Your new post is loading...
 Your new post is loading...

|
Scooped by
Gilbert C FAURE
August 8, 2020 9:55 AM
|
Enumeration and characterization of circulating tumor cells (CTC) hold the promise of a real time liquid biopsy. They are however present in a large b…

|
Scooped by
Gilbert C FAURE
April 4, 2020 3:43 AM
|
Metastases and cancer recurrence are the main causes of cancer death. Circulating Tumor Cells (CTCs) and disseminated tumor cells are the drivers of cancer cell dissemination. The assessment of CTCs’ clinical role in early metastasis prediction, diagnosis, and treatment requires more information about their biology, their roles in cancer dormancy, and immune evasion as well as in therapy resistance. Indeed, CTC functional and biochemical phenotypes have been only partially characterized using murine metastasis models and liquid biopsy in human patients. CTC detection, characterization, and enumeration represent a promising tool for tailoring the management of each patient with cancer. The comprehensive understanding of CTCs will provide more opportunities to determine their clinical utility. This review provides much-needed insights into this dynamic field of translational cancer research.

|
Scooped by
Gilbert C FAURE
March 28, 2020 2:46 PM
|
Abstract
Background
Genomic rearrangement in anaplastic lymphoma kinase (ALK) gene occurs in 3−7% of patients with non-small-cell lung cancer (NSCLC). The detection of this alteration is crucial as ALK positive NSCLC patients benefit from ALK inhibitors, which improve both the patient's quality of...

|
Scooped by
Gilbert C FAURE
November 25, 2019 12:11 PM
|
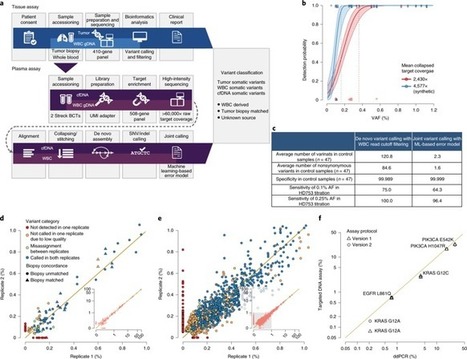
Accurate identification of tumor-derived somatic variants in plasma circulating cell-free DNA (cfDNA) requires understanding of the various biological compartments contributing to the cfDNA pool. We sought to define the technical feasibility of a high-intensity sequencing assay of cfDNA and matched white blood cell DNA covering a large genomic region (508 genes; 2 megabases; >60,000× raw depth) in a prospective study of 124 patients with metastatic cancer, with contemporaneous matched tumor tissue biopsies, and 47 controls without cancer. The assay displayed high sensitivity and specificity, allowing for de novo detection of tumor-derived mutations and inference of tumor mutational burden, microsatellite instability, mutational signatures and sources of somatic mutations identified in cfDNA. The vast majority of cfDNA mutations (81.6% in controls and 53.2% in patients with cancer) had features consistent with clonal hematopoiesis. This cfDNA sequencing approach revealed that clonal hematopoiesis constitutes a pervasive biological phenomenon, emphasizing the importance of matched cfDNA–white blood cell sequencing for accurate variant interpretation. Ultra-sensitive cell-free DNA (cfDNA) sequencing uncovers clonal hematopoiesis as a major source of somatic cfDNA variants in healthy individuals and patients with cancer, and underscores the importance of matched white blood cell DNA sequencing in liquid biopsy procedures.

|
Scooped by
Gilbert C FAURE
October 20, 2019 12:37 AM
|
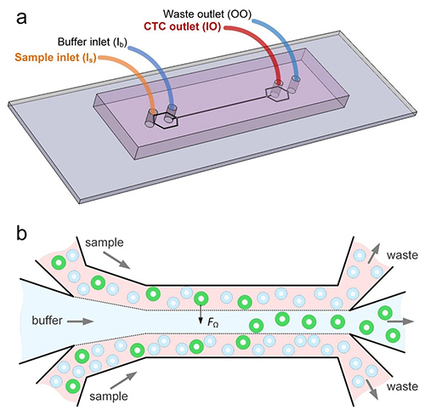
The detection of circulating tumor cells (CTCs), an approach considered to be “liquid biopsy”, is crucial in cancer diagnosis, monitoring and prognosis. However, the extremely large number of blood cells challenges the rare CTC isolation and enrichment.

|
Scooped by
Gilbert C FAURE
August 13, 2019 2:29 PM
|
University researchers develop microfluidic device that partitions cancer cells according to size in effort to create a useful liquid biopsy method.

|
Scooped by
Gilbert C FAURE
July 26, 2019 8:35 AM
|
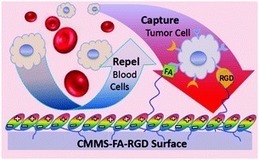
A circulating cancer cell (pink) attaches to carbon nanotube surface; white blood cells (blue) do not adhere and are later washed away.Using the tendency of circulating cancer cells to adhere to c…...

|
Scooped by
Gilbert C FAURE
June 17, 2019 2:21 AM
|
Clin Exp Med. 2019 Jun 12. doi: 10.1007/s10238-019-00563-w.[Epub ahead of print] Review...

|
Scooped by
Gilbert C FAURE
May 8, 2019 3:52 AM
|
Advancing technology is allowing scientists increasingly to search for tiny signs of cancer and other health issues in samples of patients' blood and urine. These "liquid biopsies" are less invasive than a traditional biopsy, and can provide information about what's happening...

|
Scooped by
Gilbert C FAURE
March 29, 2019 2:39 PM
|
March 2019—Cancer, liquid biopsy, pathology. The standard riff for talking about a promising new cancer test should be familiar to anyone within sneezing distance of a laboratory

|
Scooped by
Gilbert C FAURE
February 5, 2019 6:53 AM
|
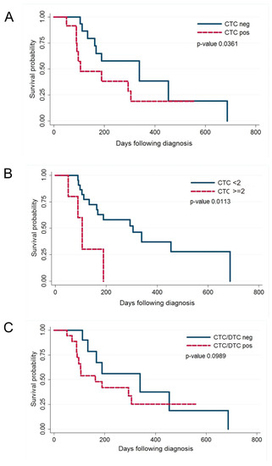
In human breast cancer, both circulating tumour cells (CTCs) in peripheral blood and disseminated tumour cells (DTCs) in the bone marrow are predictive of short survival and may be used as liquid biopsy to guide therapy.

|
Scooped by
Gilbert C FAURE
December 6, 2018 12:08 PM
|
This Review discusses the challenges in accurately identifying rare prostate circulating tumour cells or bone marrow-derived disseminated tumour cells and the resulting need for prostate-specific, sensitive biomarkers for the detection of these cells in liquid biopsy samples.

|
Scooped by
Gilbert C FAURE
November 28, 2018 1:39 PM
|
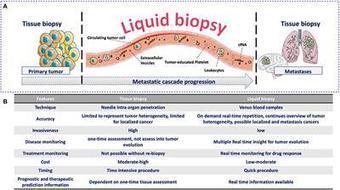
The initial results of an ongoing study show that a liquid biopsy has advantages over a tissue biopsy for people with lung cancer.
|

|
Scooped by
Gilbert C FAURE
July 21, 2020 9:57 AM
|
The metastatic cascade describes the process whereby aggressive cancer cells leave the primary tumor, travel through the bloodstream, and eventually reach distant organs to develop one or several metastases. During the last decade, innovative technologies have exploited the recent biological knowledge to identify new circulating biomarkers for the screening and early detection of cancer, real-time monitoring of treatment response, assessment of tumor relapse risk (prognosis), identification of new therapeutic targets and resistance mechanisms, patient stratification, and therapeutic decision-making. These techniques are broadly described using the term of Liquid Biopsy. This field is in constant progression and is based on the detection of circulating tumor cells, circulating free nucleic acids (e.g., circulating tumor DNA), circulating tumor-derived extracellular vesicles, and tumor-educated platelets. The aim of this review is to describe the biological principles underlying the liquid biopsy concept and to discuss how functional studies can expand the clinical applications of these circulating biomarkers.

|
Suggested by
LIGHTING
March 30, 2020 4:49 AM
|
The analysis of tumours using biomarkers in blood is beginning to transform cancer diagnosis, says Catherine Alix-Panabières. The challenge now is to make liquid biopsy a standard clinical tool.

|
Scooped by
Gilbert C FAURE
March 25, 2020 4:03 PM
|
In the era of precision oncology, liquid biopsy techniques, especially the use of plasma circulating tumour DNA (ctDNA) analysis, represent a paradigm shift in the use of genomic biomarkers with considerable implications for clinical practice. Compared with tissue-based tumour DNA analysis, plasma ctDNA is more convenient to test, more readily accessible, faster to obtain and less invasive, minimizing procedure-related risks and offering the opportunity to perform serial monitoring. Additionally, genomic profiles of ctDNA have been shown to reflect tumour heterogeneity, which has important implications for the identification of resistant clones and selection of targeted therapy well before clinical and radiographic changes occur. Moreover, plasma ctDNA testing can also be applied to cancer screening, risk stratification and quantification of minimal residual disease. These features provide an unprecedented opportunity for early treatment of patients, improving the chances of treatment success. Liquid biopsy techniques, especially the use of plasma circulating tumour DNA (ctDNA) analysis, are a convenient, fast and non-invasive approach to the diagnosis and monitoring of urological cancers, and could enable selection of targeted therapy before clinical and radiographic changes occur. In this Review, the authors discuss the uses of liquid biopsy and plasma ctDNA analysis in particular, and consider how it could be used in clinical practice now and in the future.

|
Scooped by
Gilbert C FAURE
November 23, 2019 4:08 AM
|
Cancer originates from small mistakes in our DNA. These mistakes are different for every person and dictate how aggressive their cancer is.On top of this, starting in development and continuing throughout life, new mistakes in our DNA can be …...

|
Scooped by
Gilbert C FAURE
August 18, 2019 2:21 PM
|
Room for improvement exists regarding recommendations for screening, staging, therapy selection, and frequency of surveillance of gastrointestinal cancers. Screening is costly and invasive, improved staging demands increased sensitivity and specificity to better guide therapy selection.

|
Scooped by
Gilbert C FAURE
August 2, 2019 4:44 AM
|
We perform single-cell genomic profiling of CTCs and pair the genomics to digital pathology features allowing us to see both cell phenotype and genotype

|
Scooped by
Gilbert C FAURE
June 18, 2019 6:46 AM
|
The Lund, Sweden-based company raised $4.1 million to accelerate commercialization of its portfolio of kits and services.

|
Scooped by
Gilbert C FAURE
May 8, 2019 2:31 PM
|
With the combined advantages of optical microscopy and flow cytometry, imaging flow cytometry (IFC) has quickly become an established tool for performing cytometric analysis in diverse areas of biology including microbiology, immunology, and stem cell biology1-14. IFC provides quantitative, spatially registered image data for every event analyzed, which allows the morphometric characterization of single cells in large heterogeneous populations, further advancing our understanding of cell‐to‐cell differences in physiological settings. The availability of image data produced by IFC is well aligned with the pressing need for progressively larger biomedical datasets for efficient and accurate data analysis with the help of computational tools (e.g., compressive imaging, data mining, machine/deep learning) to make better decisions in biomedical research and clinical settings. Recent studies 4, 5, 10, 12 show that IFC is highly effective for the characterization of DNA damage and repair, the accurate assessment of cell death and autophagy, fluorescence in situ hybridization (FISH), the localization and enumeration of specific molecules such as proteins, nucleic acids, and peptides, and the analysis of cell–cell interaction and cell cycle. Furthermore, a number of potential clinical applications of IFC are conceivable, such as liquid biopsy, leukemia detection, and identification of infectious diseases in which the availability of high‐content single‐cell images plays an important role. Objective and Highlights of the Special Issue Building on the excellent capabilities of IFC, the objective of this Special Issue is to provide a valuable forum where scientists, engineers, and users in the field share their most recent findings and understanding of basic principles, advanced techniques, and applications of IFC. Topics covered can be broadly categorized into (1) advanced imaging methods for IFC, (2) biological and clinical applications of IFC, and (3) computational techniques for IFC. Specifically, the first part of the Special Issue consists of nine papers (while the second part follows in a subsequent issue). Dey et al. 15 report the use of IFC to accurately differentiate between active and viable but not‐culturable forms of Psedomonas aeruginosa (a Gram‐negative bacterium abundant in the environment and water systems). Kim et al. 16 present their investigation of the physical and magnetic properties of monocytes in human blood together with plasma platelets, oxyhemoglobin red blood cells, and methemoglobin red blood cells by using IFC. Lelliott et al. 17 describe their development of an automated analysis method to rapidly evaluate cells that form neutrophil extracellular traps or precursors to neutrophil extracellular traps and differentiate them from cells that undergo other forms of cell death. Zhang et al. 18 report their development of a computational method for intelligent de‐blurring of out‐of‐focus cell images in IFC by deep learning. Gu et al. 19 present their demonstration of image‐guided cell sorting together with a cell classification system and real‐time isolation of single cells based on nuclear localization of glucocorticoid receptor, particle binding to the cell membrane, and DNA damage induced γ‐H2AX foci using the instrument. Goktas et al. 20 report a combination of IFC and angle‐resolved light scattering measurements to study the morphological features of red blood cells in sickle cell patients. Hui et al. 21 describe their development of a method, which they refer to as immuno‐flowFISH, for conducting FISH on immunophenotyped whole cells in suspension to detect chromosomal abnormalities in chronic lymphocytic leukemia. Lee et al. 22 report their development of a quantitative phase IFC system to conduct large‐scale biophysical phenotyping of numerous single cells (>1,000,000 cells) in a label‐free manner. Yaakov et al. 23 report their quantitative characterization of the kinetics of Mimivirus infection stages by IFC. History The field of IFC dates back to 1979 when Kay et al. 1 and Kachel et al. 2 first introduced and experimentally demonstrated imaging in flow. Since then, the field of IFC has slowly gained interest from the research community over the past two decades (Fig. 1). Before the year 2002, only a handful of researchers with interdisciplinary expertise in optics (in particular, optical imaging), fluidics, electronics, and cell biology were capable of building IFC‐capable systems and conducting experiments while many other researchers could appreciate the great potential of the technology. This situation dramatically changed around 2008 when the Amnis Corporation (now Luminex Corporation) successfully launched the first commercial IFC system. This has helped to democratize IFC from only a few specialist research groups at academic institutions to place IFC in the hands of many researchers worldwide, as shown by the sharp rise in the number of publications in Figure 1. As of today, IFC systems have been installed in research laboratories and flow cytometry core facilities at a number of universities and other research institutions to tackle problems unanswered by conventional (non‐imaging) flow cytometry. Future Perspective The field of IFC is expected to grow further in the next decade or so (Fig. 2). There are several driving factors that push the frontier of IFC and expand its utility in diverse areas of biology and medicine. The first driving factor is the emergence of advanced microfluidics and its integration into IFC platforms 7, 8, 12-14. Microfluidics provides higher degrees of freedom in multiplexing, automation, and parallelization than traditional capillary‐based fluidics in flow cytometry, thereby adding more functionalities for preparation, analysis, and manipulation of cells. The second driving factor is the advent of various optical imaging modalities other than traditional image acquisition based on CCD image sensors with time‐delay integration 10. They include time‐stretch imaging 8, 13, frequency‐division‐multiplexed imaging 9, 24, spatial–temporal transformation 14, temporally coded excitation 25, and stroboscopic fluorescence imaging 6, which are designed to handle sensitive image acquisition of fast‐flowing cells. The third driving factor is the availability of sophisticated digital image processing techniques, such as compressive imaging, data mining, and deep learning, powered by the rapidly evolving smartphone industry 4, 5, 7, 18, 19. The big data brought by IFC systems acts as a fuel for artificial neural networks that can characterize and classify cells with higher accuracy than classical image analysis algorithms. The fourth driving factor is the advent of image‐activated cell sorting (IACS) (also called image‐guided cell sorting) 7, 19, a technology that achieves the best of both worlds between IFC and fluorescence‐activated cell sorting (FACS). IACS extends the capabilities of FACS from one‐dimensional signal intensities to multidimensional information‐rich images, which can be used to analyze the spatial architecture of single cells. IACS offers the opportunity to study and exploit sorted cells via single‐cell analysis tools such as genomics, proteomics, and proteomics. With these driving factors, the number of researchers, applications, discoveries, and publications in the field of IFC is expected to go up in the next decade or so (Fig. 2). The future of IFC is bright. Literature Cited

|
Scooped by
Gilbert C FAURE
April 9, 2019 2:37 PM
|
Dr. David Wong, associate dean for research, and his team received regulatory approval on their EFIRM (electric field-induced release and measurement) technology as a Clinical Laboratory Improvement Amendments (CLIA)-certified, College of American Pathology (CAP)-accredited assay to be used in clinical practice. EFIRM creates a reliable, novel, and impactful method to detect tumor-causing, lung cancer mutations in saliva and blood that is non-invasive, cost-effective, and quick. For the past four years until now, the technology has been in research and testing phases. Small celebration after the technology received its regulatory approval. Left to right: Drs. David Wong, Charles Strom, Michael Tu, Paul Krebsbach (Dean), Richard Bender, and Josh Deignan. A major contributor to helping move the EFIRM testing phase along is the National Cancer Institute (NCI), who awarded Dr. Wong and his collaborators, Drs. Wayne Grody and Josh Deignan with the UCLA Department of Pathology and Laboratory Medicine and Drs. Fang Wei, Charles Strom and Michael Tu of the School of Dentistry, $2.5M in 2017. The grant’s support successfully paid off, as the technology is now approved to be used in medical offices to assist a doctor’s diagnosis of a major disease—lung cancer. The goal is to use results from EFIRM, based on a patient’s saliva sample, to adjust therapeutic strategies in real-time, improving clinical outcomes. Another grant from by the National Cancer Institute, awarded in October 2018 for $5M over five years, is continuing to develop and refine the technology for early assessment of cancer. Dr. Wong’s lab is working with Dr. Denise Aberle, professor of radiology at the David Geffen School of Medicine at UCLA, Dr. Steven Dubinett, co-investigator and chief of pulmonary and critical care medicine in the Geffen School, and We Liao, co-inventor of the technology, to test the blood and saliva of at least 300 at-risk patients for lung cancer. Dr. Wong’s team has shown that the technology can detect early-stage lung cancer with greater than 90 percent similarity with traditional tissue biopsies. In another defining moment for liquid biopsy and salivary diagnostics, Dr. Wong was invited to speak on the EFIRM Liquid Biopsy technology in an NCI-sponsored session at the annual meeting for the American Association for Cancer Research (AACR) in Atlanta, Georgia on April 2. In front of hundreds of cancer researchers from around the world, Wong was included in a session on the topic; “Liquid Biopsy: The Holy Grail for Early Cancer Detection.” Dr. Wong also chairs the Steering Committee of the NCI Liquid Biopsy consortium. Research panel at AACR, from left to right: Drs. Sudhir Srivastava (Chief NCI, Early Detection Research Network (EDRN), Lynn Sorbara (NCI Program Director), David Wong (UCLA), Bob Carter (Harvard/MGH), Changhuei Yang (CALTech), Nicholas Papadopoulos (Johns Hopkins). "Receiving regulatory approval for the EFIRM technology and also having the opportunity to present our research findings at a major cancer research meeting proves that salivary diagnostics must be integrated with mainstream, cutting-edge cancer early detection, screening and risk assessment,” Dr. Wong said.

|
Scooped by
Gilbert C FAURE
February 22, 2019 9:26 AM
|
In this Review, Pantel and Alix-Panabières provide an overview of approaches for the detection and characterization of minimal residual disease (MRD) using circulating tumour cells and circulating tumour DNA. They also discuss the clinical implications of such liquid biopsy approaches to MRD monitoring for the management of patients with cancer.

|
Scooped by
Gilbert C FAURE
January 9, 2019 8:50 AM
|
The way we think about cancer is about to change. What we used to treat as a random occurrence is becoming both predictable and preventable through an array of emerging circular biomarkers. Through the use of circulating tumor cells, cell-free circulating DNA, exosomes, exomers, oncosomes and other extracellular vesicles, we are now able to understand early stages of cancer and apply molecular profiling to clinical trial selection, patient treatment and minimal residual disease detection. The applications are expanding outside of cancer to include transplant medicine, cardiovascular disease, CNS, autoimmune and infectious disease. Encounter the latest research results and clinical trial data while networking with leaders in the field. Final Agenda Day 1 | Day 2 | Day 3 | Download Brochure Scientific Advisory Board Stefanie Jeffrey, MD, John and Marva Warnock Professor, Surgery; Chief, Surgical Oncology Research, Stanford University School of Medicine Stuart S. Martin, PhD, Professor, Physiology, Marlene and Stewart Greenebaum NCI Comprehensive Cancer Center, University of Maryland School of Medicine Catherine Alix-Panabières, PhD, Director, Laboratory of Rare Human Circulating Cells (LCCRH), Pathology and Onco-Biology Department, University Medical Center of Montpellier Klaus Pantel, MD, Professor and Founding Director, Institute of Tumor Biology, University, Medical Center Hamburg-Eppendorf Steven A. Soper, PhD, Professor, Micro and Nanofabricated Tools for Biological Discovery and Medical Diagnostics, University of Kansas Shannon L. Stott, PhD, Assistant Professor, Department of Medicine, Massachusetts General Hospital, Harvard Medical School; BioMEMS Resource Center, The MGH Cancer Center Arrive Early for: SUNDAY, MARCH 10, 2:00 - 5:00 PM (AFTERNOON SHORT COURSES) SC4: Translating CTCs and ctDNA for Clinical Use - Detailed Agenda SUNDAY, MARCH 10, 5:30 - 8:30 PM (DINNER SHORT COURSES) SC14: Liquid Biopsy Technologies and Applications - Detailed Agenda MONDAY, MARCH 11, 8:00 - 11:00 AM (MORNING SHORT COURSES) SC21: Detection and Characterization of Circulating Biomarkers - Detailed Agenda MONDAY, MARCH 11 10:30 am Conference Program Registration Open OPENING KEYNOTE SESSION: WHAT NEEDS TO BE DONE TO BRING CTCS TO THE CLINIC? NEXT STUDIES 11:50 Chairperson’s Opening Remarks Klaus Pantel, MD, Professor and Founding Director, Institute of Tumor Biology, University, Medical Center Hamburg-Eppendorf 12:00 pm Circulating Tumor Cells in Breast Cancers: Current Clinical Validity and Utility François-Clément Bidard, MD, PhD, Professor of Medical Oncology, Institut Curie & Versailles St. Quentin University Numerous clinical data have been collected regarding the clinical validity of CTC count and characterization in both early and advanced breast cancer patients. After having reviewed these data, we will discuss how CTC detection may help customize breast cancer therapy. 12:20 Critical Assessment of the Challenges of Using Blood-Based Biomarkers in Prostate Cancer from a Clinical Point of View Daniel C. Danila, MD, Medical Oncology Fellow, Genitourinary Oncology Service, Department of Medicine, Memorial Sloan Kettering Cancer Center In addition to determining whether circulating tumor cells (CTC) can be used as prognostic biomarkers for patients needing further therapy, current studies have proposed many predictive and pharmacodynamic biomarkers to facilitate the doctor’s ability to optimize and adjust dosage based on their ability to target specific pathways. Significant efforts have been focused on developing and analytically validating predictive biomarkers to select patients most likely to benefit the specific therapy. 12:40 PANEL DISCUSSION 1:00 Session Break 1:10 Luncheon Presentation I to be Announced 1:40 Luncheon Presentation II to be Announced 2:10 Session Break NUCLEIC ACID ANALYSIS IN EXTRACELLULAR VESICLES AND CTCS 2:30 Chairperson’s Remarks Klaus Pantel, MD, Professor and Founding Director, Institute of Tumor Biology, University, Medical Center Hamburg-Eppendorf 2:40 The Whole Transcriptional Landscape of Circulating Tumors Cells in Metastatic Breast Cancer Julie E. Lang, MD, Associate Professor of Clinical Surgery; Director, Breast Cancer Program, Department of Surgery, Keck School of Medicine, University of Southern California The aim of our study was to evaluate if RNA Seq of circulating tumor cells could serve as a surrogate for biopses of macrometastases. We evaluated treatment opportunities based on circulating tumor cell profiles and tracked them under the selection pressure of systemic therapy in Stage IV breast cancer. RNA Seq of circulating tumor cells may be used to discover molecular alterations that are potentially clinically actionable. 3:10 Large Oncosomes as a Source of Cancer-Specific Circulating Biomarkers Dolores Di Vizio, MD, PhD, Division of Cancer Biology and Therapeutics, Departments of Surgery, Biomedical Sciences and Pathology and Laboratory Medicine, Samuel Oschin Comprehensive Cancer Institute, Cedars-Sinai Medical Center Extracellular vesicles (EVs) are heterogeneous membrane enclosed structures harboring molecular cargo from the cell of origin. Large oncosomes are a novel subtype of EV that is released by highly migratory cancer cells. Large oncosomes contain abundant tumor-derived cargo, which can be analyzed upon purification from plasma and other biological fluids. Here we show analyses of protein, RNA and DNA in controlled system and patient plasma. 3:40 Molecular Analysis of Circulating Tumor Cells, Extracellular Vesicles and Nucleic Acids as Liquid Biopsy in the Follow-Up of Metastatic Breast Cancer Patients to Stratify Patients for Targeted Therapy Prof. Dr. Sabine Kasimir-Bauer, Head of Research Laboratory, Department of Gynecology and Obstetrics, University Hospital Essen The so-called liquid biopsy is discussed to be a surrogate marker for therapy stratification of metastatic breast cancer patients. We compared and analyzed RNA profiles enclosed in circulating tumor cells or extracellular vesicles and performed mutational analysis of cell-free DNA (cfDNA) in plasma samples (Next Generation Sequencing) in the follow-up of the disease to get insights into their feasibility for therapy stratification and to predict therapeutic options. 4:10 An Open, End to End, and Flexible Platform for Scalable CTC Collection, Identification, and Analysis Tad George, PhD, Senior Director of Platform, Applications, RareCyte RareCyte provides instrumentation and consumables that enable an exquisitely sensitive, accurate, simple, and repeatable workflow from liquid biopsy to single cell isolation. The open and end to end RareCyte CTC workflow will be the focus of this talk. 4:40 Refreshment Break and Transition to Plenary Session 5:00 Plenary Keynote Session 6:00 Grand Opening Reception in the Exhibit Hall with Poster Viewing 7:30 Close of Day Day 1 | Day 2 | Day 3 | Download Brochure TUESDAY, MARCH 12 7:30 am Registration Open and Morning Coffee 8:00 Plenary Keynote Session 9:15 Refreshment Break in the Exhibit Hall with Poster Viewing CTCS AND LIQUID BIOPSY – NEW WINDOW INTO SELECTING BEST TREATMENT FOR PATIENTS 10:15 Chairperson’s Remarks Stuart S. Martin, PhD, Professor, Physiology, Marlene and Stewart Greenebaum NCI Comprehensive Cancer Center, University of Maryland School of Medicine 10:25 Chemotherapy-Induced Pro-Metastatic Changes in Breast Cancer Microenvironment Maja H. Oktay, MD, PhD, Professor of Pathology, The L.G. Koss Division of Cytology, Montefiore Medical Center; Professor of Anatomy and Structural Biology; Director, Clinical Imaging Applications, Integrated Imaging Program, Michael F. Price Center for Genetic and Translational Medicine, Albert Einstein College of Medicine Chemotherapy may induce pro-metastatic changes within breast cancer microenvironment in both mouse and human breast cancer and subsequent hematogenous cancer cell dissemination to distant sites. Cellular and molecular mechanism involved in chemotherapy-mediated cancer cell dissemination as well as pharmacological inhibitors of this process will be discussed. 10:55 RNA Signatures in CTCs for Precision Cancer Medicine David T. Miyamoto, MD, PhD, Assistant Professor, Radiation Oncology, Harvard Medical School; Department of Radiation Oncology, Center for Cancer Research, Massachusetts General Hospital CTC analysis is a type of liquid biopsy that enables RNA expression profiling, an analysis not possible with ctDNA. Microfluidic enrichment of CTCs followed by RNA-based digital PCR enables the high-throughput and highly specific detection of CTC molecular signatures in several cancers. These CTC signatures can predict therapeutic responses and may enable the early detection of invasive cancers, thus guiding the precision management of cancer patients. 11:25 Assessing PD-L1 Expression on CTCs and Correlation with Immunotherapy Response Rajan Kulkarni, MD, PhD, Assistant Professor, Dermatology and CEDAR/Knight Cancer Institute, Oregon Health and Science University PD-1 inhibitors have promising efficacy in several cancers. Circulating tumor cell (CTC) PD-L1 levels could supplement tissue PD-L1 biopsy results by sampling from disseminated tumor sites. Towards establishing CTCs as a screening tool, we developed a protocol to isolate CTCs at high purity and immunostain for PD-L1. Monitoring of PD-L1 expression on CTCs could be an additional biomarker that may help in determining response to immunotherapies. 11:55 Molecular Heterogeneity as Expressed in Cells from Liquid Biopsies Using CellSearch and DepArray in Multiple Myeloma Steven Gross, PhD, Head, CellSearch Assay Development, Research & Development, Menarini Silicon Biosystems Inc Exploring the molecular heterogeneity as expressed in cells using CellSearch and DepArray technologies in multiple myeloma and other cancers. 12:10 pm Sponsored Presentation (Opportunity Available) 12:25 Session Break 12:35 Luncheon Presentation I : Esophageal Adenocarcinoma: Circulating Tumor Cells in Multimodal Treatment Protocols Jasmina Kuvendjiska, PhD, Surgery, Freiburg University Our study was designed as a pilot study of the ESOPEC-Trial. We evaluated the presence and morphology of CTCs during the treatment period. The experimental results will be correlated with patients’ overall and relapse-free survival. 1:05 Luncheon Presentation II to be Announced 1:35 Refreshment Break in the Exhibit Hall with Poster Viewing TECHNOLOGIES FOR LIQUID BIOPSY 2:05 Chairperson’s Remarks Steven A. Soper, PhD, Professor, Micro and Nanofabricated Tools for Biological Discovery and Medical Diagnostics, University of Kansas 2:10 Diagnosing Disease with Rare Circulating EVs: Finding Heterogeneous, Nanoscale Needles in a Nanoscale Haystack David A. Issadore, PhD, Assistant Professor, Bioengineering & Electrical & Systems Engineering, University of Pennsylvania Circulating EVs contain a wealth of proteomic and genetic information, presenting an enormous opportunity for liquid biopsy. While microfluidics have been used to successfully isolate cells from complex samples, scaling these approaches for EV isolation has been limited by the low throughput and susceptibility to clogging of nanofluidics. Moreover, the analysis of EV biomarkers is confounded by substantial heterogeneity between patients and within a disease itself. To address these challenges, we developed a multichannel nanofluidic system to analyze crude clinical samples. Using this platform, we isolated EVs, profile the RNA cargo inside of these EVs, and apply a machine learning algorithm to generate predictive panels that could provide useful diagnostics for applications in traumatic brain injury and pancreatic cancer using both murine models and clinical samples. 2:40 Single Cell Morphogenomic and Morphoproteomic Profiling of Liquid Biopsy in Prostate Cancer Paymaneh D. Malihi, PhD Candidate, USC Michelson Center for Convergent Bioscience Peter Kuhn, PhD, Dean’s Professor of Biological Sciences and Professor of Biological Sciences, Medicine, Biomedical Engineering, and Aerospace and Mechanical Engineering and Associate Director, The Bridge, University of Southern California Tumor heterogeneity is prevalent in both treatment-naïve localized and end-stage metastasized prostate cancer, and may contribute to the broad range of clinical presentation, treatment response, and disease progression. High Definition-Single Cell Analysis enables morphoproteomic and morphogenomic profiling of single cells from touch preparations of tissue cores as well as liquid samples. HD-SCA platform enables real-time molecular profiling of cells and has the potential to elucidate the origin and evolution of metastatic tumor cells. 3:10 High Precision Isolation and Analysis of Exosomes Daniel T. Chiu, PhD, A. Bruce Montgomery Professor of Chemistry and Bioengineering, University of Washington We have recently developed microfluidic and nanofluidic systems for the isolation and analysis of exosomes, offering detailed molecular information with single-exosome resolution. Here, we will describe our technical approach, device performance, and the new information we learned about exosomes as revealed by the new measurements. 3:40 Presentation to be Announced 3:55 Talk Title to be Announced Christoph Mauracher, PhD, Managing Director, STRATEC Consumables GmbH 4:10 St. Patrick’s Day Celebration in the Exhibit Hall with Poster Viewing 5:00 Breakout Discussions in the Exhibit Hall Parallel Analysis of Circulating Biomarkers in Immunotherapy Genevieve Boland, MD, PhD, Director, Melanoma Surgery Program, Massachusetts General Hospital; Director, Surgical Oncology Research Laboratories, Massachusetts General Hospital; Assistant Professor, Harvard Medical School; Associate Member, Broad Institute Clinical application of blood-based biomarkers in melanoma Unmet clinical needs in blood-based biomarkers Microvesicle applications in immunotherapy Clinical Utility and Impact of Liquid Biopsy Rajan Kulkarni, MD, PhD, Assistant Professor, Dermatology and CEDAR/Knight Cancer Institute, Oregon Health and Science University Necessary features of technologies/tests for clinical utility Tumor information that is of clinical relevance Necessity for standardization The Importance and Challenge in CTC Culture Professor Yong-Jie Lu, MBBS, MD, PhD, Professor, Centre for Molecular Oncology, Barts Cancer Institute, Queen Mary University of London Why is CTC culture important? What is the challenge? Does it worth to try it? Alternative ways to avoid it? How can we success with CTC culture? The researcher, technology development and the funding supporter/policy marker. Deploying Bioengineered 3D Tumor Models into Clinical Practice and the Pharmaceutical Pipeline Aleksander Skardal, PhD, Assistant Professor, Regenerative Medicine, Wake Forest Institute for Regenerative Medicine, Wake Forest School of Medicine Where is the clinical enterprise and/or pharma in terms of seeing value in these model systems? How do we convince clinicians and more traditional scientists that these new technologies are important and effective? 3D models are typically better representations of in vivo biology; Why the slow adoption? (besides decreased throughput) Complexity (more cell types or multiple organoids per model system) versus scale-up/high throughput Simple tumor organoids (aka spheroids) are amenable to medium-high throughput screening. What about multi-organ system (aka body-on-a-chip)? Where in the pharmaceutical pipeline do more advanced systems that are less amenable to high throughput fall? Post-preclinical/pre-human Phase I? Serve as go/no go points? Isolation and Analysis of CTCs Min Yu, MD, PhD, Assistant Professor, Stem Cell Biology and Regenerative Medicine Member, Norris Comprehensive Cancer Center, Keck School of Medicine, University of Southern California Downstream analysis of circulating tumor cells Technologies for CTC isolation Experimental mouse models for metastasis analysis Using CTCs as liquid biopsy 6:00 Close of Day Day 1 | Day 2 | Day 3 | Download Brochure WEDNESDAY, MARCH 13 7:30 am Registration Open and Morning Coffee 8:00 Plenary Keynote Session > 10:00 Refreshment Break and Poster Competition Winner Announced in the Exhibit Hall CIRCULATING BIOMARKERS INFORMING CLINICAL TRIALS 10:50 Chairperson’s Remarks Stefanie Jeffrey, MD, John and Marva Warnock Professor, Surgery; Chief, Surgical Oncology Research, Stanford University School of Medicine 11:00 Utilizing Extracellular Vesicles for Prediction and Monitoring of Immunotherapy Responses in Melanoma Genevieve Boland, MD, PhD, Director, Melanoma Surgery Program, Massachusetts General Hospital; Director, Surgical Oncology Research Laboratories, Massachusetts General Hospital; Assistant Professor, Harvard Medical School; Associate Member, Broad Institute Immune checkpoint inhibitors (ICI) show therapeutic benefit in melanoma. We identified pretreatment and early on-treatment biomarkers that incorporate tumor and host immune status. Our findings suggest that peripheral blood-derived exosomes may serve as a non-invasive biomarker to probe tumor and immune responses to ICI therapy, functioning as both a predictive marker of ICI responsiveness as well as a monitoring tool for tumor persistence and immune activation. 11:30 Molecular Assessment of Circulating Extracellular Vesicles Toward Liquid Biopsy Diagnosis of Gastrointestinal Stromal Tumor and Ewing Sarcoma Andrew K. Godwin, PhD, Chancellors Distinguished Chair in Biomedical Sciences and Endowed Professor and Director, Molecular Oncology, Professor, Department of Pathology and Laboratory Medicine, University of Kansas Medical Center, Deputy Director, University of Kansas Cancer Center There is growing interest in exploiting circulating extracellular vesicles (EVs), primarily small bioactive nanovesicles (60–120 nm), known as exosomes, as developing sources of biomarkers for diagnosis and therapeutic response. However, EVs are heterogeneous vesicles released by both healthy and tumors cells, with the vast majority of circulating vesicles being derived from normal host cells. To establish their utility as valuable tools for the diagnosis of cancer and therapeutic monitoring of disease progression and response to therapy, we used molecular approaches to profile the proteome of Gastrointestinal Stromal Tumor (GIST)- and Ewing Sarcoma (EWS)-derived EVs, as well as other types of cancer, which provided insights into the oncogenic cargo of these tumor-derived EVs. Using proteins enriched in these different sarcomas, we have developed microfluidic platforms and immunocapture approaches to target tumor-associated EVs and in turn measure exosomal proteins and/or RNAs specific to a cancer subtype to further enhance the specificity of these liquid-based biopsies. 12:00 pm Clinical Trials – Liquid Biopsy in Lung Cancer Lecia V. Sequist, MD, Hematology/Oncology, Massachusetts General Hospital We will review the wide variety of liquid biopsy assays that have been used in clinical trials for lung cancer patients, and how this is affecting practice in the real world. 12:30 Session Break 12:40 Luncheon Presentation (Sponsorship Opportunity Available) or Enjoy Lunch on Your Own 1:10 Refreshment Break in the Exhibit Hall and Last Chance for Poster Viewing NEW MODELS OF MECHANISMS OF CANCER DISSEMINATION 1:50 Chairperson’s Remarks Shannon L. Stott, PhD, Assistant Professor, Department of Medicine, Massachusetts General Hospital, Harvard Medical School; BioMEMS Resource Center, The MGH Cancer Center 2:00 Advanced in vitro Cancer Models: Metastasis-on-a-Chip and Patient-Derived Organoid Technologies Aleksander Skardal, PhD, Assistant Professor, Regenerative Medicine, Wake Forest Institute for Regenerative Medicine, Wake Forest School of Medicine Most in vitro models of human tumors are too simplistic and fail to accurately recapitulate the in vivo tumor microenvironment. This presentation will describe 1) development of a microfluidic platform supporting observable metastasis from a primary site organoid to downstream target organoids, and 2) the translation of organoid technology to support models derived from clinical tumor biospecimens for personalized ex vivo drug screening and therapeutic efficacy identification prior to treatment. 2:30 Patient-Derived Circulating Tumor Cells Inform Mechanisms of Metastasis Min Yu, MD, PhD, Assistant Professor, Stem Cell Biology and Regenerative Medicine Member, Norris Comprehensive Cancer Center, Keck School of Medicine, University of Southern California Circulating tumor cells (CTCs) are expected to contain metastasis-initiating cells that can shed light on the mechanisms of cancer metastasis. However, due to limited patient-derived material, the metastatic capability of CTCs has yet to be proved. Using our recently established patient-derived CTC lines, we found that different patient CTC lines demonstrated distinct metastatic tissue tropisms in immunodeficient mice and identified associated pathways to specific organs via RNA-seq and ATAC-seq analysis. 3:00 From Sample-to-Single Cells-to-Answer – An Integrated Microfluidics Approach for Identifying Cancer Cells Abraham “Abe” Lee, PhD, William J. Link Professor and Chair, Department of Biomedical Engineering (BME); Professor, Mechanical & Aerospace Engineering; Director, NSF I/UCRC CADMIM Research Lab: Biomolecular Microsystems and Nano Transducers (BioMiNT), University of California, Irvine Liquid biopsy is performed by a microfluidic acoustic microstreaming device to isolate and target circulating tumor cells (CTCs) in whole blood. A key bottleneck is to identify the critical subpopulation of cells, often at single cell resolution among billions of cells in circulation. An in vitro perfused vascularized 3D tissue platform can then determine which CTCs can be disseminated to distant tissues through the circulatory system. 3:30 Session Break EXPANSION OF CIRCULATING TUMOR CELLS: A CHALLENGE! 3:40 Chairperson’s Remarks Catherine Alix-Panabières, PhD, Director, Laboratory of Rare Human Circulating Cells (LCCRH), Pathology and Onco-Biology Department, University Medical Center of Montpellier 3:45 Expansion of Patient-Derived Circulating Tumor Cells from Liquid Biopsy Chwee Teck (C.T.) Lim, PhD, NUS Society (NUSS) Professor, Department of Biomedical Engineering; Director, Biomedical Institute for Global Health Research and Technology; Founding Principal Investigator, Mechanobiology Institute, National University of Singapore Personalized therapy in cancer requires monitoring of patients’ individual response to treatment. One approach is to assess drug efficacy on circulating tumor cells (CTCs) obtained from patients’ blood samples (aka liquid biopsy) following ex vivo expansion into CTC clusters. We developed a simple microfluidics-based culture platform that allows co-culture of immune cells with CTCs (obtained after red blood cell lysis of patients’ blood) to promote CTC cluster formation. 4:15 Holy Grail and Big Challenge – Culture of Circulating Tumor Cells Yong-Jie Lu, MBBS, MD, PhD, Professor, Centre for Molecular Oncology, Barts Cancer Institute, Queen Mary University of London Circulating tumor cells (CTCs) provide a minimally invasive approach to access living cancer cells. The culture of CTCs enables the investigation of their biological features, mechanisms of cancer metastasis and the opportunity for in vitro therapeutic sensitivity tests. Challenge to CTC culture includes their rarity, small proportion of viable cells and unclear favorable growth condition. I will give a brief summary of culture strategies and our own experience. 4:45 Growing Organoids from CTCs: The Importance of CTC Input Andrew Rhim, PhD, Associate Director for Translational Research Ahmed Center for Pancreatic Cancer Research Assistant Professor of Internal Medicine UT MD Anderson Cancer Center Liquid biopsies have potential to inform personalized treatments and decision making in patients with cancer. While still in its infancy, the field has exploded, featuring numerous modalities and source material, including circulating tumor cells (CTCs), circulating nucleic acids, exosomes, and others. CTCs, however, hold the greatest promise, as not only do they contain all the tumor-derived material that are in the circulation, but if they could be cultured and propagated ex vivo, the prospect of functional analysis and direct drug combination testing could direct true individualized cancer care. Moreover, since CTCs are in the circulation, serial assay of CTCs, during the throes of treatment, could allow for nimble decision-making to thwart resistance. The major hurdle in the application of CTCs in the clinic is the inability to establish CTC cultures. Our laboratory has hypothesized that the major determinant of culture success is the number of CTCs seeded. This hypothesis is driven in part by evidence suggesting that a sub population of tumor cells retain stem cell like characteristics, called cancer stem like cells. These cells are theorized to be required to initiate tumors and metastases since they possess the capacity for self-renewal. In current protocols, at most 60ml of blood is phlebotomized for CTC isolation. However, we estimate that this yields far too few cancer stem like cells to achieve success in culture. Here we will review the efforts we have undertaken to achieve robust and routine cultures of CTCs, mostly by focusing on dramatically increasing CTC input with less emphasis on epithelial cell purity. In so doing, our preliminary data suggest real promise in our approaches. 5:15 Close of Conference Program Stay Late for: MARCH 14-15 TS4: Introduction to Liquid Biopsy for Cancer - Detailed Agenda Day 1 | Day 2 | Day 3 | Download Brochure

|
Suggested by
Société Francaise d'Immunologie
November 29, 2018 4:12 AM
|
Introduction: Multiple myeloma (MM) is a malignancy of terminally differentiated plasma cells. The high heterogeneity of MM cells is one of the major cause of disease relapse. Detection of circulating MM cells (CMMC) from peripheral blood is a useful procedure to investigate tumor heterogeneity and provides a painless alternative to the classic bone marrow biopsy to monitor disease progression. Here we demonstrate that the synergy between CellSearch® (CS) and DEPArray™ (DA) technologies can be used to identify, isolate and characterize at the genetic level single and pure CMMCs .
Methods: 4.0 ml of peripheral blood samples were obtained from 3 patients with MM. Putative CMMCs were enriched with CS using anti-CD138 or anti-CD138/CD38 as positive selection marker and subsequently stained with CD38-PE, CD19/CD45-APC immunofluorescent probes. Cells detection and enumeration was performed based on the co-localization of nuclei DAPI staining and CD38-PE. Single CMMCs (CD38+/CD19- and CD45-/DAPI+) and White Blood Cells (WBCs: CD38-/CD19+ or CD45+/DAPI+) were then isolated using the DA NxT system. Single cells genomic DNA was amplified using Ampli 1™ Whole Genome Amplification (WGA) kit and Illumina®-compatible libraries were obtained using Ampli 1™ LowPass kit and a high-throughput, customized automated protocol using Hamilton STARLet Liquid handler. Highly-multiplexed, genome-wide single-cell Low-Pass Copy Number Alteration (LPCNA) analysis was performed using HiSeq 2500 Illumina® platform.
Results: CS and DA workflow* enabled the isolation of 215 single CMMC, selected for LPCNA analysis. 42 single WBCs were also included as normal controls. Copy-number profiles of single CMMCs showed relevant gains and losses of chromosomal segments, as result of a high-level genomic instability. Notably, intra-patient CMMCs revealed overall conserved CNA patterns with subclonal alterations, suggesting a certain level of branched tumor evolution. Conversely, a higher degree of heterogeneity in CMMCs CNA profiles was observed among different patients. Interestingly, CNAs detected in all patients are located in regions containing genes involved in cell cycle regulation (MAPK, NOTCH pathways) and cell signaling (IL6R), which might be involved in proliferative processes and immuno-surveillance escape.
Conclusion: The combination of CS and DA workflow* with a streamlined automated protocol allowed to obtain hundreds of genomic libraries from pure single CMMCs. The presented workflow constitutes a non-invasive, rapid and high-throughput approach for characterizing MM tumor heterogeneity and progression, suggesting a possible future implementation in clinical applications.
*For Research Use Only. Not for use in diagnostic procedures.
Disclosures Raspadori: Menarini Silicon Biosystems: Employment. Forcato: Menarini Silicon Biosystems: Employment. Edoardo: Menarini Silicon Biosystems: Employment. Papadopulos: Menarini Silicon Biosystems: Employment. Ferrarini: Menarini Silicon Biosystems: Employment. Del Monaco: Menarini Silicon Biosystems: Employment. Terracciano: Menarini Silicon Biosystems: Employment. Morano: Menarini Silicon Biosystems: Employment. Gross: Menarini Silicon Biosystems: Employment. Bolognesi: Menarini Silicon Biosystems: Employment. Buson: Menarini Silicon Biosystems: Employment. Fontana: Menarini Silicon Biosystems: Employment. Connelly: Menarini Silicon Biosystems, Inc.: Employment, Other: Chief R&D Officer, USA. Simonelli: Menarini Silicon Biosystems: Employment. Medoro: Menarini Silicon Biosystems: Employment. Manaresi: Menarini Silicon Biosystems: Employment.
|

 Your new post is loading...
Your new post is loading...
 Your new post is loading...
Your new post is loading...