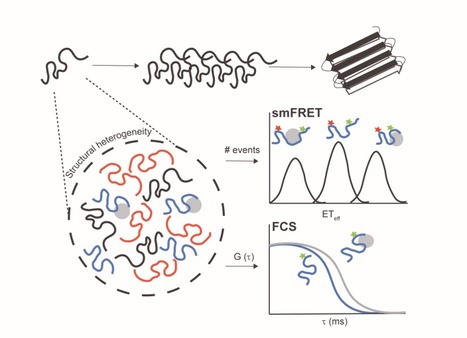
Intrinsically disordered proteins/regions (IDPs/IDRs) are widespread in the human proteome and play critical roles in distinct biological processes and dysfunction. Contrary to the classical “structure–function” paradigm, IDPs lack stable secondary/tertiary structures under physiological conditions, and instead adopt a dynamic ensemble of multiple conformations. Moreover, several neurodegenerative disorders are associated with the pathological self-assembly of neuronal IDPs, including tau (Alzheimer’s disease), α-synuclein (Parkinson’s disease), and huntingtin exon 1 (Huntington’s disease). Therefore, there is an emerging medical interest in understanding their physical and structural features. However, their characterization is inherently challenging by traditional ensemble and time-averaging methodologies. A recent review co-authored by Ana Melo from BSIRG-iBB, and published in Frontiers in Molecular Neurosciences the advantages of employing cutting-edge single-molecule fluorescence techniques are discussed to characterize the conformational ensemble of neurodegeneration-promoting IDPs under normal and disease conditions to obtain insights into their gain- or loss-of-function.



 Your new post is loading...
Your new post is loading...