A recent proof-of-concept study has brought prime editing one step closer to clinical trials. Dr. Dong Hyun Jo explains how he and his colleagues demonstrated the use of this nascent technology in vivo for the first time.
Follow, research and publish the best content
Get Started for FREE
Sign up with Facebook Sign up with X
I don't have a Facebook or a X account
Already have an account: Login
 Your new post is loading... Your new post is loading...
 Your new post is loading... Your new post is loading...
|
|








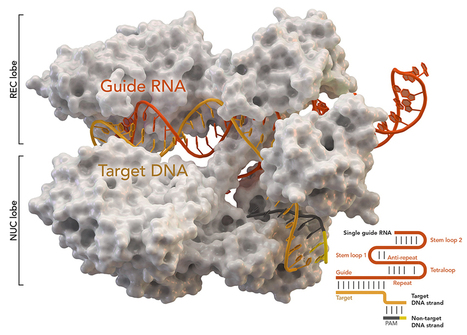









Principal editing (PE) is thought to be a safer and more precise alternative to CRISPR-Cas9. Using a Cas9 nickase and reverse transcriptase, master editors can create small insertions and deletions without creating double-strand breaks in the DNA and without requiring a donor template for repair. There are four different types of master editing systems. PE1 and PE2 use a Cas9 nickase fused to a reverse transcriptase and a master editing guide RNA (pegRNA). PE3 and PE3b are derivatives that use the PE2 system in conjunction with an additional single guide RNA (sgRNA). Master editing was tested in hereditary tyrosinemia, an autosomal recessive disease caused by a loss-of-function mutation in the fumarylacetoacetate hydrolase (Fah) gene. The pathogenic mutation prevents patients from producing the enzyme fumarylacetoacetate hydrolase, which contributes to the normal breakdown of the amino acid tyrosine. A research team delivered PE3 components by hydrodynic injection or PE2 using adenovirus in a mut-mut FAH mouse model. Compared to the original Cas9 which induces double-strand breaks, main editing, in particular PE2, was shown to be a safer strategy in terms of off-target effects. Extensive analyses showed that the off-target effects of master editing were less than 0.1%.