https://www.i2bc.paris-saclay.fr/enzymology-and-non-ribosomal-peptide-biosynthesis/

|
Scooped by
I2BC Paris-Saclay
January 9, 5:24 AM
|
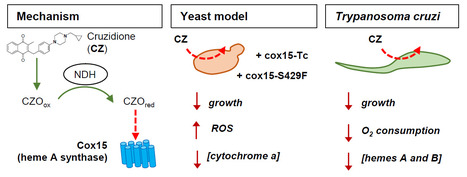
The heme A synthase Cox15, as a target of redox-active 3-benzylmenadiones with antiparasitic activity
We identified a new target for antiparasitic compounds and it is in the mitochondria. Chagas disease, caused by Trypanosoma cruzi, is a neglected parasitic infection. The very limited arsenal of anti-T. cruzi treatments calls for the development of new drugs. Recently, a library of 3-benzylmenadione derivatives was synthesized with cruzidione being the most efficient and specific compound against the parasite. To decipher its mode of action, we used the yeast Saccharomyces cerevisiae as model. Evidence pinpointed at the heme A synthase Cox15 as a primary target of cruzidione: 1) a mutation in Cox15 (i.e., S429F) renders the yeast cells highly sensitive to the drug, 2) treatment with cruzidione led to the loss of cytochrome c oxidase, an enzyme that relies on heme A as an essential cofactor and 3) replacement of the yeast Cox15 by T. cruzi enzyme resulted in a high sensitivity to cruzidione. We then investigated the effect of cruzidione in T. cruzi and observed a significant reduction of heme contents, most likely involving the inhibition of the heme A synthase. This, in turn, led to a decrease in O2 consumption by the parasite. Finally, using the yeast model, we showed that, similarly to what we previously found for the antimalarial benzylmenadione plasmodione, NADH-dehydrogenase plays a key role in cruzidione bioactivation. We proposed that the reduced benzoylmenadione metabolites produced by the reaction with NADH-dehydrogenase, act as Cox15 inhibitors. This study, through the identification of the mode of action of cruzidione, highlighted Cox15 as a novel target for antiparasitic drugs. More information: https://journals.asm.org/doi/10.1128/aac.01161-25 Contact: Brigitte Meunier, brigitte.meunier@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
January 6, 4:23 PM
|
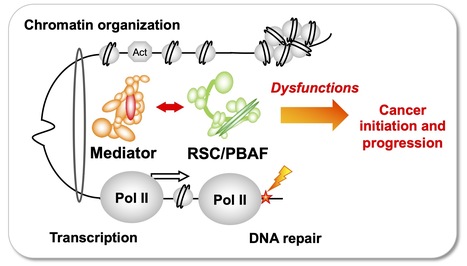
Team of J. Soutourina received the label from Ligue Nationale contre le cancer
Understanding how the essential complexes —the coregulator Mediator and the chromatin remodeling complex RSC/PBAF— cooperate in the nucleus In eukaryotes, transcription and DNA repair occur in the crowded context of chromatin. Dysfunctions of these processes can lead to cancers. Mediator is an essential and conserved multisubunit coactivator complex, mutated in many cancers. However, it remains largely unknown how Mediator and chromatin regulators coordinate their functions. A recent publication of the team suggests the novel hypothesis that Mediator acts in conjunction with the chromatin remodeling complex RSC (Remodels the Structure of Chromatin) of SWI/SNF family, homologous to PBAF (Polybromo-associated BAF) in human, representing the most frequently mutated complexes in cancers. Building on this recent publication, a new project has been supported by Ligue Nationale contre le cancer. Using the yeast model, with a perspective to extend the study to human cells, the team intends to decipher the molecular mechanisms involved in functional cooperation between these essential coregulator complexes in transcription regulation, DNA repair and chromatin organization relevant for cancer biology. Contact: Julie Soutourina julie.soutourina@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
December 8, 2025 5:52 AM
|
A tripartite protein complex promotes DNA transport during natural transformation in Firmicutes.
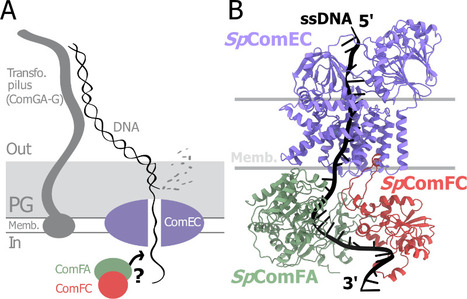
From structural modeling to natural transformation in bacteria: identification of a key protein complex for DNA transport across the membrane of Firmicutes. Transformation is a key mechanism of horizontal gene transfer, central to bacterial adaptation. This evolutionarily conserved process allows bacteria to integrate exogenous genetic material into their genome, thereby facilitating, for example, the spread of antibiotic resistance.
In a study published in the journal PNAS, scientists from I2BC (CEA/CNRS/UPSaclay, Gif-sur-Yvette) and DRCM/IBFJ (CEA/UPSaclay/UP Cité, Fontenay-aux-Roses), in collaboration with a laboratory at CBI (CNRS/University of Toulouse), identified a protein complex involved in the transport of single-stranded DNA across the membrane during transformation. Using AlphaFold, they structurally modeled this three-protein complex, which is highly conserved in the phylum Firmicutes, in interaction with single-stranded DNA. The structural model allowed them to identify a possible path for DNA through a conserved channel in one of the three proteins, a transmembrane protein, and then along a groove formed by the other two proteins. This model was validated by a robust experimental strategy in the bacterium Streptococcus pneumoniae, by measuring the impact on transformation efficiency of disruptive mutations in several protein–protein and protein–DNA interfaces. The transmembrane channel was found to be conserved in a structural model in Helicobacter pylori, and its importance for transformation was also experimentally confirmed in this bacterium. This study sheds light on the molecular mechanisms of bacterial transformation and demonstrates the power of macromolecular structure prediction to generate molecular hypotheses and guide functional experiments. More information: https://doi.org/10.1073/pnas.2511180122 Contact : Jessica Andreani jessica.andreani@i2bc.paris-saclay.fr
|

|
Scooped by
I2BC Paris-Saclay
Today, 5:00 AM
|
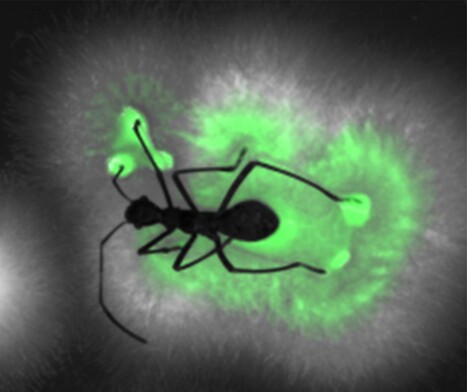
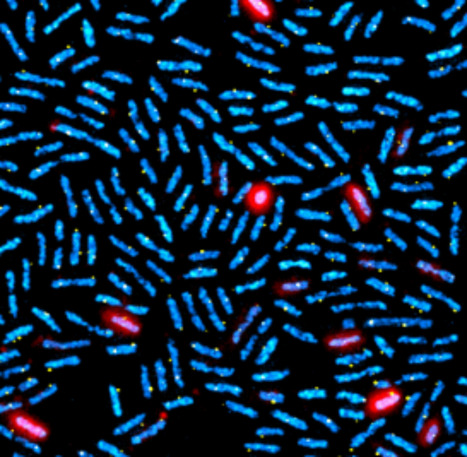
A wolf in sheep’s clothing
Distinguishing beneficial from harmful among closely related bacteria, a crucial question for organisms in nature and for biotechnology Many organisms crucially depend on the services of specific microorganisms and need to maintain these symbionts in large numbers in their body. Hosts dispose of highly specific symbiont selection mechanisms and have symbiotic organs to host safely these symbionts. It is believed that these sophisticated adaptations evolved to counter the thread of exploitation of the symbiotic association by opportunistic pathogens, although such cases are rarely reported. The association of the bean bug Riptortus pedestris with caballeronia bacteria is a model of a highly evolved symbiotic system. In a study published in PNAS, Ishigami et al report on an opportunistic pathogen of the bean bug that is phylogenetically related to symbionts. Initially, this Burholderia pathogen behaves as a true symbiont, efficiently passing all symbiont filtering mechanisms of the host, inducing the development of the symbiotic organ, and infecting and colonizing it. However, once settled, the pathogen throws off its disguise and shows its true nature: it breaks out from its confinement in the symbiotic organ, causes deadly sepsis and then escapes from the dead animal to recolonize the environment (figure). This case of a pathogen exploiting an intricate mutualistic association highlights the evolutionary pressures predicted by theory that have shaped the stringent partner-choice mechanisms usually observed in symbioses and at the same time reveals the fragility of these mutualisms despite their sophistication. Caballeronia and Burkholderia species belong both to the Burkholderia sensu latu clade of betaproteobacteria and were before phylogenetic revisions included in the single genus Burkholderia. In the 1990s, several Burkholderia strains were registered as biocontrol agents for agricultural applications but later withdrawn and a moratorium was placed on the registration of Burkholderia-containing products, after opportunistic human infections were reported with closely related strains. Thus, the Burkholderia has become a showcase of the potential health risks associated with the use of microorganisms in commercial applications. Because of the many known Burkholderia strains with potential applications in agriculture, it is important to dispose of criteria that distinguish friend from foe. A separate study by Agnoli et al, published in the ISME Journal, was conducted on a panel of 76 Burkholderia, Paraburkholderia and Caballeronia strains of the Burkholderia sensu lato. The work identified phenotypic traits and genetic markers that enable improved, strain-level evaluation of pathogenic potential and biocontrol capacity, supporting the rational selection of Burkholderia sensu latu strains for safe agricultural applications. More information :https://academic.oup.com/ismej/advance-article/doi/10.1093/ismejo/wrag081/8650978 Contact : Peter Mergaert peter.mergaert@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
April 15, 4:06 AM
|
The I2BC was well represented on Tuesday, April 14, at the Science Outreach Day co-organized by the French Society for Developmental Biology, the French Society for Cell Biology and the city of Ivry-sur-Seine. Seven members of the I2BC led workshops throughout the day. Nearly 1,200 elementary school students participated in workshops on cell, plant, and animal biology, as well as on the scientific method. Let’s hope we’ve inspired a few future scientists… one thing is certain: the children were thrilled! https://sbcf.fr/newsletter/mediation-scientifique-sbcf/ https://sfbd.fr/ressources/mediation/#1558951904425-8b59e54f-fdb5

|
Scooped by
I2BC Paris-Saclay
April 1, 5:18 AM
|
As part of the “Des Plantes et des Hommes” project led by the EUR Saclay Plant Science, a senior high school class visited the I2BC on March 25, 2026. Catherine Grandclément presented the Institute and a lecture on the symbiosis between legume plants and Rhizobium bacteria. The students visited the I2BC greenhouse to search for nodules on the roots of legumes plants before observing them with a macroscope. Finally, the microscopy platform and Amanda Edling de Barros introduced them to a confocal microscope and its use in studying symbiosis. With their interest in the subject and their fascination with the equipment, we hope we’ve inspired a passion for research! More information : Legume-rhizobia Symbiosis : https://www.i2bc.paris-saclay.fr/equipe-interactions-of-bacteria-with-plants-and-insects/ "Des Plantes et des Hommes” project : Des Plantes et des Hommes” project
Vous développez un projet ambitieux en biologie moléculaire et cellulaire, à l’interface de plusieurs disciplines, ou en biologie computationnelle ? Rejoignez l’I2BC, un institut de recherche de premier plan affilié au CEA, au CNRS et à l’Université Paris-Saclay. Nous recherchons de nouveaux responsables d’équipes autour de projets originaux, excellents scientifiquement, et capables de créer des synergies avec les recherches en cours à l’I2BC. Une attention particulière sera portée aux projets multidisciplinaires et aux approches en biologie computationnelle, notamment sur la modélisation des processus biologiques et l’intégration de données multi-échelles. L’I2BC offre un environnement multidisciplinaire dynamique, des plateformes technologiques de pointe, ainsi qu’un accompagnement fort, en particulier pour les profils en début de carrière. Les candidatures de profils plus avancés disposant déjà d’un poste permanent en France sont également bienvenues. 📅 Date limite : 22 avril 2026 📩 Contact : call2026@i2bc.paris-saclay.fr Présélection et notification aux candidats retenus : début juin 2026 - Entretiens sur place avec les candidats retenus : les 2 et 3 juillet. Les candidats doivent s'assurer d'être disponibles ces deux jours-là
- Annonce de la sélection finale d'ici la mi-juillet 2026
- Date de prise de fonction négociable à partir de l'automne 2026, en fonction du poste obtenu et de l'obtention du financement.
🔗 Toutes les informations sur notre site web
Via Life Sciences UPSaclay
La plateforme conjointe de Cryo-Microscopie Électronique de l’Institut de Biologie Intégrative de la Cellule (I2BC, Gif-sur-Yvette, Institut Joliot, CEA, Saclay) et du Synchrotron SOLEIL renforce son expertise avec l’arrivée de Heddy Soufari, recruté en tant qu’ingénieur-chercheur CEA. Spécialisé en biologie structurale et en cryo-microscopie électronique (cryo-EM), il consacre l’essentiel de son activité au développement méthodologique et au soutien scientifique des utilisateurs régionaux et nationaux sur les sites de SOLEIL et de l’I2BC. Docteur en biologie structurale de l’Université de Bordeaux, Heddy Soufari a orienté ses recherches vers la compréhension de la structure et de la dynamique des complexes macromoléculaires. Après un parcours académique entre la France et le Canada, il a acquis une solide expertise en cryo-microscopie électronique appliquée à l’étude de systèmes biologiques complexes. Il a notamment contribué à la caractérisation de ribosomes de mitochondries et de complexes protéiques chez différents modèles eucaryotes, avant de rejoindre la société NovAliX où il dirige le pôle Cryo-EM et met la microscopie au service du structure-based drug design (SBDD) sur des cibles thérapeutiques. Depuis son recrutement au CEA en septembre 2024, il accompagne les chercheurs de l’I2BC et de la région dans leurs projets de cryo-EM, de la préparation d’échantillons à l’analyse des reconstructions tridimensionnelles. Il développe parallèlement une activité méthodologique centrée sur la cryo-tomographie électronique (cryo-ET) et l’imagerie corrélative cryo-CLEM, avec pour objectif d’étudier l’organisation et la dynamique de complexes macromoléculaires in situ. Sa vision scientifique repose sur une approche multi-échelle, allant de la molécule unique à la cellule entière, afin d’explorer la tridimensionnalité des systèmes biologiques dans toute leur complexité. Avec l’arrivée d’Heddy Soufari, la plateforme renforce sa capacité à développer de nouvelles méthodologies et pipelines pour la tomographie cellulaire, l’analyse d’images 3D et la corrélation optique-électronique. Ces efforts visent à faire de la plateforme un pôle de référence pour la visualisation in situ des architectures cellulaires et des complexes macromoléculaires à haute résolution. -> Contact : Stéphane Bressanelli (plt-cryoem@i2bc.paris-saclay.fr) Plug In Labs Université Paris-Saclay : cliquer ICI Envie de (re)lire leurs précédents FOCUS PLATEFORME ? I2BC / Plateforme de cryo-microscopie électronique (CRYO-EM). La plateforme pour la cryo-microscopie électronique I2BC met à disposition une large gamme d’instruments et de services pour la biologie structurale : vitrification d’échantillons (Vitrobot, Leica GP2), microscope Tecnai Spirit (120 kV) pour les expériences préliminaires, microscopes de dernière génération Glacios (200 kV) pour les expériences à haute résolution, ainsi qu’un microscope confocal cryogénique Stellaris pour les approches corrélatives optique-électronique. La plateforme a la capacité de préparer et d’imager des échantillons biologiques de niveau de sécurité 2. Elle s’est associée au mésocentre de calcul Paris-Saclay pour l’hébergement des serveurs de stockage et d’analyse des images produites. La plateforme CryoEM de l’I2BC est associée à SOLEIL dans le cadre de la plateforme conjointe CryoEM@Paris-Saclay. Les images obtenues en cryo-microscopie électronique sur le microscope Glacios, ainsi que les moyens de calcul disponibles sur la plateforme, permettent le crible de grilles et la détermination de structures 3D à des résolutions dépassant fréquemment 3 Å et pouvant atteindre 2 Å. Ces premières structures permettent si nécessaire d'accéder aux microscopes plus puissants, notamment le Titan Krios G4 (300 kV) à SOLEIL, mais aussi les microscopes Titan Krios de l'ESRF, de l’IBS (Grenoble) ou de l'IGBMC (Strasbourg), menant aux résolutions atomiques. La plateforme CryoEM de l’I2BC permet aussi l'observation de complexes in situ (protéines à la surface d'organites purifiés ou de virus enveloppés, protéines membranaires reconstituées dans des liposomes...) par cryo-tomographie électronique. La plateforme accueille des projets issus de la recherche académique, institutionnelle et industrielle, et s’inscrit dans le réseau des infrastructures nationales en biologie structurale et intégrative (FRISBI). Ses missions couvrent la préparation d’échantillons, la collecte de données, le traitement d’images, la formation et l’accompagnement des utilisateurs. Cette plateforme fait partie du pôle des plateformes de Biologie Structurale de l'I2BC qui comprend six plateformes : cristallographie, RMN, CryoEM, mesures d'interactions macromoléculaires (PIM), expression de protéines recombinantes en systèmes eucaryotes (Prot-Ex) et sélection sur mesure de protéines artificielles comme ligands spécifiques de toute protéine d’intérêt (AlphaRep). D’autre part, les plateformes CryoEM, cristallographie, PIM et AlphaRep sont aujourd’hui labélisées IBISA sous la bannière BioStruct@UPSAY. A propos de l’Institut de Biologie Intégrative de la Cellule (I2BC - UMR 9198). L’I2BC est une Unité Mixte de Recherche (CEA, CNRS, Université Paris-Saclay), accueillant une soixantaine d’équipes de recherche et hébergeant 17 plateformes technologiques, réparties en 6 pôles. 2025 a aussi été une année clé pour l’I2BC : cette unité a fêté ses 10 ans !
Via Life Sciences UPSaclay

|
Scooped by
I2BC Paris-Saclay
March 8, 5:08 AM
|
A new-engineered integrative tool to target the terminal compartment of the Streptomyces chromosome
Pushing Streptomyces engineering to new frontiers! The Samy phage tool targets the chromosome’s farthest terminal regions, rich in specialized metabolite genes, enabling precise integration across strains. A breakthrough for antibiotic production like albonoursin. Phages are a valuable resource for the genetic engineering of Streptomyces antibiotic-producing bacteria. Indeed, a few integrative vectors based on phage integrase are available to insert transgene at specific genomic loci. Chromosome conformation captures previously demonstrated that the Streptomyces linear chromosome is organized in two spatial compartments: The central compartment encompassing most conserved and highly expressed genes in exponential phase, and the terminal compartments enriched in poorly conserved sequences including specialized metabolite biosynthetic gene clusters. This study introduces a new integrative tool based on a recently described phage, Samy, which specifically targets the terminal compartment of its native host chromosome. Samy is related to PhiC31 phage and, like the latter, encodes a serine integrase. Whereas PhiC31 targets a site generally located near the origin of replication, the Samy integration site is one of the farthest known attB sites from it. The authors demonstrated that the Samy integrase efficiently mediates the specific integration of a non-replicating plasmid in six Streptomyces strains from distinct clades. Bioinformatic analyses revealed that the Samy-attB site is rather conserved, and located in the terminal compartment of most Streptomyces chromosomes. Finally, heterologous expression of the albonoursin biosynthetic gene cluster from the Samy-, PhiC31-, and R4-attB sites yields quantitatively equivalent levels of production, though qualitative differences were observed. Altogether, these results demonstrate that the att-int Samy system expands Streptomyces genetic engineering tools by enabling targeted integration in the terminal chromosomal compartment. More information: https://doi.org/10.1007/s00253-026-13707-2 Contact: Stéphanie BURY-MONE stephanie.bury-mone@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
February 9, 5:23 AM
|
Workshop: Flow cytometry applied to microbiology (May 2026)

|
Scooped by
I2BC Paris-Saclay
January 28, 8:19 AM
|
Crosstalk between cohesins and axis proteins determines meiotic chromosome architecture in Sordaria macrospora
New discovery: a dynamic interplay between axis proteins and cohesins ensures chromosome stability during meiosis in Sordaria. Faithful chromosome segregation during meiosis requires the coordinated action of cohesin complexes and chromosome axis proteins. How these factors interact and communicate along chromosome axes, especially during meiotic prophase I, remains however, only partially understood. Researchers of the Genome Biology Department of the I2BC investigated the functional interplay between the cohesin components and regulators (Rad21, Rec8, Wapl, Sororin, Spo76/Pds5) and two meiosis-specific axis proteins Red1 and Hop1. Analysis of multiple combinations of their corresponding null mutants and of their genetic-epistasis interactions in the fungus Sordaria macrospora revealed a hierarchical regulatory network for their recruitment and releasing. Their work uncovers an unexpected role of axis proteins Red1 and Hop1, that together with Sororin, provide stage-specific protection of Spo76/Pds5 against Wapl-mediated release. Furthermore, we identify that Spo76/Pds5 is the main target of Wapl and acts as a central guardian of kleisin stability against Slx8/STUbL-dependent proteasomal degradation. Together, these findings show that a dynamic crosstalk between axis proteins and cohesins is crucial to preserve axis integrity and to ensure accurate meiotic progression. More information: https://doi.org/10.1371/journal.pgen.1012001 Contact: Stéphanie BOISNARD stephanie.boisnard@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
January 13, 3:30 PM
|
How virulence genes reorganize the Salmonella genome
Using functional genomics in sorted Salmonella populations and high-resolution microscopy, the researchers of the I2BC show that the expression of Pathogenicity Island 1 is associated with chromatin remodeling and with the repositioning of this region toward the nucleoid periphery. Chromatin provides a universal framework for organizing and regulating genomes across the three domains of life. In bacteria, it is composed of intrinsically supercoiled DNA associated with small DNA-binding proteins known as nucleoid-associated proteins (NAPs). Their binding can induce DNA bending, bridging, coating, and/or wrapping, giving rise to distinct modes of chromatin organization.
Bacterial chromatin can exist in a repressed state (silent chromatin) or in an actively transcribed state (active chromatin). Silent chromatin is largely associated with H-NS, a xenogeneic silencer that restricts the costly expression of genes acquired by horizontal transfer. In contrast, active chromatin is densely occupied by RNA polymerase and is characterized by different levels of DNA supercoiling. However, the changes in protein occupancy and chromatin organization that accompany transitions between these two states remain poorly understood.
Researchers of the Genome Biology Department of the I2BC in collaboration with the NGS and the Imaging facility of the I2BC and the Trinity College Dublin (Ireland), have unveiled the chromatin organization of horizontally acquired regions in Salmonella enterica serovar Typhimurium, which are essential for its pathogenicity. They show that expression of Pathogenicity Island 1 (SPI-1) is accompanied by local chromatin remodeling, marked by profound changes in three-dimensional organization and protein occupancy. This remodeling is also associated with the repositioning of SPI-1 toward the nucleoid periphery.
These findings provide new insights into the interplay between xenogeneic silencing, counter-silencing mechanisms, chromatin architecture, and the evolutionary integration of acquired DNA. They also reveal a finely tuned chromatin remodeling process that minimizes the cellular cost of activating pathogenicity islands, and they establish a direct link between the linear (1D) organization of the genome and its three-dimensional (3D) folding. More information: https://www.nature.com/articles/s41467-025-67746-w https://www.insb.cnrs.fr/fr/cnrsinfo/comment-les-genes-de-virulence-reorganisent-le-genome-de-salmonella-0 Contact: Vicky Lioy virginia.lioy@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
January 12, 7:46 AM
|
Cross-regulation of [2Fe–2S] cluster synthesis by ferredoxin-2 and frataxin.
Tight regulation of Fe-S clusters biosynthesis via a mutually antagonistic binding of frataxin and ferredoxin-2 to the assembly machinery, with several important implications for the Friedreich’s ataxia disease caused by frataxin deficiency. Iron-sulfur (Fe-S) clusters are essential metallocofactors that perform a multitude of biological functions. They are synthesized de novo by multi-proteins machienries and any defect in their synthesis leads to severe diseases such as Friedreich’s ataxia (FRDA), caused by defective expression of frataxin (FXN). Here, we uncover that efficient [2Fe-2S] cluster assembly requires a fine-tuned balanced ratio of FXN and Ferredoxin-2 (FDX2), an essential enzyme of the assembly process. [2Fe-2S] clusters are assembled on the scaffold protein ISCU2 with sulfur provided as a persulfide by NFS1, which is cleaved into sulfide by FDX2. FXN stimulates the whole process by accelerating persulfide transfer to ISCU2. Using an in vitro reconstituted human system, we show that any deviation from a close-to-equal amount of FXN or FDX2 downregulates Fe-S cluster synthesis. We performed a structure-function investigation, which revealed that this is due to competition between FXN and FDX2 for the same binding site on the NFS1-ISCU2 complex. We found that higher levels of FXN impair the persulfide-reductase activity of FDX2 and higher levels of FDX2 slow FXN-accelerated persulfide transfer to ISCU2. We also discovered that FDX2 directly hinders persulfide generation and transfer to ISCU2 by interacting with the persulfide-carrying mobile loop of NFS1. We further found that knocking-down FDX2 expression in a FRDA drosophila model, increases fly lifespan. Altogether, this work highlights a direct regulation of Fe-S cluster biosynthesis through antagonistic binding of FXN and FDX2 and suggests that decreasing FDX2 in the context of FXN deficiency in FRDA might constitute a novel therapeutic axis. More information: https://www.nature.com/articles/s41586-025-09822-1 Contact : Benoit D'Autréaux benoit.dautreaux@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
January 9, 5:05 AM
|
Gain and loss of gene function shaped the nickel hyperaccumulation trait in Noccaea caerulescens
Plants that hyperaccumulate nickel open the possibility to mine this metal from soils in an environmetally friendly manner. In this study, sequencing of a nickel hyperaccumulating plant together with genomic and transcriptomic comparisons reveal the molecular mechanisms underlying this extreme trait. Nickel hyperaccumulation is an extreme adaptation to ultramafic soils observed in more than 500 plant species. However, our understanding of the molecular mechanisms underlying the evolution of this trait remains limited. To shed light on these mechanisms, we have generated a high-quality genome assembly of the metal hyperaccumulator Noccaea caerulescens. We then used this genome as reference to conduct comparative intraspecific and interspecific transcriptomic analyses using various accessions of N. caerulescens and the non-accumulating relative Microthlaspi perfoliatum to identify genes associated with nickel hyperaccumulation.
Our results suggest a correlation between nickel hyperaccumulation and a decrease in the expression of genes involved in defense responses and the regulation of membrane trafficking. Surprisingly, these analyses did not reveal a significant enrichment of genes involved in the regulation of metal homeostasis. However, we found that the expression levels of selected metal transporter genes,
namely, NcHMA3, NcHMA4, and NcIREG2, are consistently elevated in N. caerulescens accessions hyperaccumulating nickel.
Furthermore, our analyses identified frameshift mutations in NcIRT1 associated with the loss of nickel hyperaccumulation in a few accessions. We further showed that the expression of a functional NcIRT1 in the roots of the La Calamine accession increases nickel accumulation in shoots. Our results demonstrate that NcIRT1 participates in nickel hyperaccumulation in N. caerulescens. They also
suggest that nickel hyperaccumulation is an ancient trait in N. caerulescens that has evolved from the high and constitutive expression of several metal transporters, including NcIREG2, and that the trait was subsequently lost in a few accessions due to mutations in NcIRT1. More information: https://doi.org/10.1093/plcell/koaf281 Contacts: Sylvain Merlot, sylvain.merlot@univ-tlse3.fr Sébastien Thomine, sebastien.thomine@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
December 10, 2025 4:16 AM
|
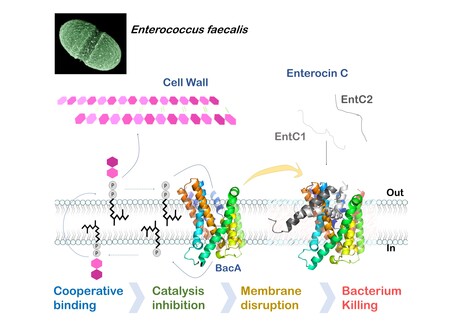
Mechanistic insights into Enterocin C bacteriocin targeting Enterococcus faecalis
Hijacking mechanism of membrane undecaprenyl phosphate recycling protein BacA in Enterococcus faecalis by a two-peptide bacteriocin revealed Antibiotic-resistance is a critical global health concern stressing the urgent need for new therapeutic strategies beyond conventional drugs. In this context, peptide-based bacteriocins constitute potential medical use antibacterial alternatives. In the present study, we have highlighted the molecular mechanism of a two-peptide bacteriocin, enterocin C, with potent activity against Enterococcus faecalis, a major opportunistic pathogen notorious for multidrug resistance. In collaboration with a group from INRAE MICALIS, the team has uncovered how enterocin C specifically exploits the membrane-embedded protein BacA as a cell surface receptor. BacA, widely conserved across bacteria, plays a central role in bacterial cell-wall biogenesis through its dual phosphatase and flippase activities, which are essential for recycling the lipid carrier undecaprenyl phosphate.
Using a combination of biochemical, biophysical, and microbiological approaches supported by AlphaFold structural modelling, we dissected the cooperative action of the two enterocin C peptides. Acting at nanomolar concentrations, peptide EntC1 inserts deeply into BacA’s outward-open catalytic pocket, blocking its enzymatic function and facilitating the subsequent binding of peptide EntC2. This dual docking event anchors the bacteriocin deep within the membrane’s hydrophobic core, ultimately triggering membrane disruption and bacterial cell death.
The findings reveal the molecular determinants of this precision targeting and represent the first detailed mechanistic description of a two-peptide bacteriocin’s mode of action. This work identifies BacA as a valuable target for bacteriocin-mediated killing and open avenues for the rational design of peptide-based antimicrobials for tailor-made antimicrobials to help combat antibiotic-resistant infections. More information : Thierry Touzé :thierry.touze@i2bc.paris-saclay.fr https://pubmed.ncbi.nlm.nih.gov/41232670/
|





 Your new post is loading...
Your new post is loading...















![Cross-regulation of [2Fe–2S] cluster synthesis by ferredoxin-2 and frataxin. | I2BC Paris-Saclay | Scoop.it](https://img.scoop.it/Ve3yN6adKt_5lzSWPuYkATl72eJkfbmt4t8yenImKBVvK0kTmF0xjctABnaLJIm9)