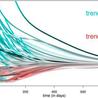
Analysis: With experts concerned over rising UK case rates, where are we also with deaths, hospital admissions, long Covid and the economy?
Get Started for FREE
Sign up with Facebook Sign up with X
I don't have a Facebook or a X account

| Tags |
|---|
 Your new post is loading... Your new post is loading...
 Your new post is loading... Your new post is loading...
Oldest known ancestor of octopuses unearthed in Montana in form of approximately 330m-year-old fossil...
Researchers at MIT have developed a synthetic gene circuit that triggers the body’s immune system to attack cancers when it detects signs of the disease. Via Gerd Moe-Behrens
From
phys
Every cell needs a shell. The cell interior is separated from its surroundings by a membrane made up of fat molecules, helping to create the environment needed for the cell to survive. Development of artificial cells is similarl Via Gerd Moe-Behrens
Phillip Trotter's insight:
Share your insight
Recent years have witnessed an explosion of extensive geolocated datasets related to human movement, enabling scientists to quantitatively study individual and collective mobility patterns, and to generate models that can capture and reproduce the spatiotemporal structures and regularities in human trajectories. The study of human mobility is especially important for applications such as estimating migratory flows, traffic forecasting, urban planning, and epidemic modeling. In this survey, we review the approaches developed to reproduce various mobility patterns, with the main focus on recent developments. This review can be used both as an introduction to the fundamental modeling principles of human mobility, and as a collection of technical methods applicable to specific mobility-related problems. The review organizes the subject by differentiating between individual and population mobility and also between short-range and long-range mobility. Throughout the text the description of the theory is intertwined with real-world applications.
Human Mobility: Models and Applications Via Complexity Digest
Phillip Trotter's insight:
This is a comprehensive review of mobility methods that are essential for emergent models of disease, transportation, traffic and economics among other applications,. Worth reading.
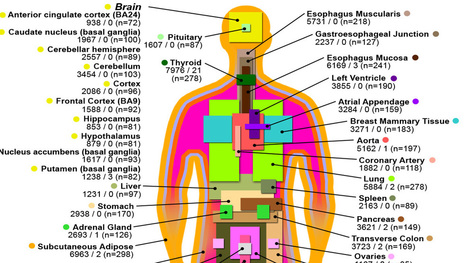
A study by an international consortium of scientists reached a major milestone in establishing a baseline understanding of gene expression across healthy human tissues, and linking genes to disease. Via Integrated DNA Technologies
Phillip Trotter's insight:
Share your insight
Which mattered first at the dawn of life: proteins or nucleic acids? Proteins may have had the edge if a theorized process let them grow long enough to become self-replicating catalysts. Via Integrated DNA Technologies
It will enter clinical trials to prevent and treat the infection next year.
Phillip Trotter's insight:
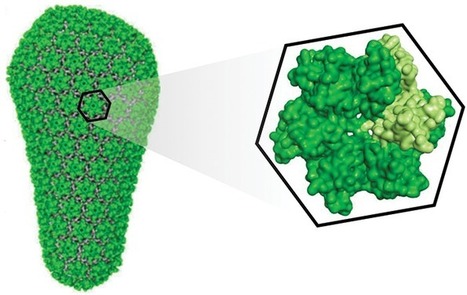
This possibly the most promising news in HIV treatment research in over a decade. A research collaboration between the US National Institutes of Health and the pharmaceutical company Sanofi has produced a new antibody for treatment of AIDS. Developed from three "broadly neutralising antibodies", that a small number of patients develop in response to HIV infection, the new antibody has been shown to be effective to 99% of HIV strains in vitro tests with monkeys. Human trials start next year. to see the BBC article click the picture. To see the full paper - see here http://science.sciencemag.org/content/early/2017/09/22/science.aan8630
First potential new novel treatment for HIV in 10 years leverages the geometry of the HIV Capsid
Network science offers a powerful language to represent and study complex systems composed of interacting elements from the Internet to social and biological systems. In its standard formulation, this framework relies on the assumption that the underlying topology is static, or changing very slowly as compared to dynamical processes taking place on it, e.g., epidemic spreading or navigation. Fuelled by the increasing availability of longitudinal networked data, recent empirical observations have shown that this assumption is not valid in a variety of situations. Instead, often the network itself presents rich temporal properties and new tools are required to properly describe and analyse their behaviour.A Guide to Temporal Networks presents recent theoretical and modelling progress in the emerging field of temporally varying networks, and provides connections between different areas of knowledge required to address this multi-disciplinary subject. After an introduction to key concepts on networks and stochastic dynamics, the authors guide the reader through a coherent selection of mathematical and computational tools for network dynamics. Perfect for students and professionals, this book is a gateway to an active field of research developing between the disciplines of applied mathematics, physics and computer science, with applications in others including social sciences, neuroscience and biology. Via Complexity Digest
From
arxiv
We present a model of contagion that unifies and generalizes existing models of the spread of social influences and micro-organismal infections. Our model incorporates individual memory of exposure to a contagious entity (e.g., a rumor or disease), variable magnitudes of exposure (dose sizes), and heterogeneity in the susceptibility of individuals. Through analysis and simulation, we examine in detail the case where individuals may recover from an infection and then immediately become susceptible again (analogous to the so-called SIS model). We identify three basic classes of contagion models which we call \textit{epidemic threshold}, \textit{vanishing critical mass}, and \textit{critical mass} classes, where each class of models corresponds to different strategies for prevention or facilitation. We find that the conditions for a particular contagion model to belong to one of the these three classes depend only on memory length and the probabilities of being infected by one and two exposures respectively. These parameters are in principle measurable for real contagious influences or entities, thus yielding empirical implications for our model. We also study the case where individuals attain permanent immunity once recovered, finding that epidemics inevitably die out but may be surprisingly persistent when individuals possess memory.
A generalized model of social and biological contagion Via Complexity Digest
From
arxiv
In this paper we present a thorough analysis of the nature of news in different mediums across the ages, introducing a unique mathematical model to fit the characteristics of information spread. This model enhances the information diffusion model to account for conflicting information and the topical distribution of news in terms of popularity for a given era. We translate this information to a separate graphical node model to determine the spread of a news item given a certain category and relevance factor. The two models are used as a base for a simulation of information dissemination for varying graph topoligies. The simulation is stress-tested and compared against real-world data to prove its relevancy. We are then able to use these simulations to deduce some conclusive statements about the optimization of information spread.
Characterizing information importance and the effect on the spread in various graph topologies Via Complexity Digest
Phillip Trotter's insight:
Share your insight
From
arxiv
How cooperation can evolve between players is an unsolved problem of biology. Here we use Hamiltonian dynamics of models of the Ising type to describe populations of cooperating and defecting players to show that the equilibrium fraction of cooperators is given by the expectation value of a thermal observable akin to a magnetization. We apply the formalism to the Public Goods game with three players, and show that a phase transition between cooperation and defection occurs that is equivalent to a transition in one-dimensional Ising crystals with long-range interactions. We also investigate the effect of punishment on cooperation and find that punishment acts like a magnetic field that leads to an "alignment" between players, thus encouraging cooperation. We suggest that a thermal Hamiltonian picture of the evolution of cooperation can generate other insights about the dynamics of evolving groups by mining the rich literature of critical dynamics in low-dimensional spin systems.
Thermodynamics of Evolutionary Games Via Complexity Digest |
The pandemic has taught us that viruses are not easy to identify – and can spread like wildfire...
Phillip Trotter's insight:
After the last two years of ever changing policies and unclear guidelines with decisions often grounded in politics rather than health science - its clear that we now need to start studying and learning the lessons from the pandemic so we are better prepared in the future.
Remember flu?Despite lockdowns holding it at bay, a small group of scientists is searching the globe for deadly new strains – and to work out what to put in next winter’s vaccines...
Observational data about human behavior is often heterogeneous, i.e., generated by subgroups within the population under study that vary in size and behavior. Heterogeneity predisposes analysis to Simpson's paradox, whereby the trends observed in data that has been aggregated over the entire population may be substantially different from those of the underlying subgroups. I illustrate Simpson's paradox with several examples coming from studies of online behavior and show that aggregate response leads to wrong conclusions about the underlying individual behavior. I then present a simple method to test whether Simpson's paradox is affecting results of analysis. The presence of Simpson's paradox in social data suggests that important behavioral differences exist within the population, and failure to take these differences into account can distort the studies' findings.
Computational Social Scientist Beware: Simpson's Paradox in Behavioral Data Via Complexity Digest
Phillip Trotter's insight:
When building models of human systems understanding Simpson's paradox is essential for creating behaviorally accurate representations
Researchers at MIT have developed a synthetic gene circuit that triggers the body's immune system to attack cancers when it detects signs of the disease. Via Gerd Moe-Behrens
Most drugs work by tinkering with the behavior of proteins. Like meddlesome coworkers, these molecules are designed to latch onto their target proteins and keep them from doing what they need to do. Via Integrated DNA Technologies
A new statistical model has enabled researchers to pinpoint 27 novel genes thought to prevent cancer from forming, in an analysis of over 2000 tumours across 12 human cancer types. The findings could help create new cancer treatments that target these genes, and open up other avenues of cancer research. Via Integrated DNA Technologies
Results are in from the first phase 1 clinical trial for a Zika vaccine, and they are very promising. A new generation DNA-based Zika vaccine, developed by Wistar scientists in collaboration with Inovio Pharmaceuticals and GeneOne Life Science, was found to be safe and well tolerated by all study participants and was able to elicit an immune response against Zika, opening the door to further and larger trials to move this vaccine forward. Via Integrated DNA Technologies
Phillip Trotter's insight:
Very welcome news regarding initial ZIka virus trials.
Most digital sensors are easy to connect, but those tucked away in hard-to-access spots will need to harvest ambient energy
Phillip Trotter's insight:
Powering sensors over the long time periods is key to sensor network related developments in environmental monitoring and industrial internet of things. By recouping ambient energy as a power sources for sensors - deployed duration times can become measured in years and decades rather than weeks and months. Good article on changes in sensor power sources designers can expect to be available soon.
From
arxiv
Poor economies not only produce less; they typically produce things that involve fewer inputs and fewer intermediate steps. Yet the supply chains of poor countries face more frequent disruptions---delivery failures, faulty parts, delays, power outages, theft, government failures---that systematically thwart the production process. To understand how these disruptions affect economic development, we model an evolving input--output network in which disruptions spread contagiously among optimizing agents. The key finding is that a poverty trap can emerge: agents adapt to frequent disruptions by producing simpler, less valuable goods, yet disruptions persist. Growing out of poverty requires that agents invest in buffers to disruptions. These buffers rise and then fall as the economy produces more complex goods, a prediction consistent with global patterns of input inventories. Large jumps in economic complexity can backfire. This result suggests why "big push" policies can fail, and it underscores the importance of reliability and of gradual increases in technological complexity.
Contagious disruptions and complexity traps in economic development Via Complexity Digest
CompleNet is an international conference that brings together researchers and practitioners from diverse disciplines—from sociology, biology, physics, and computer science—who share a passion to better understand the interdependencies within and across systems. CompleNet is a venue to discuss ideas and findings about all types networks, from biological, to technological, to informational and social. It is this interdisciplinary nature of complex networks that CompleNet aims to explore and celebrate.
CompleNet 2018 - 9th Conference on Complex Networks Boston (MA, US) March 5-8, 2018 Via Complexity Digest
Phillip Trotter's insight:
Given the growth discipline complex network analysis tools and techniques, nthis could be a must do conference for 2018. Definitely one for the calendar.
One hundred and ten Zika virus genomes from ten countries and territories involved in the Zika virus epidemic reveal rapid expansion of the epidemic within Brazil and multiple introductions to other regions.
Zika virus evolution and spread in the Americas Nature 546, 411–415 (15 June 2017) doi:10.1038/nature22402 Via Complexity Digest
From
arxiv
Biological organisms must perform computation as they grow, reproduce, and evolve. Moreover, ever since Landauer's bound was proposed it has been known that all computation has some thermodynamic cost -- and that the same computation can be achieved with greater or smaller thermodynamic cost depending on how it is implemented. Accordingly an important issue concerning the evolution of life is assessing the thermodynamic efficiency of the computations performed by organisms. This issue is interesting both from the perspective of how close life has come to maximally efficient computation (presumably under the pressure of natural selection), and from the practical perspective of what efficiencies we might hope that engineered biological computers might achieve, especially in comparison with current computational systems. Here we show that the computational efficiency of translation, defined as free energy expended per amino acid operation, outperforms the best supercomputers by several orders of magnitude, and is only about an order of magnitude worse than the Landauer bound. However this efficiency depends strongly on the size and architecture of the cell in question. In particular, we show that the {\it useful} efficiency of an amino acid operation, defined as the bulk energy per amino acid polymerization, decreases for increasing bacterial size and converges to the polymerization cost of the ribosome. This cost of the largest bacteria does not change in cells as we progress through the major evolutionary shifts to both single and multicellular eukaryotes. However, the rates of total computation per unit mass are nonmonotonic in bacteria with increasing cell size, and also change across different biological architectures including the shift from unicellular to multicellular eukaryotes.
The thermodynamic efficiency of computations made in cells across the range of life Via Complexity Digest
Phillip Trotter's curator insight,
July 7, 2017 5:25 AM
The concept of computation as it occurs in biology is fascinating and this paper is likely to become a-classic - worth reading.
|