Your new post is loading...
 Your new post is loading...

|
Scooped by
I2BC Paris-Saclay
March 9, 8:48 AM
|
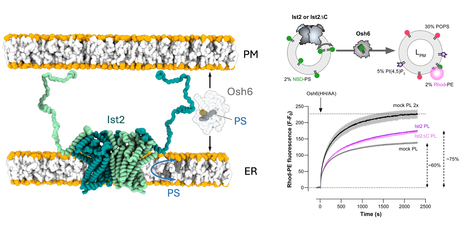
Phosphatidylserine dynamics between the endoplasmic reticulum and the plasma membrane in Saccharomyces cerevisiae.
A functional partnership between a lipid scramblase and a lipid transfer protein in the regulation of phosphatidylserine homeostasis. Phosphatidylserine is synthesised in the endoplasmic reticulum (ER), but this negatively charged lipid is highly concentrated in the plasma membrane (PM). There, it plays numerous roles in processes such as cell signalling and cell fusion, and it can also mediate apoptosis and synaptic pruning when exposed in the outer leaflet of the PM. In the yeast Saccharomyces cerevisiae, selective PS transport between the ER and the PM by the lipid transfer protein Osh6 is required for PS enrichment in the plasma membrane. Osh6 operates at membrane contact sites (MCS), where it interacts with the Ist2 protein embedded in the ER, tethering the ER to the PM via its long, disordered C-terminal tail. In two studies conducted in collaboration with the CRBM (CNRS/University of Montpellier), the IPMC (CNRS/University of Nice Côte d'Azur), and the IBCP (CNRS/University of Lyon), we show that Ist2 is a membrane transporter that catalyses rapid lipid exchange across the two leaflets of the ER – acting as a lipid ‘scramblase’ – and that this scrambling activity sustains Osh6-mediated PS transfer between the ER and the PM. First, we demonstrated that Ist2 catalyses the scrambling of different lipids in vitro after reconstituting purified Ist2 in proteoliposomes. Molecular dynamics simulations were then used to identify a cavity through which the lipid headgroup passes during transport from one leaflet to another. Further cellular studies revealed a close relationship between the COPII complex and the Ist2 protein. This was manifested by impaired yeast growth and vacuolar trafficking, as well as disruption of ER exit sites. We also found that Ist2 deletion stimulates the formation of ER-derived lipid droplets and changes their composition. Finally, using artificial, reconstituted ER-PM contact sites, we demonstrated that Ist2-mediated lipid scrambling sustains Osh6-mediated lipid transfer. Together, our studies identify Ist2 as a lipid scramblase and establish that lipid scrambling in the ER by Ist2 controls various cellular functions, such as vesicular transport and lipid droplet homeostasis. This highlights the importance of lipid dynamics for ER function. Furthermore, we reveal a functional partnership between Ist2-mediated lipid scrambling and Osh6-mediated lipid transfer at MCS. More information : https://doi-org.insb.bib.cnrs.fr/10.1083/jcb.202502112 https://www-science-org.insb.bib.cnrs.fr/doi/10.1126/sciadv.adz2217 Contact : Guillaume Lenoir guillaume.lenoir@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
January 12, 7:46 AM
|
Cross-regulation of [2Fe–2S] cluster synthesis by ferredoxin-2 and frataxin.
Tight regulation of Fe-S clusters biosynthesis via a mutually antagonistic binding of frataxin and ferredoxin-2 to the assembly machinery, with several important implications for the Friedreich’s ataxia disease caused by frataxin deficiency. Iron-sulfur (Fe-S) clusters are essential metallocofactors that perform a multitude of biological functions. They are synthesized de novo by multi-proteins machienries and any defect in their synthesis leads to severe diseases such as Friedreich’s ataxia (FRDA), caused by defective expression of frataxin (FXN). Here, we uncover that efficient [2Fe-2S] cluster assembly requires a fine-tuned balanced ratio of FXN and Ferredoxin-2 (FDX2), an essential enzyme of the assembly process. [2Fe-2S] clusters are assembled on the scaffold protein ISCU2 with sulfur provided as a persulfide by NFS1, which is cleaved into sulfide by FDX2. FXN stimulates the whole process by accelerating persulfide transfer to ISCU2. Using an in vitro reconstituted human system, we show that any deviation from a close-to-equal amount of FXN or FDX2 downregulates Fe-S cluster synthesis. We performed a structure-function investigation, which revealed that this is due to competition between FXN and FDX2 for the same binding site on the NFS1-ISCU2 complex. We found that higher levels of FXN impair the persulfide-reductase activity of FDX2 and higher levels of FDX2 slow FXN-accelerated persulfide transfer to ISCU2. We also discovered that FDX2 directly hinders persulfide generation and transfer to ISCU2 by interacting with the persulfide-carrying mobile loop of NFS1. We further found that knocking-down FDX2 expression in a FRDA drosophila model, increases fly lifespan. Altogether, this work highlights a direct regulation of Fe-S cluster biosynthesis through antagonistic binding of FXN and FDX2 and suggests that decreasing FDX2 in the context of FXN deficiency in FRDA might constitute a novel therapeutic axis. More information: https://www.nature.com/articles/s41586-025-09822-1 Contact : Benoit D'Autréaux benoit.dautreaux@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
October 15, 2025 9:07 AM
|
Moss BRCA2 lacking the canonical DNA-binding domain promotes homologous recombination and binds to DNA
Despite having low homology with BRCA2 proteins from other organisms and no folded domain, the newly discovered BRCA2 protein from Physcomitrium patens (the green yeast) promotes homologous recombination by binding to recombinases and DNA. BRCA2 is crucial for mediating homology-directed DNA repair (HDR) through its binding to single-stranded DNA (ssDNA) and the recombinases RAD51 and DMC1. Most BRCA2 orthologs have a canonical DNA-binding domain (DBD) with the exception of Drosophila melanogaster. It remains unclear whether such a noncanonical BRCA2 variant without DBD possesses a DNA-binding activity. Here, we identify a new noncanonical BRCA2 in the model plant Physcomitrium patens (PpBRCA2). We establish that PpBRCA2 is essential for genome integrity maintenance, somatic DNA double-strand break (DSB) repair, HDR-mediated gene targeting, and RAD51 foci recruitment at DNA break sites. PpBRCA2 is also critical for DSB repair during meiosis. Interestingly, PpBRCA2 interacts strongly with RAD51 but weakly with DMC1, suggesting a distinct meiotic function compared to other BRCA2 homologs. Despite lacking the canonical DBD, PpBRCA2 binds ssDNA through its disordered N-terminal region and efficiently promotes HDR. Our work highlights that the ssDNA binding capacity of BRCA2 homologs is conserved regardless of the presence of a canonical DBD and provides a deeper understanding of BRCA2’s functional diversity across species. More information: https://pmc.ncbi.nlm.nih.gov/articles/PMC12412785/ Contact: Sophie Zinn sophie.zinn@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
July 10, 2025 6:12 AM
|
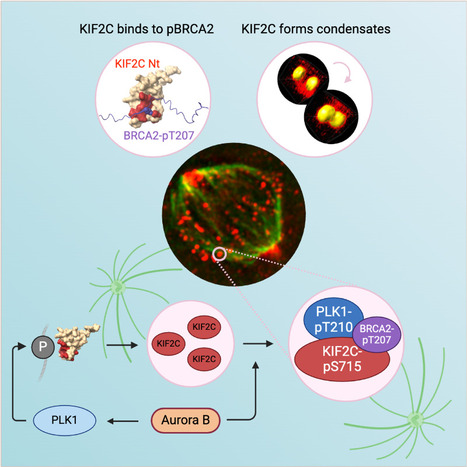
KIF2C condensation concentrates PLK1 and phosphorylated BRCA2 on kinetochore microtubules in mitosis
The microtubule depolymerase KIF2C forms membrane-less organelles at the kinetochore through its N-terminal phospho-peptide binding domain that interacts with BRCA2 and other phosphorylated targets in mitosis. During mitosis, the microtubule depolymerase KIF2C, the tumor suppressor BRCA2, and the kinase PLK1 contribute to the control of kinetochore-microtubule attachments. Both KIF2C and BRCA2 are phosphorylated by PLK1, and BRCA2 phosphorylated at T207 (BRCA2-pT207) serves as a docking site for PLK1. Reducing this interaction results in unstable microtubule-kinetochore attachments. Here we identified that KIF2C also directly interacts with BRCA2-pT207. Indeed, the N-terminal domain of KIF2C adopts a Tudor/PWWP/MBT fold that unexpectedly binds to phosphorylated motifs. Using an optogenetic platform, we found that KIF2C forms membrane-less organelles that assemble through interactions mediated by this phospho-binding domain. KIF2C condensation does not depend on BRCA2-pT207 but requires active Aurora B and PLK1 kinases. Moreover, it concentrates PLK1 and BRCA2-pT207 in an Aurora B-dependent manner. Finally, KIF2C depolymerase activity promotes the formation of KIF2C condensates, but strikingly, KIF2C condensates exclude tubulin: they are located on microtubules, especially at their extremities. Altogether, our results suggest that, during the attachment of kinetochores to microtubules, the assembly of KIF2C condensates amplifies PLK1 and KIF2C catalytic activities and spatially concentrates BRCA2-pT207 at the extremities of microtubules. We propose that this novel and highly regulated mechanism contributes to the control of microtubule-kinetochore attachments, chromosome alignment, and stability. More Information : https://academic.oup.com/nar/article/53/11/gkaf476/8160319 Contact: Sophie Zinn sophie.zinn@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
April 3, 2025 8:51 AM
|
A monomer–dimer switch modulates the activity of plant adenosine kinase
Novel regulation of adenosine kinase activity in plants Adenosine undergoes ATP-dependent phosphorylation catalyzed by adenosine kinase (ADK). In plants, ADK also phosphorylates cytokinin ribosides, transport forms of the hormone. Here, we investigated the substrate preferences, oligomeric states, and structures of ADKs from moss (Physcomitrella patens) and maize (Zea mays) alongside metabolomic and phenotypic analyses. We showed that dexamethasone-inducible ZmADK overexpressor lines in Arabidopsis can benefit from a higher number of lateral roots and larger root areas under nitrogen starvation. We discovered that maize and moss enzymes can form dimers upon increasing protein concentration, setting them apart from the monomeric human and protozoal ADKs. Structural and kinetic analyses revealed a catalytically inactive unique dimer. Within the dimer, both active sites are mutually blocked. The activity of moss ADKs, exhibiting a higher propensity to dimerize, was 10-fold lower compared with maize ADKs. Two monomeric structures in a ternary complex highlight the characteristic transition from an open to a closed state upon substrate binding. This suggests that the oligomeric state switch can modulate the activity of moss ADKs and probably other plant ADKs. Moreover, dimer association represents a novel negative feedback mechanism, helping to maintain steady levels of adenosine and AMP. More information: https://academic.oup.com/jxb/advance-article/doi/10.1093/jxb/eraf094/8068880 Contact: Solange Morera solange.morera@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
February 11, 2025 9:48 AM
|
Complete description of the biosynthetic process of [2Fe-2S] clusters
Iron-sulfur clusters are essential metallocofactors providing catalytic activities to a multitude of enzymes and proteins. Here, we report a comprehensive picture of their assembly mechanism, showing that it relies on the formation of [1Fe-1S] precursors fused into [2Fe-2S] clusters upon dimerization of the scaffold protein. Iron-sulfur (Fe-S) clusters are ubiquitous metallocofactors constituting the active site of a multitude of enzymes and proteins involved in electron transfer, catalysis, sulfur donation and signalling. They are made of iron and sulfide ions assembled into diverse structures. The [2Fe-2S] and [4Fe-4S] clusters are the most common forms in organisms. They are synthesized by multi-protein machineries which have remained highly conserved during evolution. The iron-sulfur cluster (ISC) assembly machinery present in eukaryotes and prokaryotes synthesizes [2Fe-2S] clusters, which serve as building blocks for the assembly of [4Fe-4S] clusters. The core ISC machinery assembles [2Fe-2S] clusters on the scaffold protein IscU, which requires iron provided by an unknown source, sulfur provided in the form of cysteine bound persulfides (Cys-SSH) by the cysteine desulfurase IscS, and electrons provided by the ferredoxin–ferredoxin reductase complex Fdx-FdxR from NADPH. Then, specialized chaperones transfer [2Fe-2S] clusters to recipient acceptors. Despite previous studies on the core assembly machinery, the mechanistic details of the [2Fe-2S] cluster assembly process have remained poorly understood due to the experimental difficulties in trapping the relevant intermediates.
The team Biochemistry of Metalloproteins and Associated Diseases at the Institute of Integrative Biology of the Cell in Gif-Sur-Yvette (I2BC, UMR 9198, CNRS – CEA - Paris-Saclay University) managed to dissect this process step by step and to isolate several key intermediates using a functional reconstitution of the Escherichia coli ISC machinery. They used a combination of biochemical techniques to trap these intermediates: anaerobic reconstitution, persulfide detection assays, kinetics, UV-visible, circular dichroism, and to characterize them by spectroscopic methods: electron paramagnetic spectroscopy (EPR), nuclear magnetic resonance (NMR) and native mass spectrometry (nMS) in collaboration with teams at Aix-Marseille University (BIP), Gif-Sur-Yvette (ICSN) and Strasbourg (IPHC). They show that the assembly of [2Fe-2S] clusters is initiated by iron binding to IscU, which triggers persulfide insertion by IscS in the vicinity of the iron-binding site of IscU upon the formation of a complex between IscU and IscS. The persulfide in IscU binds to the iron center and is cleaved into sulfide by the Fdx-FdxR complex, which leads to the formation of a Fe-SH intermediate, referred to as the [1Fe-1S] precursor. Then, IscU dissociates from IscS, dimerizes and generates a bridging [2Fe-2S] cluster by fusion of two [1Fe-1S] precursors. The IscU dimer ultimately dissociates into a monomer, ready to transfer its [2Fe-2S] cluster to acceptors. The data also indicate that the bridging cluster is initially in the super-reduced state [2Fe-2S]0 and releases two electrons to the ferredoxin enzyme, thereby leading to an oxidised [2Fe-2S]2+ state as the final product.
These data provide a comprehensive description of mechanism of [2Fe-2S] clusters assembly by the bacterial ISC machinery, highlighting the formation of key intermediates through a tightly concerted process. This stepwise dissection further supports findings in eukaryotes, including iron loading, persulfidation and dimerization of IscU, which point to an evolutionary conservation of the assembly process. More information: https://www.nature.com/articles/s41589-024-01818-8 Contact: Benoit D'Autréaux benoit.dautreaux@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
January 17, 2025 4:17 AM
|
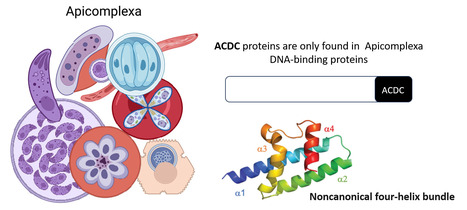
Rocking Science: New ACDC Protein Fold Unveiled
The Apicomplexa-specific ACDC domain adopts a never before seen protein fold. In collaboration with the Nessler team at the I2BC and the Llinás lab at PSU, US, we reveal that the ACDC domain, only found in DNA-binding proteins of the Apicomplexa phylum, grouping several important human pathogens including malaria, has a never before seen protein fold. We also identify potential ligands that may be optimized in the future as protein inhibitors. More information:https://journals.iucr.org/paper?S2059798324012518 Conatct: Joana SANTOS joana.santos@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
December 4, 2024 11:38 AM
|
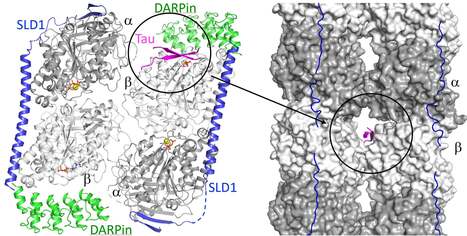
The structure of a Tau fragment bound to tubulin prompts new hypotheses on Tau mechanism and oligomerization
We have identified a novel tubulin surface targeted by the Tau protein. From these results, we formulate new hypotheses on the regulation of microtubules by Tau and on Tau oligomerization. Tau is a protein involved in the regulation of axonal microtubules in neurons. In pathological conditions, it forms filamentous aggregates which are molecular markers of the Alzheimer’s disease and of related neurodegenerative disorders collectively known as tauopathies. Structures of Tau in fibrils or bound to the microtubule have been reported. We have determined the structure of a Tau construct comprising the PHF6 motif, an hexapeptide involved in Tau aggregation, as a complex with tubulin. This Tau fragment binds as a dimer to a new site which, when transposed to the microtubule, would correspond to a pore between protofilaments. These results raise new hypotheses on Tau-induced microtubule assembly and stabilization and on Tau oligomerization. They reconcile apparently contradictory data from the literature, in particular concerning the coexistence of different binding modes of Tau to the microtubule. More information : https://academic.oup.com/pnasnexus/article/3/11/pgae487/7850850 Contact : Benoît GIGANT benoit.gigant@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
October 17, 2024 6:14 AM
|
Introduction to biological crystallography
Following the French MOOC 'Voyage au cœur du vivant avec des rayons X : la cristallographie', available on FUN-MOOC from 2017 to 2022, and the book in French 'Introduction à la cristallographie biologique', we now present an English version. Following the French MOOC 'Voyage au cœur du vivant avec des rayons X : la cristallographie', available on FUN-MOOC from 2017 to 2022, and the book in French 'Introduction à la cristallographie biologique', we now present an English version.
This book is primarily designed for beginners in biological crystallography.
It provides an introduction to the different steps of biological crystallography, from the crystallisation to the three-dimensional structure of a macromolecule. The QR codes at the end of each chapter give you access to the videos in French, that are part of the MOOC. English subtitles are available for all the videos. In the early stages of learning biological crystallography, the digital version of this book can therefore be a particularly useful companion. - Marie-Hélène Le Du, Pierre Legrand, Serena Sirigu, Sylvain Ravy - EDP Sciences & Science Press Introduction to biological crystallography link: https://bit.ly/3zBC6p5

|
Scooped by
I2BC Paris-Saclay
September 23, 2024 10:22 AM
|
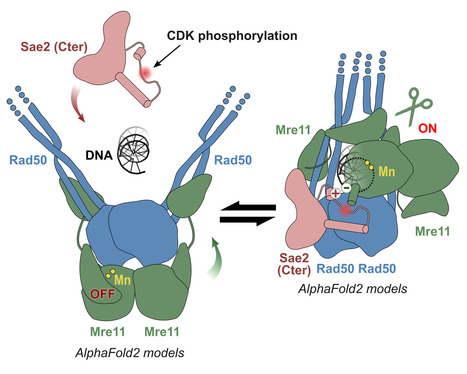
Molecular insights into the activation of Mre11-Rad50 endonuclease activity by Sae2/CtIP
Early Steps of DNA Recombination: AlphaFold2 breaks a lock. The human MRN nuclease, formed by the association of two proteins, Mre11 and Rad50, is a key player in homologous recombination and meiosis, two processes that utilize DNA break repair mechanisms. In eukaryotes, the Mre11 nuclease is equipped with a molecular lock that controls the activation of the enzyme: its activation is triggered during certain phases of the cell cycle (entry into G2, when DNA is duplicated to allow for the recombination process), when the lock is phosphorylated. This locking protein is called CtIP in humans and Sae2 in the yeast Saccharomyces cerevisiae. It is a protein with a largely disordered structure, which remained a significant challenge for structural characterization for a number of years. In a study published in Molecular Cell, teams from I2BC, the Curie Institute, and IRB shed light on how the human MRN nuclease functions. The I2BC team modeled the structure of the yeast MRN complex using the AlphaFold2 algorithm. Interestingly, the program hesitates when modeling between two states: inactive (auto-inhibited) and active. Notably, the addition of phosphorylated Sae2 favors the active conformation in AlphaFold2’s predictions: Sae2 appears to unlock the MRN complex by establishing a synergistic set of interactions centered on the phosphorylation of a serine residue in Sae2. The structural predictions obtained with AlphaFold2, supported by in vitro and in vivo experiments, highlight that the phosphorylated Sae2/CtIP protein creates a network of interactions with MRN that promotes the release of its auto-inhibition. All these findings illustrate how an apparently disordered protein can lift the auto-inhibition of a nuclease and thus control the switch between different repair pathways, which is important in humans for cell cycle progression and meiosis. More information: https://www.sciencedirect.com/science/article/pii/S1097276524004428?via%3Dihub Contact: Raphaël GUEROIS raphael.guerois@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
July 2, 2024 10:31 AM
|
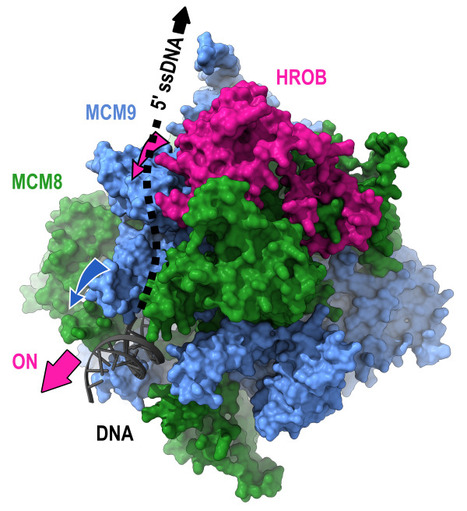
Mechanism of DNA unwinding by MCM8-9 in complex with HROB
Researchers from the AMIG team (I2BC department), in collaboration with the IRB (Switzerland), have modeled the interaction between HROB and the helicases MCM8-MCM9, some mutations of which predispose individuals to infertility or cancer. They demonstrate that HROB promotes the catalytic activity of the MCM8-MCM9 complex but does not play a role in its recruitment or stability. The proteins MCM8 and MCM9 have recently been discovered as involved in multiple processes, normal and pathological, related to DNA replication, meiosis, homologous recombination, and mismatch repair. These proteins are helicases that have the ability to move along DNA and separate the two DNA strands. Variants of these proteins may predispose carriers to disorders such as infertility and cancer. In 2019, a third partner, HROB, was identified as associated with these two helicases without its mechanisms of action being understood. Since HROB is capable of interacting with DNA, three functional hypotheses can be considered:
1) HROB participates in the recruitment of helicases on DNA,
2) HROB stabilizes the assembly of MCM8 and MCM9 on DNA,
3) HROB promotes the catalytic activity of pre-assembled helicases. In a study published in the journal Nature Communications, researchers from the AMIG team at the Institute of Integrative Biology of the Cell (I2BC) collaborated with a team from the Institute of Research in Biomedicine (IRB) in Switzerland to establish different structural models of the MCM8-MCM9-HROB complex. Guided by these models, a set of simple and compensatory mutations allowed the dissection of the functional role of HROB. The teams showed that only the third hypothesis was correct and that HROB is an essential factor for activating the MCM8-MCM9 helicase but not for its recruitment or stability. The structural model suggests that by transiently altering the conformation of a MCM9 subunit, HROB stimulates the translocation of the helicase along the DNA. Based on this model, it was even possible to design mutations that increase the efficiency of the helicase in the presence of HROB. More information: https://www.nature.com/articles/s41467-024-47936-8 Contact: Raphaël Guerois raphael.guerois@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
April 15, 2024 9:03 AM
|
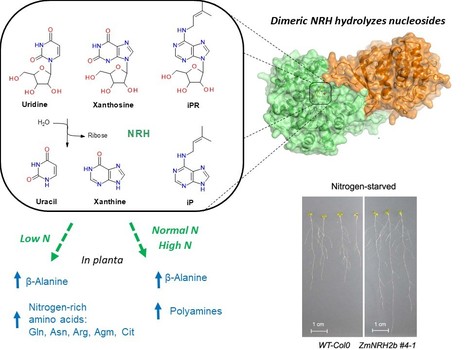
Plant nucleoside N-ribohydrolases: riboside binding and role in nitrogen storage mobilization
Plants can benefit from fast nucleoside breakdown Cells save their energy during nitrogen starvation by selective autophagy of ribosomes and degradation of RNA to ribonucleotides and nucleosides. Nucleosides are hydrolyzed by nucleoside N-ribohydrolases (nucleosidases, NRHs). Subclass I of NRHs preferentially hydrolyzes the purine ribosides while subclass II is more active towards uridine and xanthosine. Here, we performed a crystallographic and kinetic study to shed light on nucleoside preferences among plant NRHs followed by in vivo metabolomic and phenotyping analyses to reveal the consequences of enhanced nucleoside break-down. We report the crystal structure of Zea mays NRH2b (subclass II) and NRH3 (subclass I) in complexes with the substrate analog forodesine. Purine and pyrimidine catabolism are inseparable because nucleobase binding in the active site of ZmNRH is mediated via a water network and is thus unspecific. Dexamethasone-inducible ZmNRH overexpressor lines of Arabidopsis thaliana, as well as double nrh knockout lines of moss Physcomitrium patents, reveal a fine control of adenosine in contrast to other ribosides. ZmNRH overexpressor lines display an accelerated early vegetative phase including faster root and rosette growth upon nitrogen starvation or osmotic stress. Moreover, the lines enter the bolting and flowering phase much earlier. We observe changes in the pathways related to nitrogen-containing compounds such as β-alanine and several polyamines, which allow plants to reprogram their metabolism to escape stress. Taken together, crop plant breeding targetting enhanced NRH-mediated nitrogen recycling could therefore be a strategy to enhance plant growth tolerance and productivity under adverse growth conditions. more information: https://onlinelibrary.wiley.com/doi/10.1111/tpj.16572 contact: Solange Morera solange.morera@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
April 15, 2024 8:44 AM
|
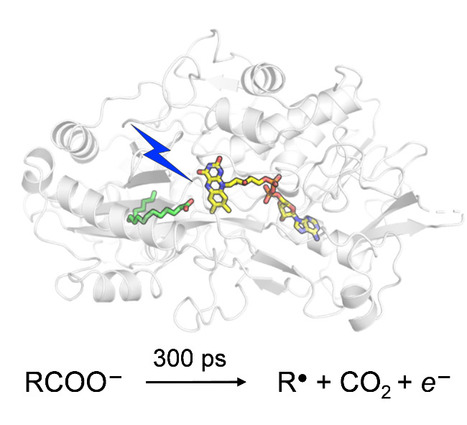
Catalytic Mechanism of Fatty Acid Photodecarboxylase: on the Detection and Stability of the Initial Carbonyloxy Radical Intermediate
Quality trumps quantity: photoenzymatic decarboxylation of fatty acids does indeed occur in 300 ps. Despite the dismissal by our "slightly overproductive" competitors, citing it as 30x slower, our original findings stand strong! In fatty acid photodecarboxylase (FAP), light-induced formation of the primary radical product RCOO● from fatty acid RCOO– occurs in 300 ps, upon which CO2 is released quasi-immediately. Based on the hypothesis that aliphatic RCOO●(spectroscopically uncharacterized because unstable) absorbs in the red similarly to aromatic carbonyloxy radicals such as 2,6-dichlorobenzoyloxy radical (DCB●), much longer-lived linear RCOO● has been suggested recently. We performed quantum chemical reaction pathway and spectral calculations. Thesecalculations are in line with the experimental DCB● decarboxylation dynamics and spectral properties and show that in contrast to DCB●, aliphatic RCOO● radicals a) decarboxylate with a very low energetic barrier and on the timescale of a few ps and b) exhibit little red absorption. A time-resolved infrared spectroscopy experiment confirms very rapid, <<300 ps RCOO● decarboxylation in FAP. We argue that this property is required for the observed high quantum yield of hydrocarbons formation by FAP. more information: https://onlinelibrary.wiley.com/doi/10.1002/anie.202401376 contact: Pavel MULLER <pavel.muller@i2bc.paris-saclay.fr>
|

|
Scooped by
I2BC Paris-Saclay
February 16, 10:02 AM
|
Snapshots of cotranslational N-myristoylation reveal NMT as a ribosome-associated chaperone.
NMT in a new light: associated with the ribosome via the NAC complex, this enzyme not only adds lipid tags to the N-termini of nascent proteins but also acts as a chaperone, cooperating with MetAPs to ensure proper folding and delivery. N-myristoylation is an essential cotranslational lipid modification catalyzed by N-myristoyltransferases (NMTs). Structural and cellular analyses reveal that NMT1 associates with the ribosomal tunnel exit via the nascent polypeptide–associated complex (NAC) and acts sequentially after MetAP-mediated initiator methionine removal, in contrast to previously described simultaneous cotranslational modification assemblies. Unexpectedly, NMT1 also exhibits chaperone-like activity, expanding its functional repertoire in cotranslational protein biogenesis. More information : https://www-nature-com.insb.bib.cnrs.fr/articles/s41467-025-67962-4 Contact : Thierry Meinnel and Carmela Giglione thierry.meinnel@i2bc.paris-saclay.fr and carmela.giglione@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
December 8, 2025 5:52 AM
|
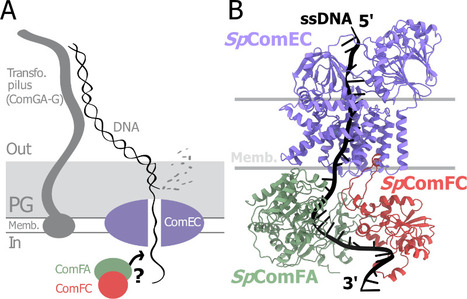
A tripartite protein complex promotes DNA transport during natural transformation in Firmicutes.
From structural modeling to natural transformation in bacteria: identification of a key protein complex for DNA transport across the membrane of Firmicutes. Transformation is a key mechanism of horizontal gene transfer, central to bacterial adaptation. This evolutionarily conserved process allows bacteria to integrate exogenous genetic material into their genome, thereby facilitating, for example, the spread of antibiotic resistance.
In a study published in the journal PNAS, scientists from I2BC (CEA/CNRS/UPSaclay, Gif-sur-Yvette) and DRCM/IBFJ (CEA/UPSaclay/UP Cité, Fontenay-aux-Roses), in collaboration with a laboratory at CBI (CNRS/University of Toulouse), identified a protein complex involved in the transport of single-stranded DNA across the membrane during transformation. Using AlphaFold, they structurally modeled this three-protein complex, which is highly conserved in the phylum Firmicutes, in interaction with single-stranded DNA. The structural model allowed them to identify a possible path for DNA through a conserved channel in one of the three proteins, a transmembrane protein, and then along a groove formed by the other two proteins. This model was validated by a robust experimental strategy in the bacterium Streptococcus pneumoniae, by measuring the impact on transformation efficiency of disruptive mutations in several protein–protein and protein–DNA interfaces. The transmembrane channel was found to be conserved in a structural model in Helicobacter pylori, and its importance for transformation was also experimentally confirmed in this bacterium. This study sheds light on the molecular mechanisms of bacterial transformation and demonstrates the power of macromolecular structure prediction to generate molecular hypotheses and guide functional experiments. More information: https://doi.org/10.1073/pnas.2511180122 Contact : Jessica Andreani jessica.andreani@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
September 5, 2025 3:50 AM
|
Restriction of Ku translocation protects telomere ends
How Ku inward translocation can be restricted to protect telomere ends from NHEJ without altering other important functions. Safeguarding chromosome ends against fusions via nonhomologous end joining (NHEJ) is essential for genome integrity. Paradoxically, the conserved NHEJ core factor Ku binds telomere ends. How it is prevented from promoting NHEJ remains unclear, as does the mechanism that allows Ku to coexist with telomere-protective DNA binding proteins, Rap1 in Saccharomyces cerevisiae. Here, we find that Rap1 directly inhibits Ku’s NHEJ function at telomeres. A single Rap1 molecule near a double-stand break suppresses NHEJ without displacing Ku in cells. Furthermore, Rap1 and Ku form a complex on short DNA duplexes in vitro. Cryo-EM shows Rap1 blocks Ku’s inward translocation on DNA – an essential step for NHEJ at DSBs. Nanopore sequencing of telomere fusions confirms this mechanism protects native telomere ends. These find- ings uncover a telomere protection mechanism where Rap1 restricts Ku’s inward translocation. This switches Ku from a repair-promoting to a protective role preventing NHEJ at telomeres. More information : https://doi.org/10.1038/s41467-025-61864-1 Contact : Philippe Cuniasse philippe.cuniasse@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
April 11, 2025 5:59 AM
|
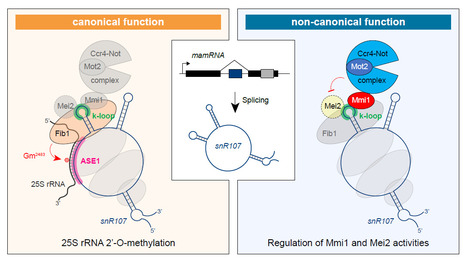
A bifunctional snoRNA guides rRNA 2’-O-methylation and scaffolds gametogenesis effectors
This study uncovers a fission yeast small nucleolar RNA (snoRNA) that guides ribosomal RNA 2’- O-methylation and modulates the activities of RNA-binding proteins involved in gametogenesis, expanding our vision of the non-canonical functions exerted by snoRNAs. Small nucleolar RNAs are non-coding transcripts that guide chemical modifications of RNA substrates and modulate gene expression at the epigenetic and post-transcriptional levels. However, the extent of their regulatory potential and the underlying molecular mechanisms remain poorly understood. In a collaborative work with the I2BC B3S department and NGS facility, the epiRNA-Seq facility in Nancy and the Palancade lab at the Institut Jacques Monod, we have identified a conserved, previously unannotated intronic C/D-box snoRNA, termed snR107, hosted in the fission yeast long non-coding RNA mamRNA and carrying two independent cellular functions. On the one hand, snR107 guides site-specific 25S rRNA 2’-O-methylation and promotes pre-rRNA processing and 60S subunit biogenesis. On the other hand, snR107 associates with the gametogenic RNA-binding proteins Mmi1 and Mei2, mediating their reciprocal inhibition and restricting meiotic gene expression during sexual differentiation. Both functions require distinct cis-motifs within snR107, including a conserved 2’-O-methylation guiding sequence. Together, our results position snR107 as a dual regulator of rRNA modification and gametogenesis effectors, expanding our vision on the non-canonical functions exerted by snoRNAs in cell fate decisions. More Information: https://www.nature.com/articles/s41467-025-58664-y Contact: Mathieu Rougemaille mathieu.rougemaille@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
April 3, 2025 6:36 AM
|
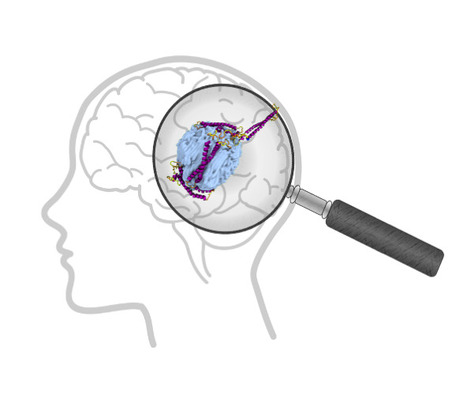
Exploring the Interaction of Human α‐Synuclein with Polyethylene Nanoplastics: Insights from Computational Modeling and Experimental Corroboration
Can the presence of microplastics in the brain affect proteins? Plastics are now part of our daily lives because they are used in so many different ways. They are therefore a major source of pollution. Their chemical stability has consequently become a major concern for the environment and health, given their presence in all ecosystems. The penetration of plastic particles into living organisms through ingestion or inhalation has now been widely demonstrated. Micro- and nanoplastics are found throughout the human body, including in the brain, which raises the question of their potential toxicity. In a biological environment, the plastic particle does not remain naked but interacts with the surrounding molecules to form a biocorona. In this study, Tripathi et al. investigate the binding behavior of human α-synuclein (hαSn) with polyethylene (PE)-based plastics using molecular dynamics simulations and experimental methods. Their simulations show that (a) hαSn folds into a compact conformation to enhance intramolecular interactions, (b) non-oxidised PE nanoplastics facilitate the rapid adsorption of hαSn onto its surface with a change in the structural properties of hαSn, and (c) oxidised nanoplastics fail to capture hαSn. The experimental dynamic light scattering and adsorption isotherms are in good agreement with simulations. The observed formation of the plastic nanoparticle complex with hαSn can be proposed as a plausible pathogenic driving force in neuronal dysfunction and subsequent neurological damage. More information : https://doi.org/10.1021/acs.biomac.4c00918 Contact: Yves Boulard yves.boulard@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
February 10, 2025 9:09 AM
|
Tutorial videos for ChimeraX

|
Scooped by
I2BC Paris-Saclay
December 17, 2024 11:56 AM
|
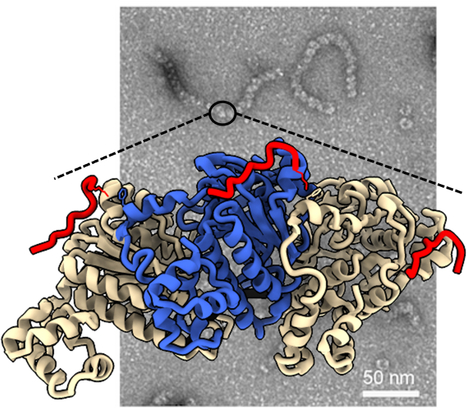
DciA, the Bacterial Replicative Helicase Loader, promotes LLPS in the presence of ssDNA.
This study shows that DciA, the bacterial replicative helicase loader, promotes condensates in the presence of DNA. This opens the way to the possibility of non-membrane compartments in bacteria. The goal could be to concentrate the players involved in replication and thus facilitate it. The loading of the bacterial replicative helicase DnaB is an essential step for genome replication and depends on the assistance of accessory proteins. Several of these proteins have been identified across the bacterial phyla. DciA is the most common loading protein in bacteria, yet the one whose mechanism is the least understood. We have previously shown that DciA from Vibrio cholerae is composed of a globular domain followed by an unfolded extension and demonstrated its strong affinity for DNA. Here, drawing on the skills of two I2BC facilities, Light microscopy (Imagerie-Gif) and PIM (Structural biology), we characterize the condensates formed by VcDciA upon interaction with a short single-stranded DNA substrate. We demonstrate the fluidity of these condensates using light microscopy and address their network organization through electron microscopy, thereby bridging events to conclude on a liquid-liquid phase separation behavior. Additionally, we observe the recruitment of DnaB in the droplets, concomitant with the release of DciA. We show that the well-known helicase loader DnaC from Escherichia coli is also competent to form these phase-separated condensates in the presence of ssDNA. Our phenomenological data are still preliminary as regards the existence of these condensates in vivo, but open the way for exploring the potential involvement of DciA in the formation of non-membrane compartments within the bacterium to facilitate the assembly of replication players on chromosomal DNA. More information : https://pubmed.ncbi.nlm.nih.gov/39603490/ Contact : Sophie CHERUEL sophie.quevillon-cheruel@i2bc.paris-saclay.fr and Stéphanie MARSIN stephanie.marsin@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
November 26, 2024 3:28 AM
|
Perylene-derivative singlet exciton fission in water solution
Ultrafast formation of triplet statates by singlet fission in water solution. We provide evidence that singlet fission—a process where one excited state splits into two—can occur in water-soluble compounds. Specifically, we studied perylene-3,4,9,10-tetracarboxylate, a molecule that forms transient disordered dimers. Using a combination of advanced techniques like time-resolved absorption and fluorescence spectroscopy, nuclear magnetic resonance (NMR) spectroscopy, and theoretical modeling, we were able to detect the key features of singlet fission and map out the behavior of the molecular assemblies. Our results show that the constant structural changes within these dimers play a crucial role in steering the process toward either singlet fission or charge separation. The quantum efficiency of producing triplet states is over 100%, with these states lasting for nanoseconds—highlighting how disordered systems can drive highly efficient photophysical processes. More information: https://pubs.rsc.org/en/content/articlehtml/2024/sc/d4sc04732j Contact: Manuel LLANSOLA <manuel.llansola@I2bc.paris-saclay.fr>

|
Scooped by
I2BC Paris-Saclay
October 17, 2024 5:43 AM
|
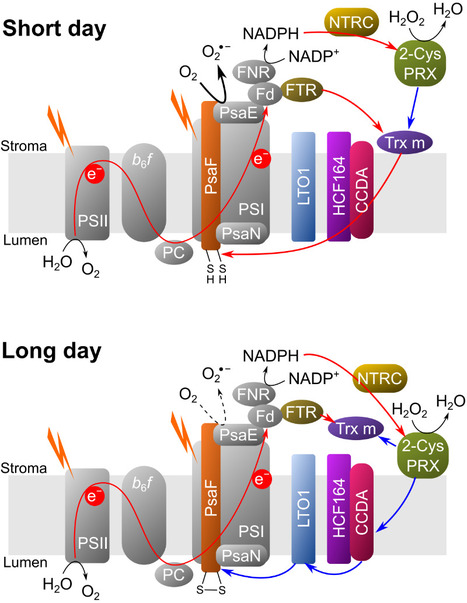
A complex and dynamic redox network regulates oxygen reduction at photosystem I in Arabidopsis
A system of redox enzymes facilitates a series of reactions that regulate oxygen reduction at photosystem I in plants. In a study published in Plant Physiology, scientists from I2BC and IPS2 studied the redox regulation of superoxide production at the photosynthetic electron transfer level (pseudocyclic flow). Plants must optimise their molecular strategies to adapt to the light conditions of their environment. Reactive oxygen species (ROS) can play a role as signalling molecules but can also be potentially harmful. To manage light-induced ROS and maintain proper photosynthetic function, thiol-dependent redox enzymes play a crucial role in the redox regulation of the chloroplast. It has been shown that the main redox enzymes involved in controlling the ROS generated during photosynthesis are m-type thioredoxins, NADPH-dependent reductase C (NTRC) and peroxiredoxin 2-Cys. The interaction of these enzymes functions as a redox regulatory network, where 2-Cys PRX can be reduced by NTRC and Trx but can also re-oxidise the reduced thioredoxins, thus providing a system for rapid acclimatisation to changes in the light regime. The researchers showed that the membrane localisation of Trx m and NTRC varies as a function of photoperiod. In addition, their results show that it is the PSI itself that is redox regulated. The new results of this study make it possible to generate a new model of PSI redox regulation. This model may guide further research into the redox regulation of alternative electron transport pathways under conditions of fluctuating light or abiotic stress. More information: DOI: 10.1093/plphys/kiae501 Contact: Anja KRIEGER-LISZKAY anja.liszkay@i2bc.paris-saclay.fr

|
Scooped by
I2BC Paris-Saclay
July 26, 2024 8:24 AM
|
BRCA2 stabilizes DMC1 nucleoprotein filaments in meiosis
To repair programmed double-strand breaks in meiosis, the DNA repair protein BRCA2 binds to the recombinase DMC1 either monomeric or assembled on single-stranded DNA through two different interfaces, and stabilizes DMC1 nucleoprotein filaments. The BReast CAncer type 2 susceptibility protein (BRCA2), a tumor suppressor mutated in breast, ovarian and prostate cancers, plays a major role in the repair of DNA double-strand breaks by homologous recombination, both in somatic cells and during meiosis. BRCA2 interacts with the ubiquitous recombinase RAD51 , as well as the meiotic recombinase DMC1, and facilitates their loading at double-strand break sites. BRCA2 interacts with these recombinases via FxxA and FxPP motifs (called A and P motifs, respectively). In a study published recently in the journal Nucleic Acids Research, scientists from the INTGEN team at the I2BC (Université Paris Saclay, CEA, CNRS, Gif-sur-Yvette) and the PROXIMA-1 beamline at the SOLEIL synchrotron (Université Paris Saclay) solved the crystal structure of the complex between a BRCA2 fragment containing a P motif (PhePP) and the DMC1 protein. Together with the team of A. Zelensky and R. Kanaar (Erasmus Medical Center, Rotterdam), they showed that A and P motifs bind to distinct sites on the ATPase domain of recombinases. The P motif interacts with a site that is accessible in the octamers of DMC1 and the nucleoprotein filaments formed by DMC1 loaded onto single-stranded DNA. Furthermore, in collaboration with scientists from the Institut Gustave Roussy (Université Paris Saclay), they revealed that this interaction involves two adjacent DMC1 protomers, thereby increasing the stability of the nucleoprotein filaments. These results help to explain why the region encoded by exons 12 to 14 of the BRCA2 gene (the PhePP motif being encoded by exon 14) is essential during meiotic homologous recombination in mice (work by the teams of A. Zelensky and W. Baarends, Erasmus Medical Center, Rotterdam, Netherlands). More information: https://doi.org/10.1093/nar/gkae452 Contact: Simona MIRON <simona.miron@i2bc.paris-saclay.fr> & Sophie ZINN <sophie.zinn@i2bc.paris-saclay.fr>

|
Scooped by
I2BC Paris-Saclay
April 15, 2024 9:06 AM
|
Physics and Biological Systems 2024
Mark your calendars for Jun. 26-28, 2024! Join us at Ecole Polytechnique in Palaiseau for the 7th International Conference on Physics and Biological Systems, where top-notch scientists converge to explore the dynamic interplay between physics and life sciences. The 7th International Conference on Physics and Biological Systems will be held on Jun. 26-28 2024 at Ecole Polytechnique in Palaiseau, in the south of Paris. It aims to bring together a broad range of physical and life scientists working at the interface between the two disciplines around in-depth talks by first-rate international speakers. Attendance will be limited to 200 participants. We look forward to welcoming you in Palaiseau! more information: https://www.lptms.universite-paris-saclay.fr/physbio2024/ contact: Pavel MULLER <pavel.muller@i2bc.paris-saclay.fr>

|
Scooped by
I2BC Paris-Saclay
April 15, 2024 8:50 AM
|
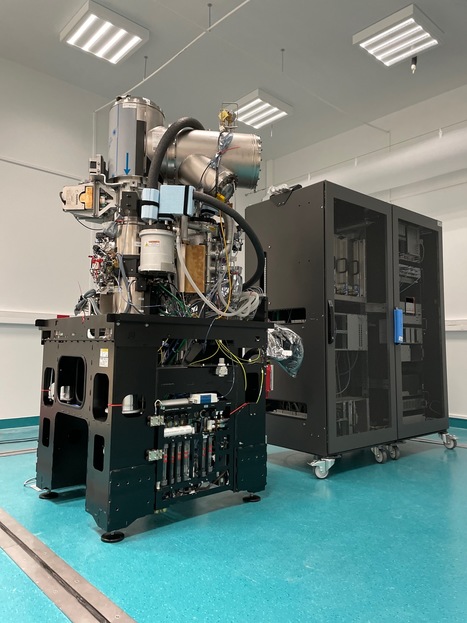
A high-end Glacios2 cryo-electron microscope at I2BC
A Glacios2 200 kV has been delivered to I2BC, heralding, together with a Titan Krios 300 kV at synchrotron SOLEIL, a new era for cryo-electron microscopy in the Paris-Saclay area Cryo-electron microscopy (cryo-EM) has become an indispensable tool in structural biology, enabling researchers to unravel the complexities of biological macromolecules and understand their roles in various cellular processes and diseases. These last ten years the field has undergone a revolution, with technical advances that have made atomic resolution reconstructions of large macromolecular complexes possible (the so-called 'resolution revolution'), I2BC received the latest generation 200kV microscope, Glacios2, on March 27, 2024. Joint operation is planned with the 300kV Titan Krios also just delivered at synchrotron SOLEIL. This will allow researchers to study structures of living machines at work in fine detail, both for purified and/or reconstituted complexes and in cells. These two cryo-electron microscopes are gathering scientists from all over the Paris-Saclay area, sparking further interest in future developments at the interface beween structural and cellular biology. contact: Stéphane Bressanelli (plt-cryoem@i2bc.paris-saclay.fr)
|



 Your new post is loading...
Your new post is loading...


![Cross-regulation of [2Fe–2S] cluster synthesis by ferredoxin-2 and frataxin. | I2BC Paris-Saclay | Scoop.it](https://img.scoop.it/Ve3yN6adKt_5lzSWPuYkATl72eJkfbmt4t8yenImKBVvK0kTmF0xjctABnaLJIm9)


![Complete description of the biosynthetic process of [2Fe-2S] clusters | I2BC Paris-Saclay | Scoop.it](https://img.scoop.it/THKpAN1-eQiW-1DEJF_ZmDl72eJkfbmt4t8yenImKBVvK0kTmF0xjctABnaLJIm9)